Step 2: SNP QC [7.batches_myo]
Belinda Cornes
2023-04-13
Last updated: 2023-04-13
Checks: 5 2
Knit directory: Serreze-T1D_Workflow/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown is untracked by Git. To know which version of the R
Markdown file created these results, you’ll want to first commit it to
the Git repo. If you’re still working on the analysis, you can ignore
this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20220210) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Using absolute paths to the files within your workflowr project makes it difficult for you and others to run your code on a different machine. Change the absolute path(s) below to the suggested relative path(s) to make your code more reproducible.
| absolute | relative |
|---|---|
| /Users/corneb/Documents/MyJax/CS/Projects/Serreze/qc/workflowr/Serreze-T1D_Workflow | . |
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 51e3845. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .DS_Store
Ignored: analysis/.DS_Store
Untracked files:
Untracked: analysis/0.1.1_preparing.data_bqc_7.batches_myo.R
Untracked: analysis/0.1.1_preparing.data_bqc_7.batches_myo.Rmd
Untracked: analysis/0.1_samples_batch_20230124.Rmd
Untracked: analysis/0.1_samples_batch_20230225.Rmd
Untracked: analysis/0.1_samples_batch_20230313.Rmd
Untracked: analysis/0.2_haplotype_comparison_bqc_7.batches_myo_minprob.R
Untracked: analysis/0.2_haplotype_comparison_bqc_7.batches_myo_minprob.Rmd
Untracked: analysis/2.1_sample_bqc_7.batches_myo.R
Untracked: analysis/2.1_sample_bqc_7.batches_myo.Rmd
Untracked: analysis/2.1_sample_bqc_7.batches_myo.Rmd.R
Untracked: analysis/2.2.1_snp_qc_7.batches_myo.R
Untracked: analysis/2.2.1_snp_qc_7.batches_myo.Rmd
Untracked: analysis/2.2.1_snp_qc_7.batches_myo_mis.R
Untracked: analysis/2.2.1_snp_qc_7.batches_myo_mis.Rmd
Untracked: analysis/2.4_preparing.data_aqc_7.batches_myo.Rmd
Untracked: analysis/2.4_preparing.data_aqc_7.batches_myo_mis.Rmd
Untracked: analysis/2.4_preparing.data_aqc_7.batches_myo_misd.R
Untracked: analysis/3.1_phenotype.qc_corrected_7.batches_myo.Rmd
Untracked: analysis/3.1_phenotype.qc_corrected_7.batches_myo_mis.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_het-ici-myo-yes.vs.het-ici-myo-no_snpsqc_dis_no-x_updated_7.batches_myo.R
Untracked: analysis/4.1.1_qtl.analysis_binary_het-ici-myo-yes.vs.het-ici-myo-no_snpsqc_dis_no-x_updated_7.batches_myo.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_het-ici-myo-yes.vs.het-ici-myo-no_snpsqc_dis_no-x_updated_7.batches_myo_mis.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_het-ici.vs.het-pbs_snpsqc_dis_no-x_updated_7.batches_myo.R
Untracked: analysis/4.1.1_qtl.analysis_binary_het-ici.vs.het-pbs_snpsqc_dis_no-x_updated_7.batches_myo.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_het-ici.vs.het-pbs_snpsqc_dis_no-x_updated_7.batches_myo_mis.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici-myo-yes.vs.ici-myo-no_snpsqc_dis_no-x_updated_7.batches_myo.R
Untracked: analysis/4.1.1_qtl.analysis_binary_ici-myo-yes.vs.ici-myo-no_snpsqc_dis_no-x_updated_7.batches_myo.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici-myo-yes.vs.ici-myo-no_snpsqc_dis_no-x_updated_7.batches_myo_mis.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici-sick.vs.ici-eoi_snpsqc_dis_no-x_updated_7.batches_myo.R
Untracked: analysis/4.1.1_qtl.analysis_binary_ici-sick.vs.ici-eoi_snpsqc_dis_no-x_updated_7.batches_myo.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici-sick.vs.ici-eoi_snpsqc_dis_no-x_updated_7.batches_myo_mis.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici.vs.pbs_snpsqc_dis_no-x_updated_7.batches_myo.R
Untracked: analysis/4.1.1_qtl.analysis_binary_ici.vs.pbs_snpsqc_dis_no-x_updated_7.batches_myo.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici.vs.pbs_snpsqc_dis_no-x_updated_7.batches_myo_mis.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_myo-yes.vs.myo-no_snpsqc_dis_no-x_updated_7.batches_myo.R
Untracked: analysis/4.1.1_qtl.analysis_binary_myo-yes.vs.myo-no_snpsqc_dis_no-x_updated_7.batches_myo.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_myo-yes.vs.myo-no_snpsqc_dis_no-x_updated_7.batches_myo_mis.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_pbs-myo-yes.vs.pbs-myo-no_snpsqc_dis_no-x_updated_7.batches_myo.R
Untracked: analysis/4.1.1_qtl.analysis_binary_pbs-myo-yes.vs.pbs-myo-no_snpsqc_dis_no-x_updated_7.batches_myo.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_pbs-myo-yes.vs.pbs-myo-no_snpsqc_dis_no-x_updated_7.batches_myo_mis.Rmd
Untracked: analysis/debugging_comparing_old_Rmd/
Untracked: analysis/genotype.frequencies_het-ici-myo-yes.vs.het-ici-myo-no_7.batches_myo.Rmd
Untracked: analysis/genotype.frequencies_het-ici-myo-yes.vs.het-ici-myo-no_7.batches_myo_mis.Rmd
Untracked: analysis/genotype.frequencies_het-ici.vs.het-pbs_7.batches_myo.Rmd
Untracked: analysis/genotype.frequencies_het-ici.vs.het-pbs_7.batches_myo_mis.Rmd
Untracked: analysis/genotype.frequencies_ici-myo-yes.vs.ici-myo-no_7.batches_myo.Rmd
Untracked: analysis/genotype.frequencies_ici-myo-yes.vs.ici-myo-no_7.batches_myo_mis.Rmd
Untracked: analysis/genotype.frequencies_ici-sick.vs.ici-eoi_7.batches_myo.Rmd
Untracked: analysis/genotype.frequencies_ici-sick.vs.ici-eoi_7.batches_myo_mis.Rmd
Untracked: analysis/genotype.frequencies_ici.vs.pbs_7.batches_myo.Rmd
Untracked: analysis/genotype.frequencies_ici.vs.pbs_7.batches_myo_mis.Rmd
Untracked: analysis/genotype.frequencies_myo-yes.vs.myo-no_7.batches_myo.Rmd
Untracked: analysis/genotype.frequencies_myo-yes.vs.myo-no_7.batches_myo_mis.Rmd
Untracked: analysis/genotype.frequencies_pbs-myo-yes.vs.pbs-myo-no_7.batches_myo.Rmd
Untracked: analysis/genotype.frequencies_pbs-myo-yes.vs.pbs-myo-no_7.batches_myo_mis.Rmd
Untracked: analysis/index_7.batches_myo.Rmd
Untracked: analysis/power.analysis_het-ici-myo-yes.vs.het-ici-myo-no_7.batches_myo.Rmd
Untracked: analysis/power.analysis_het-ici-myo-yes.vs.het-ici-myo-no_7.batches_myo_mis.Rmd
Untracked: analysis/power.analysis_het-ici.vs.het-pbs_7.batches_myo.Rmd
Untracked: analysis/power.analysis_het-ici.vs.het-pbs_7.batches_myo_mis.Rmd
Untracked: analysis/power.analysis_ici-myo-yes.vs.ici-myo-no_7.batches_myo.Rmd
Untracked: analysis/power.analysis_ici-myo-yes.vs.ici-myo-no_7.batches_myo_mis.Rmd
Untracked: analysis/power.analysis_ici-sick.vs.ici-eoi_7.batches_myo.Rmd
Untracked: analysis/power.analysis_ici-sick.vs.ici-eoi_7.batches_myo_mis.Rmd
Untracked: analysis/power.analysis_ici.vs.pbs_7.batches_myo.Rmd
Untracked: analysis/power.analysis_ici.vs.pbs_7batches_myo_mis.Rmd
Untracked: analysis/power.analysis_myo-yes.vs.myo-no_7.batches_myo.Rmd
Untracked: analysis/power.analysis_myo-yes.vs.myo-no_7.batches_myo_mis.Rmd
Untracked: analysis/power.analysis_pbs-myo-yes.vs.pbs-myo-no_7.batches_myo.Rmd
Untracked: analysis/power.analysis_pbs-myo-yes.vs.pbs-myo-no_7.batches_myo_mis.Rmd
Untracked: data/GM_covar_4.batches_myo.csv
Untracked: data/GM_covar_7.batches_myo.csv
Untracked: data/bad_markers_all_4.batches_myo.RData
Untracked: data/bad_markers_all_7.batches_myo.RData
Untracked: data/covar_corrected.cleaned_het-ici-myo-yes.vs.het-ici-myo-no_7.batches_myo.csv
Untracked: data/covar_corrected.cleaned_het-ici-myo-yes.vs.het-ici-myo-no_7.batches_myo_mis.csv
Untracked: data/covar_corrected.cleaned_het-ici.vs.het-pbs_7.batches_myo.csv
Untracked: data/covar_corrected.cleaned_het-ici.vs.het-pbs_7.batches_myo_mis.csv
Untracked: data/covar_corrected.cleaned_ici-myo-yes.vs.ici-myo-no_7.batches_myo.csv
Untracked: data/covar_corrected.cleaned_ici-myo-yes.vs.ici-myo-no_7.batches_myo_mis.csv
Untracked: data/covar_corrected.cleaned_ici-sick.vs.ici-eoi_7.batches_myo.csv
Untracked: data/covar_corrected.cleaned_ici-sick.vs.ici-eoi_7.batches_myo_mis.csv
Untracked: data/covar_corrected.cleaned_ici.vs.pbs_7.batches_myo.csv
Untracked: data/covar_corrected.cleaned_ici.vs.pbs_7.batches_myo_mis.csv
Untracked: data/covar_corrected.cleaned_myo-yes.vs.myo-no_7.batches_myo.csv
Untracked: data/covar_corrected.cleaned_myo-yes.vs.myo-no_7.batches_myo_mis.csv
Untracked: data/covar_corrected.cleaned_pbs-myo-yes.vs.pbs-myo-no_7.batches_myo.csv
Untracked: data/covar_corrected.cleaned_pbs-myo-yes.vs.pbs-myo-no_7.batches_myo_mis.csv
Untracked: data/covar_corrected_het-ici-myo-yes.vs.het-ici-myo-no_7.batches_myo.csv
Untracked: data/covar_corrected_het-ici-myo-yes.vs.het-ici-myo-no_7.batches_myo_mis.csv
Untracked: data/covar_corrected_het-ici.vs.het-pbs_7.batches_myo.csv
Untracked: data/covar_corrected_het-ici.vs.het-pbs_7.batches_myo_mis.csv
Untracked: data/covar_corrected_ici-myo-yes.vs.ici-myo-no_7.batches_myo.csv
Untracked: data/covar_corrected_ici-myo-yes.vs.ici-myo-no_7.batches_myo_mis.csv
Untracked: data/covar_corrected_ici-sick.vs.ici-eoi_7.batches_myo.csv
Untracked: data/covar_corrected_ici-sick.vs.ici-eoi_7.batches_myo_mis.csv
Untracked: data/covar_corrected_ici.vs.pbs_7.batches_myo.csv
Untracked: data/covar_corrected_ici.vs.pbs_7.batches_myo_mis.csv
Untracked: data/covar_corrected_myo-yes.vs.myo-no_7.batches_myo.csv
Untracked: data/covar_corrected_myo-yes.vs.myo-no_7.batches_myo_mis.csv
Untracked: data/covar_corrected_pbs-myo-yes.vs.pbs-myo-no_7.batches_myo.csv
Untracked: data/covar_corrected_pbs-myo-yes.vs.pbs-myo-no_7.batches_myo_mis.csv
Untracked: data/e_4.batches_myo.RData
Untracked: data/e_7.batches_myo.RData
Untracked: data/e_snpg_samqc_4.batches_myo.RData
Untracked: data/e_snpg_samqc_7.batches_myo.RData
Untracked: data/errors_ind_4.batches_myo.RData
Untracked: data/errors_ind_7.batches_myo.RData
Untracked: data/genetic_map_4.batches_myo.csv
Untracked: data/genetic_map_7.batches_myo.csv
Untracked: data/genotype_errors_marker_4.batches_myo.RData
Untracked: data/genotype_errors_marker_7.batches_myo.RData
Untracked: data/genotype_freq_marker_4.batches_myo.RData
Untracked: data/genotype_freq_marker_7.batches_myo.RData
Untracked: data/gm_allqc_4.batches_myo.RData
Untracked: data/gm_allqc_4.batches_myo_mis.RData
Untracked: data/gm_allqc_7.batches_myo.RData
Untracked: data/gm_allqc_7.batches_myo_mis.RData
Untracked: data/gm_samqc_4.batches_myo.RData
Untracked: data/gm_samqc_7.batches_myo.RData
Untracked: data/gm_serreze.BC312.RData
Untracked: data/gm_serreze.BC366.RData
Untracked: data/gm_serreze_7.batches_myo.RData
Untracked: data/gm_serreze_bc_7.batches_myo.RData
Untracked: data/het-ici-myo-yes.vs.het-ici-myo-no_marker.freq_low.geno.freq.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/het-ici-myo-yes.vs.het-ici-myo-no_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/het-ici-myo-yes.vs.het-ici-myo-no_marker.freq_low.probs.freq.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/het-ici-myo-yes.vs.het-ici-myo-no_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/het-ici-myo-yes.vs.het-ici-myo-no_sample.genos_marker.freq_low.geno.freq.removed_7.batches_myo_mis.csv
Untracked: data/het-ici-myo-yes.vs.het-ici-myo-no_sample.genos_marker.freq_low.geno.freq.removed_sample.outliers.removed_7.batches_myo_mis.csv
Untracked: data/het-ici-myo-yes.vs.het-ici-myo-no_sample.genos_marker.freq_low.probs.freq.removed_7.batches_myo_mis.csv
Untracked: data/het-ici-myo-yes.vs.het-ici-myo-no_sample.genos_marker.freq_low.probs.freq.removed_sample.outliers.removed_7.batches_myo_mis.csv
Untracked: data/het-ici.vs.het-pbs_marker.freq_low.geno.freq.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/het-ici.vs.het-pbs_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/het-ici.vs.het-pbs_marker.freq_low.probs.freq.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/het-ici.vs.het-pbs_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/het-ici.vs.het-pbs_sample.genos_marker.freq_low.geno.freq.removed_7.batches_myo_mis.csv
Untracked: data/het-ici.vs.het-pbs_sample.genos_marker.freq_low.geno.freq.removed_sample.outliers.removed_7.batches_myo_mis.csv
Untracked: data/het-ici.vs.het-pbs_sample.genos_marker.freq_low.probs.freq.removed_7.batches_myo_mis.csv
Untracked: data/het-ici.vs.het-pbs_sample.genos_marker.freq_low.probs.freq.removed_sample.outliers.removed_7.batches_myo_mis.csv
Untracked: data/ici-myo-yes.vs.ici-myo-no_marker.freq_low.geno.freq.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/ici-myo-yes.vs.ici-myo-no_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/ici-myo-yes.vs.ici-myo-no_marker.freq_low.probs.freq.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/ici-myo-yes.vs.ici-myo-no_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/ici-myo-yes.vs.ici-myo-no_sample.genos_marker.freq_low.geno.freq.removed_7.batches_myo_mis.csv
Untracked: data/ici-myo-yes.vs.ici-myo-no_sample.genos_marker.freq_low.geno.freq.removed_sample.outliers.removed_7.batches_myo_mis.csv
Untracked: data/ici-myo-yes.vs.ici-myo-no_sample.genos_marker.freq_low.probs.freq.removed_7.batches_myo_mis.csv
Untracked: data/ici-myo-yes.vs.ici-myo-no_sample.genos_marker.freq_low.probs.freq.removed_sample.outliers.removed_7.batches_myo_mis.csv
Untracked: data/myo-yes.vs.myo-no_marker.freq_low.geno.freq.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/myo-yes.vs.myo-no_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/myo-yes.vs.myo-no_marker.freq_low.probs.freq.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/myo-yes.vs.myo-no_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/myo-yes.vs.myo-no_sample.genos_marker.freq_low.geno.freq.removed_7.batches_myo_mis.csv
Untracked: data/myo-yes.vs.myo-no_sample.genos_marker.freq_low.geno.freq.removed_sample.outliers.removed_7.batches_myo_mis.csv
Untracked: data/myo-yes.vs.myo-no_sample.genos_marker.freq_low.probs.freq.removed_7.batches_myo_mis.csv
Untracked: data/myo-yes.vs.myo-no_sample.genos_marker.freq_low.probs.freq.removed_sample.outliers.removed_7.batches_myo_mis.csv
Untracked: data/pbs-myo-yes.vs.pbs-myo-no_marker.freq_low.geno.freq.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/pbs-myo-yes.vs.pbs-myo-no_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/pbs-myo-yes.vs.pbs-myo-no_marker.freq_low.probs.freq.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/pbs-myo-yes.vs.pbs-myo-no_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_7.batches_myo_mis.csv
Untracked: data/pbs-myo-yes.vs.pbs-myo-no_sample.genos_marker.freq_low.geno.freq.removed_7.batches_myo_mis.csv
Untracked: data/pbs-myo-yes.vs.pbs-myo-no_sample.genos_marker.freq_low.geno.freq.removed_sample.outliers.removed_7.batches_myo_mis.csv
Untracked: data/pbs-myo-yes.vs.pbs-myo-no_sample.genos_marker.freq_low.probs.freq.removed_7.batches_myo_mis.csv
Untracked: data/pbs-myo-yes.vs.pbs-myo-no_sample.genos_marker.freq_low.probs.freq.removed_sample.outliers.removed_7.batches_myo_mis.csv
Untracked: data/percent_missing_id_4.batches_myo.RData
Untracked: data/percent_missing_id_7.batches_myo.RData
Untracked: data/percent_missing_marker_4.batches_myo.RData
Untracked: data/percent_missing_marker_7.batches_myo.RData
Untracked: data/pheno_4.batches_myo.csv
Untracked: data/pheno_7.batches_myo.csv
Untracked: data/physical_map_4.batches_myo.csv
Untracked: data/physical_map_7.batches_myo.csv
Untracked: data/probs_36state_bc_7.batches_myo.rds
Untracked: data/probs_8state_bc_7.batches_myo.rds
Untracked: data/qc_info_bad_sample_4.batches_myo.RData
Untracked: data/qc_info_bad_sample_7.batches_myo.RData
Untracked: data/sample_geno_AHB_4.batches_myo.csv
Untracked: data/sample_geno_AHB_7.batches_myo.csv
Untracked: data/sample_geno_AHB_7.batches_myo_orig.id.csv
Untracked: data/sample_geno_bc_4.batches_myo.csv
Untracked: data/sample_geno_bc_7.batches_myo.csv
Untracked: data/sample_geno_bc_7.batches_myo_orig.id.csv
Untracked: data/serreze_probs_4.batches_myo.rds
Untracked: data/serreze_probs_7.batches_myo.rds
Untracked: data/serreze_probs_allqc_4.batches_myo.rds
Untracked: data/serreze_probs_allqc_4.batches_myo_mis.rds
Untracked: data/serreze_probs_allqc_7.batches_myo.rds
Untracked: data/serreze_probs_allqc_7.batches_myo_mis.rds
Untracked: data/summary.cg_4.batches_myo.RData
Untracked: data/summary.cg_7.batches_myo.RData
Untracked: output/Percent_missing_genotype_data_4.batches_myo.pdf
Untracked: output/Percent_missing_genotype_data_7.batches_myo.pdf
Untracked: output/Percent_missing_genotype_data_per_marker_4.batches_myo.pdf
Untracked: output/Percent_missing_genotype_data_per_marker_7.batches_myo.pdf
Untracked: output/Proportion_matching_genotypes_before_removal_of_bad_samples_4.batches_myo.pdf
Untracked: output/Proportion_matching_genotypes_before_removal_of_bad_samples_7.batches_myo.pdf
Untracked: output/genotype_error_marker_4.batches_myo.pdf
Untracked: output/genotype_error_marker_7.batches_myo.pdf
Untracked: output/genotype_frequency_marker_4.batches_myo.pdf
Untracked: output/genotype_frequency_marker_7.batches_myo.pdf
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
There are no past versions. Publish this analysis with
wflow_publish() to start tracking its development.
Loading Project
load("data/e_snpg_samqc_7.batches_myo.RData")
gm <- get(load("data/gm_samqc_7.batches_myo.RData"))
gmWarning in check_cross2(object): 1249 invalid genotypes in crossObject of class cross2 (crosstype "bc")
Total individuals 357
No. genotyped individuals 357
No. phenotyped individuals 357
No. with both geno & pheno 357
No. phenotypes 1
No. covariates 11
No. phenotype covariates 0
No. chromosomes 20
Total markers 133716
No. markers by chr:
1 2 3 4 5 6 7 8 9 10 11 12 13
10159 10172 7987 7736 7778 7911 7548 6561 6823 6471 7276 6226 6177
14 15 16 17 18 19 X
6082 5421 5075 5162 4682 3612 4857 It can also be useful to look at the proportion of missing genotypes by marker. Markers with a lot of missing data were likely difficult to call, and so the genotypes that were called may contain a lot of errors.
Marker Missing Data
pmis_mar <- n_missing(gm, "marker", "proportion")*100
save(pmis_mar, file = "data/percent_missing_marker_7.batches_myo.RData")
par(mar=c(5.1,0.6,0.6, 0.6))
hist(pmis_mar, breaks=seq(0, 100, length=201),
main="", yaxt="n", ylab="", xlab="Percent missing genotypes")
rug(pmis_mar)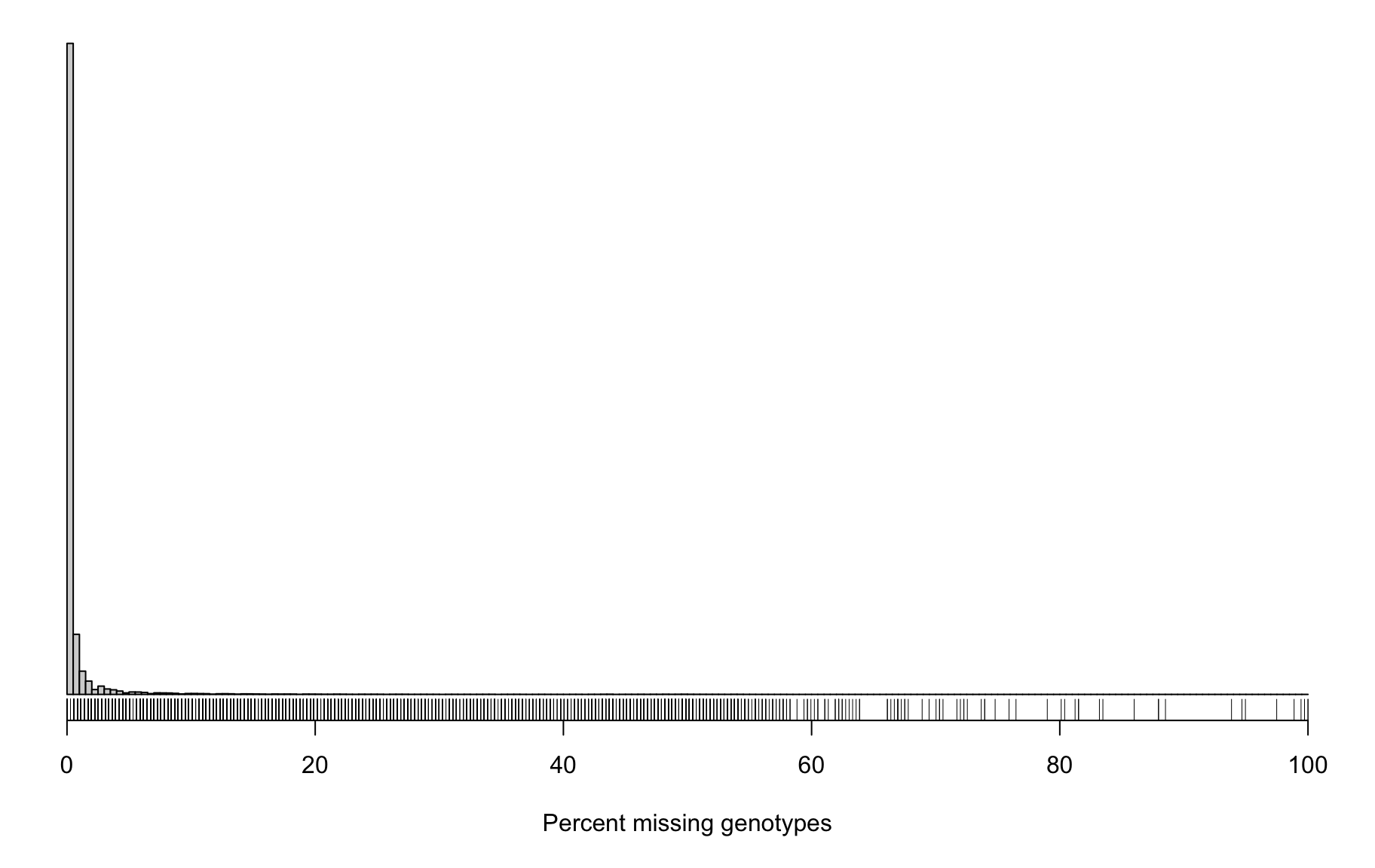
pdf(file = "output/Percent_missing_genotype_data_per_marker_7.batches_myo.pdf")
par(mar=c(5.1,0.6,0.6, 0.6))
hist(pmis_mar, breaks=seq(0, 100, length=201),
main="", yaxt="n", ylab="", xlab="Percent missing genotypes")
rug(pmis_mar)
dev.off()quartz_off_screen
2 | count | |
|---|---|
| pmis_mar_5 | 5204 |
| pmis_mar_10 | 2826 |
| pmis_mar_15 | 1851 |
| pmis_mar_25 | 1043 |
| pmis_mar_50 | 257 |
| pmis_mar_75 | 23 |
| total_snps | 133716 |
Marker Genotype Frequencies
g <- do.call("cbind", gm$geno[1:19])
#fg <- do.call("cbind", gm$founder_geno[1:19])
#g <- g[,colSums(g)!=0]
#fg <- fg[,colSums(fg==0)==0]
#fgn <- colSums(g==2)
gf_mar <- t(apply(g, 2, function(a) table(factor(a, 1:2))/sum(a != 0)))
gn_mar <- t(apply(g, 2, function(a) table(factor(a, 1:2))))
gf_mar <- gf_mar[gf_mar[,2] != "NaN",]
MAF <- apply(gf_mar, 1, function(x) min(x))
MAF <- as.data.frame(MAF)
MAF$index <- 1:nrow(gf_mar)
gf_mar_maf <- merge(gf_mar,as.data.frame(MAF), by="row.names")
gf_mar_maf <- gf_mar_maf[order(gf_mar_maf$index),]
pdf(file = "output/genotype_frequency_marker_7.batches_myo.pdf")
par(mar=c(5.1,0.6,0.6, 0.6))
hist(gf_mar_maf$MAF, breaks=seq(0, 1, length=201),
main="", yaxt="n", ylab="", xlab="MAF")
rug(gf_mar_maf$MAF)
dev.off()quartz_off_screen
2 par(mar=c(5.1,0.6,0.6, 0.6))
hist(gf_mar_maf$MAF, breaks=seq(0, 1, length=201),
main="", yaxt="n", ylab="", xlab="MAF")
rug(gf_mar_maf$MAF)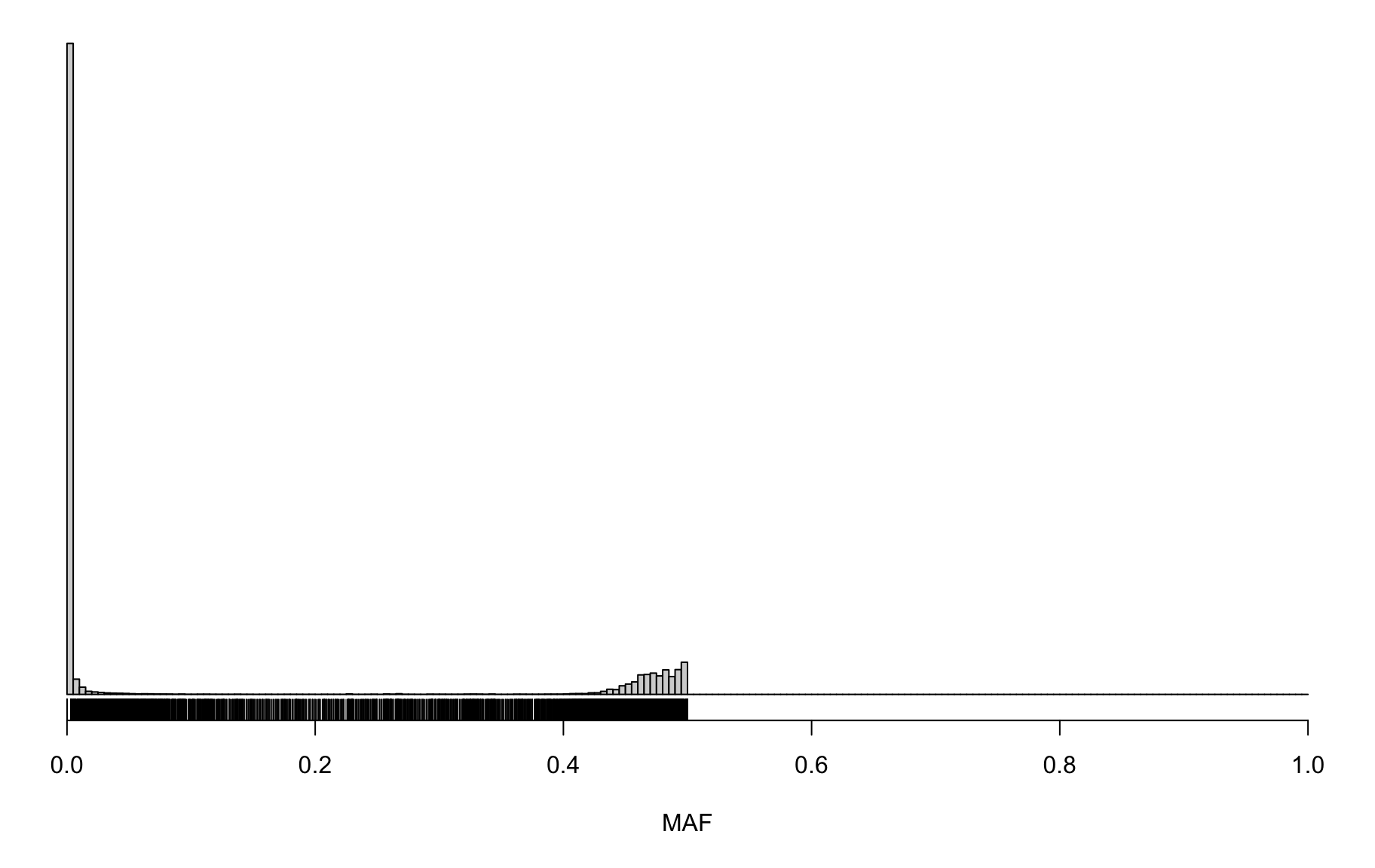
gfmar <- NULL
gfmar$gfmar_mar_0 <- sum(gf_mar_maf$MAF==0)
gfmar$gfmar_mar_1 <- sum(gf_mar_maf$MAF< 0.01)
gfmar$gfmar_mar_5 <- sum(gf_mar_maf$MAF< 0.05)
gfmar$gfmar_mar_10 <- sum(gf_mar_maf$MAF< 0.10)
gfmar$gfmar_mar_15 <- sum(gf_mar_maf$MAF< 0.15)
gfmar$gfmar_mar_25 <- sum(gf_mar_maf$MAF< 0.25)
gfmar$gfmar_mar_50 <- sum(gf_mar_maf$MAF<= 0.50)
gfmar$total_snps <- nrow(as.data.frame(gf_mar_maf))
gfmar <- t(as.data.frame(gfmar))
gfmar <- as.data.frame(gfmar)
gfmar$count <- gfmar$V1
gfmar[c(2)] %>%
kable(escape = F,align = c("ccccccccc"),linesep ="\\hline") %>%
kable_styling(full_width = F) %>%
kable_styling("striped", full_width = F) %>%
row_spec(8 ,bold=T,color= "white",background = "black")| count | |
|---|---|
| gfmar_mar_0 | 87114 |
| gfmar_mar_1 | 92158 |
| gfmar_mar_5 | 94870 |
| gfmar_mar_10 | 95480 |
| gfmar_mar_15 | 95744 |
| gfmar_mar_25 | 96154 |
| gfmar_mar_50 | 128857 |
| total_snps | 128857 |
save(gf_mar, file = "data/genotype_freq_marker_7.batches_myo.RData")Marker Genotype Errors
errors_mar <- colSums(e>2)/n_typed(gm, "marker")*100
grayplot(pmis_mar, errors_mar,
xlab="Proportion missing", ylab="Proportion genotyping errors")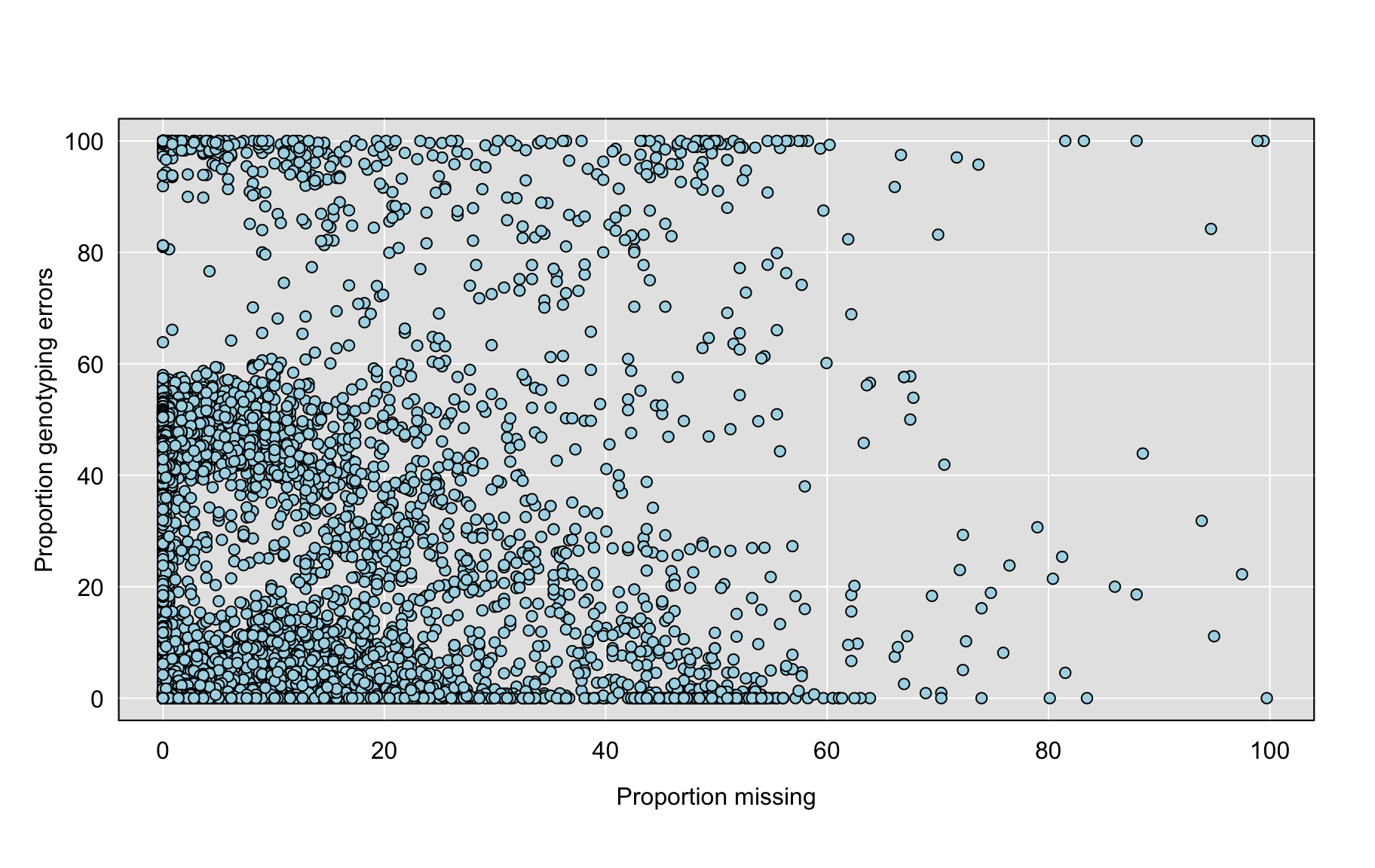
pdf(file = "output/genotype_error_marker_7.batches_myo.pdf")
grayplot(pmis_mar, errors_mar,
xlab="Proportion missing", ylab="Proportion genotyping errors")
dev.off()quartz_off_screen
2 save(errors_mar, file = "data/genotype_errors_marker_7.batches_myo.RData")Markers with higher rates of missing genotypes tend to show higher errors rates.
Removing Markers
Missingness
#Missingness
length(pmis_mar[pmis_mar >= 10])[1] 2826high_miss <- find_markerpos(gm, names(pmis_mar[pmis_mar >= 10]))
high_miss$id <- rownames(high_miss)
high_miss_df <- as.data.frame(pmis_mar[pmis_mar >= 10])
high_miss_df$index = 1: nrow(high_miss_df)
high_miss_df$id <- rownames(high_miss_df)
high_miss_bad <- merge(high_miss,high_miss_df, by=c("id"),all=T)
names(high_miss_bad)[5] <- c("high_miss")
names(high_miss_bad)[1] <- c("marker")
high_miss_bad <- high_miss_bad[order(high_miss_bad$index),]Monomorphic/Low Frequency markers
#Monomorphic/Low Frequency markers
#low_freq_df <- as.data.frame(gf_mar)
count <- rowSums(gf_mar <= 0.01)
#count <- as.data.frame(count)
low_freq_df <- merge(as.data.frame(gf_mar),as.data.frame(count), by="row.names",all=T)
low_freq_df[is.na(low_freq_df)] <- ''
low_freq_df <- low_freq_df[low_freq_df$count == 1,]
rownames(low_freq_df) <- low_freq_df$Row.names
#low_freq_df$id <- rownames(low_freq_df)
#low_freq_df$index = 1: nrow(low_freq_df)
low_freq <- find_markerpos(gm, rownames(low_freq_df))
low_freq$id <- rownames(low_freq)
nrow(low_freq)[1] 92159low_freq_bad <- merge(low_freq,low_freq_df, by="row.names",all=T)
#names(low_freq_bad)[5] <- c("AA_freq")
#names(low_freq_bad)[6] <- c("AB_freq")
#names(low_freq_bad)[7] <- c("BB_freq")
names(low_freq_bad)[1] <- c("marker")
#low_freq_bad <- low_freq_bad[order(low_freq_bad$index),]Genotyping Error
##Genotyping Error
length(errors_mar[errors_mar > 5])[1] 11469error_markers_names <- names(errors_mar[errors_mar > 5])
error_markers_names <- error_markers_names[complete.cases(error_markers_names)]
error_markers <- find_markerpos(gm, error_markers_names)
error_markers$id <- rownames(error_markers)
#rne <- rownames(as.data.frame(errors_mar))
error_mars_df <- as.data.frame(errors_mar[errors_mar > 5])
error_mars_df <- error_mars_df[complete.cases(error_mars_df$"errors_mar[errors_mar > 5]"),]
error_mars_df <- as.data.frame(error_mars_df)
#error_mars_df$id = rownames(error_mars_df)
error_mars_df$index = 1: nrow(error_mars_df)
#error_markers_bad <- merge(error_markers,error_mars_df, by=c("id"),all=T)
error_markers_bad <- cbind(error_markers,error_mars_df)
names(error_markers_bad)[5] <- c("error_mars")
names(error_markers_bad)[4] <- c("marker")
error_markers_bad <- error_markers_bad[order(error_markers_bad$index),]Total
### merge all
bad_markers <- rbind(high_miss_bad[c("marker","chr","gmap","pmap")], low_freq_bad[c("marker","chr","gmap","pmap")], error_markers_bad[c("marker","chr","gmap","pmap")])
#nrow(bad_markers)
duplicate <- bad_markers[duplicated(bad_markers),]
bad_markers <- bad_markers[!duplicated(bad_markers),]
nrow(bad_markers)[1] 101056save(bad_markers, file = "data/bad_markers_all_7.batches_myo.RData")Only removing markers that are missing in at least 10% of the samples as well those that come as genotyping errors and have a allele frequency of less than 1%
#missing in at least 10% of the samples
gm_allqc2 <- drop_markers(gm_samqc, bad_markers$marker)
gm_allqc <- drop_nullmarkers(gm_allqc2)
gm_allqcWarning in check_cross2(object): 461 invalid genotypes in crossObject of class cross2 (crosstype "bc")
Total individuals 357
No. genotyped individuals 357
No. phenotyped individuals 357
No. with both geno & pheno 357
No. phenotypes 1
No. covariates 11
No. phenotype covariates 0
No. chromosomes 20
Total markers 32660
No. markers by chr:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16
2482 2412 1742 1772 1648 1842 1541 1517 1769 1092 1754 1230 1441 1505 1128 842
17 18 19 X
655 820 945 4523 save(gm_allqc, file = "data/gm_allqc_7.batches_myo.RData")
sessionInfo()R version 4.2.2 (2022-10-31)
Platform: x86_64-apple-darwin17.0 (64-bit)
Running under: macOS Big Sur ... 10.16
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] fst_0.9.8 knitr_1.41 kableExtra_1.3.4 mclust_6.0.0
[5] ggrepel_0.9.2 ggplot2_3.4.0 qtlcharts_0.16 qtl2_0.30
[9] broman_0.80 workflowr_1.7.0
loaded via a namespace (and not attached):
[1] Rcpp_1.0.9 svglite_2.1.0 getPass_0.2-2 ps_1.7.2
[5] assertthat_0.2.1 rprojroot_2.0.3 digest_0.6.30 utf8_1.2.2
[9] R6_2.5.1 RSQLite_2.2.19 evaluate_0.18 httr_1.4.4
[13] highr_0.9 pillar_1.8.1 rlang_1.0.6 rstudioapi_0.14
[17] data.table_1.14.6 whisker_0.4.1 callr_3.7.3 jquerylib_0.1.4
[21] blob_1.2.3 fstcore_0.9.12 rmarkdown_2.18 webshot_0.5.4
[25] qtl_1.54 stringr_1.5.0 bit_4.0.5 munsell_0.5.0
[29] compiler_4.2.2 httpuv_1.6.6 xfun_0.35 systemfonts_1.0.4
[33] pkgconfig_2.0.3 htmltools_0.5.3 tidyselect_1.2.0 tibble_3.1.8
[37] viridisLite_0.4.1 fansi_1.0.3 dplyr_1.0.10 withr_2.5.0
[41] later_1.3.0 grid_4.2.2 jsonlite_1.8.4 gtable_0.3.1
[45] lifecycle_1.0.3 DBI_1.1.3 git2r_0.30.1 magrittr_2.0.3
[49] scales_1.2.1 cli_3.4.1 stringi_1.7.8 cachem_1.0.6
[53] fs_1.5.2 promises_1.2.0.1 xml2_1.3.3 bslib_0.4.1
[57] vctrs_0.5.1 generics_0.1.3 tools_4.2.2 bit64_4.0.5
[61] glue_1.6.2 processx_3.8.0 parallel_4.2.2 fastmap_1.1.0
[65] yaml_2.3.6 colorspace_2.0-3 rvest_1.0.3 memoise_2.0.1
[69] sass_0.4.4