QTL Analysis - rz transformed age of onset (rz.age) [ICI vs EOI] (corrected phenotype with outliers removed if any)
Belinda Cornes
2022-04-14
Last updated: 2022-04-14
Checks: 5 2
Knit directory: Serreze-T1D_Workflow/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown is untracked by Git. To know which version of the R Markdown file created these results, you’ll want to first commit it to the Git repo. If you’re still working on the analysis, you can ignore this warning. When you’re finished, you can run wflow_publish to commit the R Markdown file and build the HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20220210) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Using absolute paths to the files within your workflowr project makes it difficult for you and others to run your code on a different machine. Change the absolute path(s) below to the suggested relative path(s) to make your code more reproducible.
| absolute | relative |
|---|---|
| /Users/corneb/Documents/MyJax/CS/Projects/Serreze/qc/workflowr/Serreze-T1D_Workflow | . |
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version a8c2d1a. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .DS_Store
Ignored: analysis/.DS_Store
Ignored: data/.DS_Store
Untracked files:
Untracked: analysis/3.1_phenotype.qc_corrected_5.batches_0.Rmd
Untracked: analysis/3.1_phenotype.qc_corrected_5.batches_52.Rmd
Untracked: analysis/3.1_phenotype.qc_corrected_5.batches_mis_0.Rmd
Untracked: analysis/3.1_phenotype.qc_corrected_5.batches_mis_52.Rmd
Untracked: analysis/3.1_phenotype.qc_corrected_5.batches_mis_vo.Rmd
Untracked: analysis/3.1_phenotype.qc_corrected_5.batches_vo.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici.vs.eoi_snpsqc_dis_no-x_updated_5.batches_11.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici.vs.eoi_snpsqc_dis_no-x_updated_5.batches_mis_11.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici.vs.pbs_snpsqc_5.batches_1.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici.vs.pbs_snpsqc_5.batches_mis_1.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici.vs.pbs_snpsqc_dis_no-x_updated_5.batches_1.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici.vs.pbs_snpsqc_dis_no-x_updated_5.batches_11.Rmd
Untracked: analysis/4.1.1_qtl.analysis_binary_ici.vs.pbs_snpsqc_dis_no-x_updated_5.batches_mis_11.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_changed.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_mis_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_mis_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_mis_changed.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_pheno.corrected.cleaned_scanone_5.batches.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_pheno.corrected.cleaned_scanone_5.batches_mis.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_mis_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_mis_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_oops.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis.Rmd.R
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_mis_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_mis_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_test.with.4.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_test.with.4.Rmd.R
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_test.with.4_miss.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_test.with.4_miss.Rmd.R
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_test.with.5.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_test.with.5.Rmd.R
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_test.with.5_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_test.with.5_0.Rmd.R
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_mis_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_mis_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_mis_vo.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_mis_vo1.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_vo.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_vo1.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_mis_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_mis_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_52.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_mis_0.Rmd
Untracked: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_mis_52.Rmd
Untracked: analysis/genotype.frequencies_ici.vs.eoi_5.batches_0.Rmd
Untracked: analysis/genotype.frequencies_ici.vs.eoi_5.batches_52.Rmd
Untracked: analysis/genotype.frequencies_ici.vs.eoi_5.batches_mis_0.Rmd
Untracked: analysis/genotype.frequencies_ici.vs.eoi_5.batches_mis_52.Rmd
Untracked: analysis/index_5.batches_additional_vo.Rmd
Untracked: data/GM_covar.csv
Untracked: data/GM_covar_BC312.csv
Untracked: data/bad_markers_all_4.batches.RData
Untracked: data/bad_markers_all_5.batches.RData
Untracked: data/blup_sub_chr10_lod.drop-1.5_5.batches.csv
Untracked: data/blup_sub_chr3_lod.drop-1.5.csv
Untracked: data/blup_sub_chr3_lod.drop-1.5_5.batches.csv
Untracked: data/blup_sub_chr4_lod.drop-1.5.csv
Untracked: data/blup_sub_chr4_lod.drop-1.5_5.batches.csv
Untracked: data/covar_cleaned_ici.vs.eoi.csv
Untracked: data/covar_cleaned_ici.vs.pbs.csv
Untracked: data/covar_corrected.cleaned_ici-early.vs.pbs_5.batches.csv
Untracked: data/covar_corrected.cleaned_ici-early.vs.pbs_5.batches_0.csv
Untracked: data/covar_corrected.cleaned_ici-early.vs.pbs_5.batches_52.csv
Untracked: data/covar_corrected.cleaned_ici-early.vs.pbs_5.batches_mis.csv
Untracked: data/covar_corrected.cleaned_ici-early.vs.pbs_5.batches_mis_0.csv
Untracked: data/covar_corrected.cleaned_ici-early.vs.pbs_5.batches_mis_52.csv
Untracked: data/covar_corrected.cleaned_ici.vs.eoi.csv
Untracked: data/covar_corrected.cleaned_ici.vs.eoi1.csv
Untracked: data/covar_corrected.cleaned_ici.vs.eoi_5.batches.csv
Untracked: data/covar_corrected.cleaned_ici.vs.eoi_5.batches_0.csv
Untracked: data/covar_corrected.cleaned_ici.vs.eoi_5.batches_52.csv
Untracked: data/covar_corrected.cleaned_ici.vs.eoi_5.batches_mis.csv
Untracked: data/covar_corrected.cleaned_ici.vs.eoi_5.batches_mis_0.csv
Untracked: data/covar_corrected.cleaned_ici.vs.eoi_5.batches_mis_52.csv
Untracked: data/covar_corrected.cleaned_ici.vs.pbs.csv
Untracked: data/covar_corrected.cleaned_ici.vs.pbs1.csv
Untracked: data/covar_corrected.cleaned_ici.vs.pbs_5.batches.csv
Untracked: data/covar_corrected.cleaned_ici.vs.pbs_5.batches_0.csv
Untracked: data/covar_corrected.cleaned_ici.vs.pbs_5.batches_52.csv
Untracked: data/covar_corrected.cleaned_ici.vs.pbs_5.batches_mis.csv
Untracked: data/covar_corrected.cleaned_ici.vs.pbs_5.batches_mis_0.csv
Untracked: data/covar_corrected.cleaned_ici.vs.pbs_5.batches_mis_52.csv
Untracked: data/covar_corrected_ici-early.vs.pbs_5.batches.csv
Untracked: data/covar_corrected_ici-early.vs.pbs_5.batches_0.csv
Untracked: data/covar_corrected_ici-early.vs.pbs_5.batches_52.csv
Untracked: data/covar_corrected_ici-early.vs.pbs_5.batches_mis.csv
Untracked: data/covar_corrected_ici-early.vs.pbs_5.batches_mis_0.csv
Untracked: data/covar_corrected_ici-early.vs.pbs_5.batches_mis_52.csv
Untracked: data/covar_corrected_ici.vs.eoi.csv
Untracked: data/covar_corrected_ici.vs.eoi1.csv
Untracked: data/covar_corrected_ici.vs.eoi_5.batches.csv
Untracked: data/covar_corrected_ici.vs.eoi_5.batches_0.csv
Untracked: data/covar_corrected_ici.vs.eoi_5.batches_52.csv
Untracked: data/covar_corrected_ici.vs.eoi_5.batches_mis.csv
Untracked: data/covar_corrected_ici.vs.eoi_5.batches_mis_0.csv
Untracked: data/covar_corrected_ici.vs.eoi_5.batches_mis_52.csv
Untracked: data/covar_corrected_ici.vs.pbs.csv
Untracked: data/covar_corrected_ici.vs.pbs1.csv
Untracked: data/covar_corrected_ici.vs.pbs_5.batches.csv
Untracked: data/covar_corrected_ici.vs.pbs_5.batches_0.csv
Untracked: data/covar_corrected_ici.vs.pbs_5.batches_52.csv
Untracked: data/covar_corrected_ici.vs.pbs_5.batches_mis.csv
Untracked: data/covar_corrected_ici.vs.pbs_5.batches_mis_0.csv
Untracked: data/covar_corrected_ici.vs.pbs_5.batches_mis_52.csv
Untracked: data/e.RData
Untracked: data/e_BC312.RData
Untracked: data/e_snpg_samqc_4.batches.RData
Untracked: data/e_snpg_samqc_4.batches_bc.RData
Untracked: data/e_snpg_samqc_5.batches.RData
Untracked: data/errors_ind_4.batches.RData
Untracked: data/errors_ind_4.batches_bc.RData
Untracked: data/errors_ind_5.batches.RData
Untracked: data/files.to.sync.txt
Untracked: data/fitqtl_chr3.peak_chr4.peak_additive.txt
Untracked: data/fitqtl_chr3.peak_chr4.peak_interacting.txt
Untracked: data/fitqtl_chr3.peak_chr4.peak_sex_additive.txt
Untracked: data/fitqtl_chr3.peak_chr4.peak_sex_interacting.txt
Untracked: data/g2blup_effects.csv
Untracked: data/g2blup_effects.xlsx
Untracked: data/genes_chr10_lod.drop-1.5_5.batches.csv
Untracked: data/genes_chr3_lod.drop-1.5.csv
Untracked: data/genes_chr3_lod.drop-1.5_5.batches.csv
Untracked: data/genes_chr4_lod.drop-1.5.csv
Untracked: data/genetic_map.csv
Untracked: data/genetic_map_BC312.csv
Untracked: data/genotype_errors_marker_4.batches.RData
Untracked: data/genotype_errors_marker_5.batches.RData
Untracked: data/genotype_freq_marker_4.batches.RData
Untracked: data/genotype_freq_marker_5.batches.RData
Untracked: data/gm_allqc_4.batches.RData
Untracked: data/gm_allqc_5.batches.RData
Untracked: data/gm_allqc_5.batches_mis.RData
Untracked: data/gm_samqc_3.batches.RData
Untracked: data/gm_samqc_4.batches.RData
Untracked: data/gm_samqc_4.batches_bc.RData
Untracked: data/gm_samqc_5.batches.RData
Untracked: data/gm_serreze.192.RData
Untracked: data/gm_serreze.BC312.RData
Untracked: data/ici-early.vs.pbs_age.of.onset-additive.covariates_blup_sub_chr-7_peak.marker-UNC13388811_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-additive.covariates_genes_chr-7_peak.marker-UNC13388811_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-17_peak.marker-UNCJPD006614_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-no.covariates_genes_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-no.covariates_genes_chr-17_peak.marker-UNCJPD006614_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-no.covariates_genes_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-no.covariates_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_age.of.onset-no.covariates_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-10_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-10_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-10_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-10_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-10_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-10_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-11_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-11_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-11_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-11_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-11_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-11_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-12_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-12_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-12_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-12_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-12_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-12_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-13_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-13_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-13_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-13_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-13_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-13_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-14_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-14_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-14_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-14_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-14_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-14_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-15_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-15_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-15_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-15_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-15_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-15_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-16_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-16_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-16_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-16_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-16_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-16_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-17_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-17_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-17_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-17_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-17_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-17_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-18_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-18_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-18_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-18_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-18_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-18_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-19_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-19_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-19_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-19_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-19_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-19_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-1_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-1_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-1_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-1_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-1_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-1_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-2_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-2_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-2_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-2_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-2_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-2_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-3_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-3_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-3_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-3_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-3_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-3_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-4_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-4_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-4_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-4_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-4_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-4_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-5_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-5_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-5_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-6_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-6_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-6_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-6_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-6_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-6_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-7_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-7_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-7_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-7_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-7_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-7_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-8_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-8_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-8_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-8_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-8_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-8_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-9_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-9_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-9_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-9_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-9_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-9_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-X_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-X_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-X_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-X_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-X_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.full_chr-X_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-10_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-10_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-10_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-10_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-10_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-10_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-11_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-11_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-11_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-11_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-11_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-11_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-12_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-12_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-12_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-12_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-12_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-12_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-13_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-13_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-13_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-13_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-13_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-13_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-14_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-14_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-14_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-14_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-14_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-14_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-15_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-15_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-15_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-15_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-16_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-16_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-16_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-16_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-16_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-16_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-17_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-17_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-17_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-17_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-18_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-18_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-18_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-18_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-18_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-18_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-19_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-19_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-19_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-19_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-19_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-19_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-1_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-1_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-1_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-1_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-1_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-1_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-2_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-2_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-2_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-2_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-2_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-2_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-3_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-3_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-3_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-3_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-3_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-3_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-4_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-4_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-4_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-4_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-4_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-4_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-5_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-5_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-5_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-6_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-6_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-6_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-6_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-6_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-6_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-7_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-7_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-7_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-7_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-7_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-7_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-8_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-8_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-8_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-8_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-8_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-8_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-9_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-9_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-9_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-9_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-9_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-9_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-X_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-X_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-X_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-X_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-X_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_blup.qc_chr-X_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_blup_sub_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_genes_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_gm_qtl_5.batches.csv
Untracked: data/ici-early.vs.pbs_gm_qtl_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_gm_qtl_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_gm_qtl_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_gm_qtl_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_gm_qtl_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_marker.freq_low.geno.freq.removed_geno.ratio_5.batches.csv
Untracked: data/ici-early.vs.pbs_marker.freq_low.geno.freq.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_5.batches.csv
Untracked: data/ici-early.vs.pbs_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_marker.freq_low.probs.freq.removed_geno.ratio_5.batches.csv
Untracked: data/ici-early.vs.pbs_marker.freq_low.probs.freq.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_5.batches.csv
Untracked: data/ici-early.vs.pbs_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_rz.age-additive.covariates_blup_sub_chr-7_peak.marker-UNC13239362_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_rz.age-additive.covariates_blup_sub_chr-7_peak.marker-UNC13239362_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_rz.age-additive.covariates_blup_sub_chr-7_peak.marker-UNC13239362_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_rz.age-additive.covariates_blup_sub_chr-7_peak.marker-UNC13239362_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_rz.age-additive.covariates_genes_chr-7_peak.marker-UNC13239362_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici-early.vs.pbs_rz.age-additive.covariates_genes_chr-7_peak.marker-UNC13239362_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_rz.age-additive.covariates_genes_chr-7_peak.marker-UNC13239362_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici-early.vs.pbs_rz.age-additive.covariates_genes_chr-7_peak.marker-UNC13239362_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici-early.vs.pbs_scanone_5.batches.Rdata
Untracked: data/ici-early.vs.pbs_scanone_5.batches_mis.Rdata
Untracked: data/ici-early.vs.pbs_scanone_snpsqc_5.batches.Rdata
Untracked: data/ici-early.vs.pbs_scanone_snpsqc_5.batches_mis.Rdata
Untracked: data/ici-early.vs.pbs_scanone_snpsqc_dis_no-x_updated_5.batches.Rdata
Untracked: data/ici-early.vs.pbs_scanone_snpsqc_dis_no-x_updated_5.batches_mis.Rdata
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-JAX00020646_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-JAX00292499_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-JAX00292927_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-JAX00292927_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-JAX00294019_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18216614_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18240977_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18311938_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18311938_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18311938_lod.drop-1.5_snpsqc_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18311938_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18343181_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18363544_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18363544_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18363544_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNC18376338_lod.drop-1.5_snpsqc_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNCHS028236_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNCHS028236_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNCHS028536_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-10_peak.marker-UNCHS028536_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-11_peak.marker-UNC19970181_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-11_peak.marker-UNC19970181_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-11_peak.marker-UNC20090524_lod.drop-1.5_snpsqc_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-13_peak.marker-UNCHS036773_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-13_peak.marker-UNCHS036773_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-15_peak.marker-UNC26070435_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-15_peak.marker-UNC26070435_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-17_peak.marker-UNCJPD006614_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-17_peak.marker-UNCrs47191360_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-2_peak.marker-UNC4609527_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-2_peak.marker-UNC4609527_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-2_peak.marker-UNCHS008007_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-2_peak.marker-UNCHS008007_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-ICR5263_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-ICR5263_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00105649_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00105649_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00106210_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00107193_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00220631_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00220631_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00514662_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00514662_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00516812_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00516812_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00517674_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00517674_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00518911r_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00518911r_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00522922_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00524205_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00529707r_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00529707r_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-JAX00535764_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4809586_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4809586_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4820379_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4820379_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4836550_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4836550_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4836550_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4875653_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4875653_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4901083_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4901083_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4909519_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4909519_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4912039_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4912039_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4912288_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4912288_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4915446_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4915446_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4923398_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4923398_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4938565_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4938565_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4945186_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4945186_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4947492_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4947492_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4956618_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4956618_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4957168_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4957859_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4957859_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4959266_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4959266_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4960612_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4960612_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4961542_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4961542_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4979914_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4979914_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4990111_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC4990111_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5046545_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5055259_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5062524_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5068215_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5076935_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5082757_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5173293_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5192582_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5192582_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5192582_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5193970_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5213888_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5213888_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5257268_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5262472_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5262472_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5262472_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5264696_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5267813_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5270225_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5277061_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5278229_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5278229_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5282259_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5287977_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5301460_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5309136_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5321559_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5322455_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5329171_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5360055_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5778977_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5808246_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNC5916997_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008182_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008182_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008182_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008193_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008193_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008209_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008209_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008215_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008215_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008219_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008219_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008229_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008229_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008234_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008234_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008242_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008260_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008260_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008271_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008271_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008281_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008281_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008338_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008338_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008363_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008363_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008379_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008379_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008409_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008432_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008511_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008511_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008609_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008613_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008613_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008614_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008627_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008659_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008725_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009066_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009067_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009068_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009112_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009224_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009224_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009383_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009383_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009693_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009821_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCJPD001170_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCJPD001170_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCJPD001198_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCJPD001198_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-UNCJPD001276_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-sanger2496q_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-3_peak.marker-sanger2496q_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-4_peak.marker-UNC8250659_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-4_peak.marker-UNC8439633_lod.drop-1.5_snpsqc_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-4_peak.marker-UNC8439633_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-4_peak.marker-UNCHS012955_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-6_peak.marker-UNC11108920_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-6_peak.marker-UNC11108920_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-9_peak.marker-UNC17203597_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-9_peak.marker-UNC17203597_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-JAX00020646_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-JAX00292499_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-JAX00292927_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-JAX00292927_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-JAX00294019_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18216614_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18240977_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18311938_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18311938_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18311938_lod.drop-1.5_snpsqc_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18311938_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18343181_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18363544_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18363544_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18363544_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNC18376338_lod.drop-1.5_snpsqc_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNCHS028236_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNCHS028236_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNCHS028536_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-10_peak.marker-UNCHS028536_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-11_peak.marker-UNC19970181_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-11_peak.marker-UNC19970181_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-11_peak.marker-UNC20090524_lod.drop-1.5_snpsqc_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-13_peak.marker-UNCHS036773_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-13_peak.marker-UNCHS036773_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-15_peak.marker-UNC26070435_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-15_peak.marker-UNC26070435_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-17_peak.marker-UNCJPD006614_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-17_peak.marker-UNCrs47191360_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-2_peak.marker-UNC4609527_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-2_peak.marker-UNC4609527_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-2_peak.marker-UNCHS008007_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-2_peak.marker-UNCHS008007_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-ICR5263_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-ICR5263_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00105649_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00105649_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00106210_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00107193_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00220631_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00220631_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00514662_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00514662_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00516812_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00516812_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00517674_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00517674_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00518911r_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00518911r_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00522922_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00524205_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00529707r_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00529707r_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-JAX00535764_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4809586_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4809586_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4820379_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4820379_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4836550_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4836550_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4836550_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4875653_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4875653_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4901083_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4901083_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4909519_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4909519_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4912039_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4912039_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4912288_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4912288_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4915446_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4915446_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4923398_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4923398_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4938565_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4938565_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4945186_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4945186_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4947492_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4947492_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4956618_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4956618_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4957168_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4957859_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4957859_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4959266_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4959266_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4960612_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4960612_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4961542_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4961542_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4979914_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4979914_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4990111_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC4990111_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5046545_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5055259_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5062524_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5068215_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5076935_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5082757_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5173293_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5192582_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5192582_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5192582_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5193970_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5213888_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5213888_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5257268_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5262472_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5262472_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5262472_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5264696_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5267813_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5270225_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5277061_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5278229_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5278229_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5282259_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5287977_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5301460_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5309136_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5321559_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5322455_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5329171_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5360055_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5778977_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5808246_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNC5916997_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008182_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008182_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008182_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008193_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008193_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008209_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008209_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008215_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008215_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008219_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008219_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008229_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008229_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008234_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008234_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008242_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008260_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008260_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008271_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008271_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008281_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008281_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008338_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008338_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008363_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008363_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008379_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008379_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008409_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008432_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008511_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008511_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008609_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008613_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008613_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008614_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008627_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008659_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008725_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS009066_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS009067_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS009068_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS009112_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS009224_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS009224_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS009383_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS009383_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS009693_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCHS009821_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCJPD001170_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCJPD001170_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCJPD001198_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCJPD001198_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-UNCJPD001276_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-sanger2496q_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-3_peak.marker-sanger2496q_lod.drop-1.5_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-4_peak.marker-UNC8250659_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-4_peak.marker-UNC8439633_lod.drop-1.5_snpsqc_5.batches_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-4_peak.marker-UNC8439633_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-4_peak.marker-UNCHS012955_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-6_peak.marker-UNC11108920_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-6_peak.marker-UNC11108920_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-9_peak.marker-UNC17203597_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-9_peak.marker-UNC17203597_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_age.of.onset-no.covariates_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_blup.full_chr-10_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-10_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-11_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-11_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-12_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-12_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-13_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-13_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-14_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-14_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-15_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-15_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-16_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-16_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-17_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-17_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-18_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-18_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-19_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-19_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-1_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-1_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-2_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-2_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-3_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-3_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-4_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-4_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-6_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-6_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-7_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-7_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-8_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-8_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-9_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-9_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.full_chr-X_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.full_chr-X_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-10_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-10_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-10_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-10_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-10_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-10_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-10_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-11_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-11_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-11_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-11_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-11_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-11_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-11_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-12_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-12_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-12_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-12_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-12_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-12_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-12_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-13_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-13_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-13_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-13_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-13_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-13_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-13_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-14_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-14_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-14_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-14_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-14_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-14_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-15_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-15_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-15_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-15_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-15_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-16_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-16_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-16_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-16_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-16_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-16_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-17_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-17_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-17_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-17_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-17_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-17_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-18_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-18_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-18_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-18_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-18_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-18_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-19_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-19_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-19_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-19_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-19_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-19_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-1_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-1_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-1_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-1_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-1_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-1_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-1_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-2_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-2_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-2_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-2_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-2_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-2_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-2_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-3_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-3_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-3_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-3_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-3_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-3_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-3_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-4_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-4_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-4_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-4_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-4_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-4_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-4_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-5_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-5_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-6_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-6_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-6_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-6_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-6_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-6_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-6_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-7_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-7_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-7_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-7_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-7_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-7_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-7_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-8_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-8_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-8_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-8_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-8_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-8_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-8_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-9_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-9_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-9_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-9_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-9_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-9_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-9_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-X_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-X_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-X_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-X_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-X_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup.qc_chr-X_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-ICR4295_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-JAX00292499_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-JAX00292927_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-JAX00292927_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-JAX00292927_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-UNC18154330_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-UNC18195765_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-UNC18216614_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-UNC18240977_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-UNC18311938_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-UNC18856953_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-UNCHS028236_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-UNCHS028536_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-UNCJPD004316_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-10_peak.marker-UNCJPD009532_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-11_peak.marker-UNC19155926_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-11_peak.marker-UNC19481366_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-11_peak.marker-UNC19970181_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-11_peak.marker-UNC19970181_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-11_peak.marker-UNCJPD004792_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-12_peak.marker-ICR4497_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-13_peak.marker-UNCHS036773_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-13_peak.marker-UNCHS036964_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-14_peak.marker-UNC24056202_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-15_peak.marker-UNC26070435_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-17_peak.marker-UNCrs47191360_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-17_peak.marker-UNCrs47191360_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-18_peak.marker-UNCHS045344_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-19_peak.marker-UNCJPD007087_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-1_peak.marker-ICR010_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-1_peak.marker-ICR010_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-2_peak.marker-UNC4609527_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-2_peak.marker-UNC4609660_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-B6_rs31740628_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-ICR1269_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-ICR5325_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-ICR5347_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-ICR5348_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00104971_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00105059_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00105203_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00107930_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00107980_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00108068_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00108114_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00109750_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00189245_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00514662_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00517218_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00517674_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00517906_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00518418_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00518684_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00518911r_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00520122_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00524717_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00524828_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00525380_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00525709_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00527291_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00527587_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00529707_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00529707r_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00531601_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-JAX00532122_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4673471_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4681411_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4793677_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4799475_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4809586_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4820379_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4875653_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4909830_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4912288_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4914130_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4923398_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4945186_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4947492_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4957168_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4957859_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4964062_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4979914_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4985064_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4990111_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC4991565_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5001723_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5042361_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5055259_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5076935_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5082757_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5173293_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5182782_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5192582_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5193970_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5213888_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5240484_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5256922_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5266664_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5266711_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5267813_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5274554_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5277061_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5282259_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5287977_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5301460_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5321559_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5322455_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5328596_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5350384_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5354724_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5375208_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5384072_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5400314_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5433945_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5435874_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5466669_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5517972_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5522257_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5523265_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5532424_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5553882_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5572051_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5582753_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5589260_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5595555_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5623239_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5639830_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5680234_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5682314_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5725356_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5727351_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5734420_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5743257_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5751567_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5752623_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5776974_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5778977_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5788507_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5792468_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5794598_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5795561_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5802298_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5806628_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5808246_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5817478_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5836999_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5841501_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5847927_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNC5966417_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008129_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008173_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008182_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008193_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008209_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008215_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008219_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008231_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008235_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008242_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008253_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008256_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008259_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008271_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008311_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008317_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008372_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008379_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008410_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008415_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008431_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008511_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008609_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008614_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008649_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008651_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008656_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008723_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008802_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008803_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008808_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008834_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008838_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008914_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008928_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008948_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008959_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008973_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS008987_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009001_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009025_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009034_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009087_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009109_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009142_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009207_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009216_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009224_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009306_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009316_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009364_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009377_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009382_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009383_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009387_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009395_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009413_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009428_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009463_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009503_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009532_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009541_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009560_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCHS009574_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCJPD001170_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCJPD001198_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCJPD001276_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCJPD001315_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCJPD001355_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCJPD001373_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-3_peak.marker-UNCJPD001382_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNC6851205_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNC8250659_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNC8299912_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNC8300499_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNC8381981_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNC8436434_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNC8439633_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNC8530763_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNCHS012948_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNCHS012976_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNCHS013118_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNCHS013118_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNCHS013118_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNCHS013130_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNCHS013139_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-4_peak.marker-UNCHS013304_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-5_peak.marker-JAX00590770_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-5_peak.marker-UNC10044126_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-7_peak.marker-ICR1830_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-7_peak.marker-UNC13170794_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-7_peak.marker-UNCHS019480_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-7_peak.marker-cr31snv87_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-8_peak.marker-JAX00667121_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-8_peak.marker-UNC15430711_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-9_peak.marker-UNC17203329_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-9_peak.marker-UNC17203597_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-9_peak.marker-UNCHS024593_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-X_peak.marker-UNC30627130_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-X_peak.marker-UNCHS048313_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_fitqtl_peaks_additive_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_additive_5.batches_mis.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_additive_snpsqc_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_additive_snpsqc_dis_no-x_updated_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_additive_snpsqc_dis_no-x_updated_5.batches_mis.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_interacting_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_interacting_5.batches_mis.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_interacting_snpsqc_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_interacting_snpsqc_dis_no-x_updated_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_interacting_snpsqc_dis_no-x_updated_5.batches_mis.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_sex_additive_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_sex_additive_5.batches_mis.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_sex_additive_snpsqc_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_sex_additive_snpsqc_dis_no-x_updated_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_sex_additive_snpsqc_dis_no-x_updated_5.batches_mis.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_sex_interacting_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_sex_interacting_5.batches_mis.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_sex_interacting_snpsqc_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_sex_interacting_snpsqc_dis_no-x_updated_5.batches.txt
Untracked: data/ici.vs.eoi_fitqtl_peaks_sex_interacting_snpsqc_dis_no-x_updated_5.batches_mis.mis.txt
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-ICR4295_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-JAX00292499_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-JAX00292927_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-JAX00292927_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-JAX00292927_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-UNC18154330_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-UNC18195765_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-UNC18216614_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-UNC18240977_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-UNC18311938_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-UNC18856953_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-UNCHS028236_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-UNCHS028536_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-UNCJPD004316_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-10_peak.marker-UNCJPD009532_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-11_peak.marker-UNC19155926_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-11_peak.marker-UNC19481366_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-11_peak.marker-UNC19970181_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-11_peak.marker-UNC19970181_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-11_peak.marker-UNCJPD004792_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-12_peak.marker-ICR4497_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-13_peak.marker-UNCHS036773_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-13_peak.marker-UNCHS036964_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-14_peak.marker-UNC24056202_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-15_peak.marker-UNC26070435_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-17_peak.marker-UNCrs47191360_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-17_peak.marker-UNCrs47191360_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-18_peak.marker-UNCHS045344_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-19_peak.marker-UNCJPD007087_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-1_peak.marker-ICR010_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-1_peak.marker-ICR010_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-2_peak.marker-UNC4609527_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-2_peak.marker-UNC4609660_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-B6_rs31740628_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-ICR1269_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-ICR5325_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-ICR5347_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-ICR5348_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00104971_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00105059_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00105203_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00107930_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00107980_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00108068_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00108114_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00109750_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00189245_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00514662_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00517218_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00517674_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00517906_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00518418_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00518684_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00518911r_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00520122_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00524717_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00524828_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00525380_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00525709_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00527291_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00527587_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00529707_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00529707r_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00531601_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-JAX00532122_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4673471_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4681411_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4793677_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4799475_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4809586_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4820379_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4875653_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4909830_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4912288_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4914130_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4923398_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4945186_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4947492_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4957168_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4957859_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4964062_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4979914_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4985064_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4990111_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC4991565_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5001723_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5042361_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5055259_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5076935_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5082757_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5173293_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5182782_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5192582_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5193970_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5213888_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5240484_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5256922_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5266664_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5266711_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5267813_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5274554_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5277061_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5282259_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5287977_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5301460_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5321559_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5322455_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5328596_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5350384_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5354724_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5375208_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5384072_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5400314_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5433945_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5435874_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5466669_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5517972_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5522257_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5523265_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5532424_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5553882_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5572051_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5582753_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5589260_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5595555_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5623239_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5639830_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5680234_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5682314_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5725356_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5727351_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5734420_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5743257_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5751567_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5752623_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5776974_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5778977_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5788507_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5792468_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5794598_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5795561_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5802298_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5806628_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5808246_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5817478_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5836999_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5841501_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5847927_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNC5966417_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008129_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008173_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008182_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008193_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008209_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008215_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008219_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008231_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008235_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008242_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008253_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008256_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008259_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008271_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008311_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008317_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008372_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008379_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008410_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008415_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008431_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008511_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008609_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008614_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008649_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008651_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008656_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008723_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008802_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008803_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008808_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008834_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008838_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008914_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008928_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008948_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008959_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008973_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS008987_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009001_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009025_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009034_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009087_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009109_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009142_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009207_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009216_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009224_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009306_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009316_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009364_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009377_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009382_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009383_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009387_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009395_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009413_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009428_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009463_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009503_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009532_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009541_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009560_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCHS009574_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCJPD001170_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCJPD001198_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCJPD001276_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCJPD001315_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCJPD001355_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCJPD001373_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-3_peak.marker-UNCJPD001382_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNC6851205_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNC8250659_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNC8299912_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNC8300499_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNC8381981_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNC8436434_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNC8439633_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNC8530763_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNCHS012948_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNCHS012976_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNCHS013118_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNCHS013118_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNCHS013118_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNCHS013130_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNCHS013139_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-4_peak.marker-UNCHS013304_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-5_peak.marker-JAX00590770_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-5_peak.marker-UNC10044126_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-7_peak.marker-ICR1830_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-7_peak.marker-UNC13170794_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-7_peak.marker-UNCHS019480_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.eoi_genes_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-7_peak.marker-cr31snv87_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-8_peak.marker-JAX00667121_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-8_peak.marker-UNC15430711_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-9_peak.marker-UNC17203329_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-9_peak.marker-UNC17203597_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-9_peak.marker-UNCHS024593_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-X_peak.marker-UNC30627130_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-X_peak.marker-UNCHS048313_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_gm_qtl_5.batches.csv
Untracked: data/ici.vs.eoi_gm_qtl_5.batches_mis.csv
Untracked: data/ici.vs.eoi_gm_qtl_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_gm_qtl_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.eoi_gm_qtl_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.eoi_gm_qtl_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_geno.ratio.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_geno.ratio_5.batches.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_geno.ratio_5.batches_0.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_geno.ratio_5.batches_52.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_geno.ratio_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_geno.ratio_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_5.batches.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_5.batches_0.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_5.batches_52.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_geno.ratio.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_geno.ratio_5.batches.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_geno.ratio_5.batches_0.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_geno.ratio_5.batches_52.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_geno.ratio_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_geno.ratio_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_5.batches.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_5.batches_0.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_5.batches_52.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-additive.covariates_blup_sub_chr-7_peak.marker-UNC13239362_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_rz.age-additive.covariates_genes_chr-7_peak.marker-UNC13239362_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-JAX00292927_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-UNC18311938_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-UNC18343181_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-UNC18376338_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-UNC18376338_lod.drop-1.5_snpsqc_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-UNC18376338_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-UNC18382395_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-UNCHS028236_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-UNCHS028236_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-UNCHS028536_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-10_peak.marker-UNCHS029024_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-11_peak.marker-UNC19970181_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-11_peak.marker-UNC19970181_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-13_peak.marker-UNCHS036773_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-13_peak.marker-UNCHS036773_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-15_peak.marker-UNC26070435_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-15_peak.marker-UNC26070435_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-17_peak.marker-UNCrs47191360_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-1_peak.marker-UNC228500_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-1_peak.marker-UNC228500_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-2_peak.marker-UNC4609527_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-2_peak.marker-UNC4609527_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-ICR2047_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-JAX00112499_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-JAX00219554_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-JAX00220631_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-JAX00514662_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-JAX00529707_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC4836550_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC4837069_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC4912039_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC4912039_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC4956618_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC4957168_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5006948_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5192582_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5213888_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5213888_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5262472_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5262472_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5266664_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5300878_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5360055_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5679155_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5808246_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC5945286_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNC6227106_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008182_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008511_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008511_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008595_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008613_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008613_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009067_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009383_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCHS009693_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCJPD001246_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-UNCJPD001341_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-3_peak.marker-sanger2623_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-4_peak.marker-UNCHS013125_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-4_peak.marker-UNCHS013125_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-6_peak.marker-UNCHS018170_lod.drop-1.5_snpsqc_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-6_peak.marker-UNCHS018231_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-9_peak.marker-UNC17203597_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-9_peak.marker-UNC17203597_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-JAX00292927_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-UNC18311938_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-UNC18343181_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-UNC18376338_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-UNC18376338_lod.drop-1.5_snpsqc_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-UNC18376338_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-UNC18382395_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-UNCHS028236_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-UNCHS028236_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-UNCHS028536_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-10_peak.marker-UNCHS029024_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-11_peak.marker-UNC19970181_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-11_peak.marker-UNC19970181_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-12_peak.marker-ICR499_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-13_peak.marker-UNCHS036773_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-13_peak.marker-UNCHS036773_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-15_peak.marker-UNC26070435_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-15_peak.marker-UNC26070435_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-17_peak.marker-UNCHS044241_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-17_peak.marker-UNCrs47191360_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-1_peak.marker-UNC228500_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-1_peak.marker-UNC228500_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-2_peak.marker-UNC4609527_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-2_peak.marker-UNC4609527_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-ICR1338_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-ICR2047_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-JAX00106957_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-JAX00112499_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-JAX00219554_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-JAX00220631_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-JAX00514662_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-JAX00529707_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC4813186_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC4836550_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC4837069_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC4912039_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC4912039_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC4924377_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC4956618_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC4957168_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5006948_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5008656_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5083168_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5192582_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5213888_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5213888_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5262472_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5262472_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5266664_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5300878_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5360055_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5454839_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5645844_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5679155_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5808246_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC5945286_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNC6227106_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008182_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008286_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008487_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008511_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008511_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008595_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008613_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008613_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS008815_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS009067_lod.drop-1.5_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS009383_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCHS009693_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCJPD001246_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-UNCJPD001341_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-3_peak.marker-sanger2623_lod.drop-1.5_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-4_peak.marker-UNCHS013125_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-4_peak.marker-UNCHS013125_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-6_peak.marker-UNCHS018170_lod.drop-1.5_snpsqc_5.batches_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-6_peak.marker-UNCHS018231_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-7_peak.marker-UNCHS020066_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-9_peak.marker-UNC17203597_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-9_peak.marker-UNC17203597_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_0.csv
Untracked: data/ici.vs.eoi_rz.age-no.covariates_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_52.csv
Untracked: data/ici.vs.eoi_sample.genos_marker.freq_low.geno.freq.removed.csv
Untracked: data/ici.vs.eoi_sample.genos_marker.freq_low.geno.freq.removed_sample.outliers.removed.csv
Untracked: data/ici.vs.eoi_sample.genos_marker.freq_low.probs.freq.removed.csv
Untracked: data/ici.vs.eoi_sample.genos_marker.freq_low.probs.freq.removed_sample.outliers.removed.csv
Untracked: data/ici.vs.eoi_scanone_5.batches.Rdata
Untracked: data/ici.vs.eoi_scanone_5.batches_mis.Rdata
Untracked: data/ici.vs.eoi_scanone_snpsqc_5.batches.Rdata
Untracked: data/ici.vs.eoi_scanone_snpsqc_5.batches_mis.Rdata
Untracked: data/ici.vs.eoi_scanone_snpsqc_dis_no-x_updated_5.batches.Rdata
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-14_peak.marker-UNC24056202_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-17_peak.marker-UNCJPD006614_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-18_peak.marker-UNC28655293_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-5_peak.marker-UNC10044126_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-5_peak.marker-UNC8675939_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-8_peak.marker-JAX00667121_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_blup_sub_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_genes_chr-14_peak.marker-UNC24056202_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_genes_chr-14_peak.marker-UNC24056202_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_genes_chr-17_peak.marker-UNCJPD006614_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_genes_chr-18_peak.marker-UNC28655293_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_genes_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_genes_chr-5_peak.marker-UNC10044126_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_genes_chr-5_peak.marker-UNC8675939_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_genes_chr-8_peak.marker-JAX00667121_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_genes_chr-8_peak.marker-UNC15524531_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_age.of.onset-no.covariates_genes_chr-X_peak.marker-UNCHS048314_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_bad.markers_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-10_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-10_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-10_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-10_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-10_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-11_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-11_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-11_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-11_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-11_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-12_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-12_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-12_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-12_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-12_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-13_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-13_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-13_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-13_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-13_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-14_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-14_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-14_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-14_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-14_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-15_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-15_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-15_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-15_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-15_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-16_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-16_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-16_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-16_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-16_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-17_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-17_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-17_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-17_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-17_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-18_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-18_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-18_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-18_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-18_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-19_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-19_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-19_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-19_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-19_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-1_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-1_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-1_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-1_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-1_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-2_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-2_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-2_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-2_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-2_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-3_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-3_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-3_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-3_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-3_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-4_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-4_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-4_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-4_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-4_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-5_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-6_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-6_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-6_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-6_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-6_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-7_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-7_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-7_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-7_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-7_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-8_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-8_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-8_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-8_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-8_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-9_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-9_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-9_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-9_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-9_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-X_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-X_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-X_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.full_chr-X_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.full_chr-X_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-10_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-10_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-10_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-10_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-10_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-10_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-10_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-10_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-11_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-11_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-11_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-11_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-11_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-11_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-11_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-11_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-12_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-12_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-12_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-12_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-12_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-12_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-12_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-12_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-13_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-13_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-13_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-13_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-13_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-13_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-13_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-13_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-14_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-14_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-14_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-14_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-14_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-14_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-14_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-15_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-15_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-15_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-15_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-15_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-16_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-16_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-16_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-16_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-16_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-16_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-16_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-17_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-17_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-17_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-17_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-17_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-18_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-18_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-18_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-18_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-18_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-18_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-18_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-19_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-19_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-19_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-19_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-19_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-19_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-19_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-1_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-1_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-1_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-1_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-1_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-1_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-1_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-1_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-2_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-2_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-2_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-2_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-2_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-2_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-2_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-2_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-3_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-3_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-3_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-3_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-3_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-3_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-3_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-3_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-4_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-4_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-4_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-4_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-4_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-4_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-4_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-4_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-5_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-5_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-6_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-6_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-6_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-6_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-6_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-6_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-6_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-6_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-7_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-7_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-7_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-7_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-7_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-7_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-7_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-7_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-8_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-8_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-8_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-8_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-8_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-8_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-8_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-8_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-9_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-9_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-9_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-9_lod.drop-1.5_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-9_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-9_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-9_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-9_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-X_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-X_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-X_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-X_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-X_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-X_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup.qc_chr-X_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-10_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-10_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-10_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-10_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-10_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-11_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-11_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-11_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-11_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-11_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-12_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-12_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-12_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-12_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-12_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-13_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-13_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-13_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-13_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-13_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-14_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-14_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-14_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-14_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-14_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-15_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-15_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-15_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-16_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-16_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-16_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-16_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-16_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-17_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-17_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-17_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-18_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-18_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-18_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-18_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-18_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-19_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-19_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-19_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-19_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-19_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-1_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-1_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-1_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-1_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-1_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-2_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-2_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-2_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-2_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-2_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-3_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-3_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-3_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-3_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-3_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-4_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-4_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-4_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-4_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-4_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-5_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-5_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-5_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-5_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-5_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-6_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-6_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-6_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-6_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-6_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-7_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-7_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-7_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-7_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-7_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-8_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-8_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-8_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-8_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-8_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-9_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-9_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-9_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-9_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-9_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-X_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-X_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-X_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_chr-X_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_blup_chr-X_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_sub_chr-13_peak.marker-UNC22384241_lod.drop-1.5_snpsqc_snpsqc_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_blup_sub_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_blup_sub_chr-19_peak.marker-UNC29893228_lod.drop-1.5_snpsqc_snpsqc_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_blup_sub_chr-19_peak.marker-UNC29990033_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_blup_sub_chr-19_peak.marker-UNCHS047192_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_peaks.csv
Untracked: data/ici.vs.pbs_blup_sub_chr-8_peak.marker-UNCHS023353_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_peaks.csv
Untracked: data/ici.vs.pbs_blup_sub_chr-8_peak.marker-UNCHS023353_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_blup_sub_chr-8_peak.marker-UNCHS023364_lod.drop-1.5_snpsqc_snpsqc_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_fitqtl_peaks_additive_snpsqc_dis_no-x_updated_5.batches_mis_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_additive_snpsqc_dis_no-x_updated_5.batches_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_additive_snpsqc_snpsqc_5.batches_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_interacting_snpsqc_dis_no-x_updated_5.batches_mis_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_interacting_snpsqc_dis_no-x_updated_5.batches_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_interacting_snpsqc_snpsqc_5.batches_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_sex_additive_snpsqc_dis_no-x_updated_5.batches_mis_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_sex_additive_snpsqc_dis_no-x_updated_5.batches_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_sex_additive_snpsqc_snpsqc_5.batches_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_sex_interacting_snpsqc_dis_no-x_updated_5.batches_mis_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_sex_interacting_snpsqc_dis_no-x_updated_5.batches_peaks.txt
Untracked: data/ici.vs.pbs_fitqtl_peaks_sex_interacting_snpsqc_snpsqc_5.batches_peaks.txt
Untracked: data/ici.vs.pbs_genes_chr-13_peak.marker-UNC22384241_lod.drop-1.5_snpsqc_snpsqc_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_genes_chr-18_peak.marker-UNCHS045343_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_genes_chr-19_peak.marker-UNC29893228_lod.drop-1.5_snpsqc_snpsqc_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_genes_chr-19_peak.marker-UNC29990033_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_genes_chr-19_peak.marker-UNCHS047192_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_peaks.csv
Untracked: data/ici.vs.pbs_genes_chr-8_peak.marker-UNCHS023353_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis_peaks.csv
Untracked: data/ici.vs.pbs_genes_chr-8_peak.marker-UNCHS023353_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_genes_chr-8_peak.marker-UNCHS023364_lod.drop-1.5_snpsqc_snpsqc_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_gm_qtl_5.batches.csv
Untracked: data/ici.vs.pbs_gm_qtl_5.batches_mis.csv
Untracked: data/ici.vs.pbs_gm_qtl_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_gm_qtl_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_gm_qtl_snpsqc_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_gm_qtl_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_gm_qtl_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_gm_qtl_snpsqc_dis_no-x_updated_5.batches_mis_peaks.csv
Untracked: data/ici.vs.pbs_gm_qtl_snpsqc_dis_no-x_updated_5.batches_peaks.csv
Untracked: data/ici.vs.pbs_marker.freq_low.geno.freq.removed_geno.ratio.csv
Untracked: data/ici.vs.pbs_marker.freq_low.geno.freq.removed_geno.ratio_5.batches.csv
Untracked: data/ici.vs.pbs_marker.freq_low.geno.freq.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici.vs.pbs_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio.csv
Untracked: data/ici.vs.pbs_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_5.batches.csv
Untracked: data/ici.vs.pbs_marker.freq_low.geno.freq.removed_sample.outliers.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici.vs.pbs_marker.freq_low.probs.freq.removed_geno.ratio.csv
Untracked: data/ici.vs.pbs_marker.freq_low.probs.freq.removed_geno.ratio_5.batches.csv
Untracked: data/ici.vs.pbs_marker.freq_low.probs.freq.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici.vs.pbs_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio.csv
Untracked: data/ici.vs.pbs_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_5.batches.csv
Untracked: data/ici.vs.pbs_marker.freq_low.probs.freq.removed_sample.outliers.removed_geno.ratio_5.batches_mis.csv
Untracked: data/ici.vs.pbs_rz.age-additive.covariates_blup_sub_chr-7_peak.marker-JAX00153527_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_rz.age-additive.covariates_blup_sub_chr-7_peak.marker-JAX00153527_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_rz.age-additive.covariates_blup_sub_chr-7_peak.marker-UNCHS020443_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_rz.age-additive.covariates_blup_sub_chr-7_peak.marker-UNCHS020443_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_rz.age-additive.covariates_blup_sub_chr-7_peak.marker-UNCHS020486_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_rz.age-additive.covariates_genes_chr-7_peak.marker-JAX00153527_lod.drop-1.5_5.batches.csv
Untracked: data/ici.vs.pbs_rz.age-additive.covariates_genes_chr-7_peak.marker-JAX00153527_lod.drop-1.5_snpsqc_5.batches.csv
Untracked: data/ici.vs.pbs_rz.age-additive.covariates_genes_chr-7_peak.marker-UNCHS020443_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches.csv
Untracked: data/ici.vs.pbs_rz.age-additive.covariates_genes_chr-7_peak.marker-UNCHS020443_lod.drop-1.5_snpsqc_dis_no-x_updated_5.batches_mis.csv
Untracked: data/ici.vs.pbs_rz.age-additive.covariates_genes_chr-7_peak.marker-UNCHS020486_lod.drop-1.5_snpsqc_5.batches_mis.csv
Untracked: data/ici.vs.pbs_sample.genos_marker.freq_low.geno.freq.removed.csv
Untracked: data/ici.vs.pbs_sample.genos_marker.freq_low.geno.freq.removed_sample.outliers.removed.csv
Untracked: data/ici.vs.pbs_sample.genos_marker.freq_low.probs.freq.removed.csv
Untracked: data/ici.vs.pbs_sample.genos_marker.freq_low.probs.freq.removed_sample.outliers.removed.csv
Untracked: data/ici.vs.pbs_scanone_5.batches.Rdata
Untracked: data/ici.vs.pbs_scanone_5.batches_mis.Rdata
Untracked: data/ici.vs.pbs_scanone_snpsqc_5.batches.Rdata
Untracked: data/ici.vs.pbs_scanone_snpsqc_5.batches_mis.Rdata
Untracked: data/ici.vs.pbs_scanone_snpsqc_dis_no-x_updated_5.batches.Rdata
Untracked: data/ici.vs.pbs_scanone_snpsqc_dis_no-x_updated_5.batches_mis.Rdata
Untracked: data/percent_missing_id_3.batches.RData
Untracked: data/percent_missing_id_4.batches.RData
Untracked: data/percent_missing_id_4.batches_bc.RData
Untracked: data/percent_missing_id_5.batches.RData
Untracked: data/percent_missing_marker_4.batches.RData
Untracked: data/percent_missing_marker_5.batches.RData
Untracked: data/pheno.csv
Untracked: data/pheno_BC312.csv
Untracked: data/physical_map.csv
Untracked: data/physical_map_BC312.csv
Untracked: data/qc_info_bad_sample_3.batches.RData
Untracked: data/qc_info_bad_sample_4.batches.RData
Untracked: data/qc_info_bad_sample_4.batches_bc.RData
Untracked: data/qc_info_bad_sample_5.batches.RData
Untracked: data/remaining.markers_geno.freq.xlsx
Untracked: data/sample_geno.csv
Untracked: data/sample_geno_AHB_BC312.csv
Untracked: data/sample_geno_bc.csv
Untracked: data/sample_geno_bc_BC312.csv
Untracked: data/serreze_probs.rds
Untracked: data/serreze_probs_BC312.rds
Untracked: data/serreze_probs_allqc.rds
Untracked: data/serreze_probs_allqc_5.batches.rds
Untracked: data/serreze_probs_allqc_5.batches_mis.rds
Untracked: data/summary.cg_3.batches.RData
Untracked: data/summary.cg_4.batches.RData
Untracked: data/summary.cg_4.batches_bc.RData
Untracked: data/summary.cg_5.batches.RData
Untracked: output/Percent_missing_genotype_data_5.batches.pdf
Untracked: output/Percent_missing_genotype_data_per_marker_5.batches.pdf
Untracked: output/Proportion_matching_genotypes_before_removal_of_bad_samples_5.batches.pdf
Untracked: output/genotype_error_marker_5.batches.pdf
Untracked: output/genotype_frequency_marker_5.batches.pdf
Unstaged changes:
Modified: analysis/4.1.1_qtl.analysis_binary_ici.vs.eoi_snpsqc_dis_no-x_updated.Rmd
Modified: analysis/4.1.1_qtl.analysis_binary_ici.vs.pbs_snpsqc_dis_no-x_updated.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici-early.vs.pbs_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici-early.vs.pbs_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici-early.vs.pbs_snpsqc_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici-early.vs.pbs_snpsqc_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici-early.vs.pbs_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici-early.vs.pbs_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.pbs_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.pbs_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.pbs_snpsqc_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.pbs_snpsqc_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.pbs_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_age_ici.vs.pbs_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici-early.vs.pbs_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici-early.vs.pbs_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici-early.vs.pbs_snpsqc_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici-early.vs.pbs_snpsqc_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici-early.vs.pbs_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici-early.vs.pbs_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.eoi_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.pbs_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.pbs_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.pbs_snpsqc_pheno.corrected.cleaned_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.pbs_snpsqc_pheno.corrected.cleaned_5.batches_mis.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.pbs_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches.Rmd
Modified: analysis/4.1.2_qtl.analysis_cont_rz.age_ici.vs.pbs_snpsqc_pheno.corrected.cleaned_dis_no-xk_5.batches_mis.Rmd
Modified: analysis/index_5.batches.Rmd
Modified: analysis/index_5.batches_additional.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
There are no past versions. Publish this analysis with wflow_publish() to start tracking its development.
Loading Data
load("data/gm_allqc_5.batches.RData")
#gm_allqc
gm=gm_allqc
gmObject of class cross2 (crosstype "bc")
Total individuals 308
No. genotyped individuals 308
No. phenotyped individuals 308
No. with both geno & pheno 308
No. phenotypes 1
No. covariates 6
No. phenotype covariates 0
No. chromosomes 20
Total markers 34537
No. markers by chr:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16
2643 2629 1857 1890 1774 1941 1672 1627 1878 1176 1871 1300 1549 1578 1257 935
17 18 19 X
501 913 1014 4532 #pr <- readRDS("data/serreze_probs_allqc.rds")
#pr <- readRDS("data/serreze_probs.rds")
##extracting animals with ici and eoi group status
miceinfo <- gm$covar[gm$covar$group == "EOI" | gm$covar$group == "ICI",]
table(miceinfo$group)
EOI ICI
164 104 mice.ids <- rownames(miceinfo)
gm <- gm[mice.ids]
gmObject of class cross2 (crosstype "bc")
Total individuals 268
No. genotyped individuals 268
No. phenotyped individuals 268
No. with both geno & pheno 268
No. phenotypes 1
No. covariates 6
No. phenotype covariates 0
No. chromosomes 20
Total markers 34537
No. markers by chr:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16
2643 2629 1857 1890 1774 1941 1672 1627 1878 1176 1871 1300 1549 1578 1257 935
17 18 19 X
501 913 1014 4532 table(gm$covar$group)
EOI ICI
164 104 gm.full <- gm
covars <- read_csv("data/covar_corrected.cleaned_ici.vs.eoi_5.batches_52.csv")
#removing any missing info
missing_ids <- covars[which(covars$"rz.age"==""),]$Mouse.ID
length(missing_ids)[1] 1missing_ids[1] "NG01228-ICI"if(length(missing_ids) != 0){
gm <- gm[paste0("-",as.character(missing_ids)),]
gm
table(gm$covar$group)
#keeping only mouseids in covars
covars <- covars[gm$covar$id,]
table(covars$group)
}
EOI ICI
164 103 #covars[covars$age.of.onset==0,]$age.of.onset <- ''
query_variants <- create_variant_query_func("/Users/corneb/Documents/MyJax/CS/Projects/support.files/qtl2/cc_variants.sqlite")
query_genes <- create_gene_query_func("/Users/corneb/Documents/MyJax/CS/Projects/support.files/qtl2/mouse_genes_mgi.sqlite")
##removing problmetic marker
gm <- drop_markers(gm, "UNCHS013106")
#pmap_interp = interp_map(map, gm$gmap, gm$pmap)
markers <- marker_names(gm)
gmapdf <- read.csv("/Users/corneb/Documents/MyJax/CS/Projects/Serreze/haplotype.reconstruction/output_5.batches/genetic_map.csv")
pmapdf <- read.csv("/Users/corneb/Documents/MyJax/CS/Projects/Serreze/haplotype.reconstruction/output_5.batches/physical_map.csv")
#mapdf <- merge(gmapdf,pmapdf, by=c("marker","chr"), all=T)
#rownames(mapdf) <- mapdf$marker
#mapdf <- mapdf[markers,]
#names(mapdf) <- c('marker','chr','gmapdf','pmapdf')
#mapdfnd <- mapdf[!duplicated(mapdf[c(2:3)]),]
pr.qc <- calc_genoprob(gm)
gmObject of class cross2 (crosstype "bc")
Total individuals 267
No. genotyped individuals 267
No. phenotyped individuals 267
No. with both geno & pheno 267
No. phenotypes 1
No. covariates 6
No. phenotype covariates 0
No. chromosomes 20
Total markers 34537
No. markers by chr:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16
2643 2629 1857 1890 1774 1941 1672 1627 1878 1176 1871 1300 1549 1578 1257 935
17 18 19 X
501 913 1014 4532 Preparing necessary info
covars$rz.age <- as.numeric(covars$rz.age)
Xcovar <- get_x_covar(gm)
addcovar = model.matrix(~ICI.vs.EOI, data = covars)[,-1]
#addcovar.i = model.matrix(~ICI.vs.EOI, data = covars)[,-1]
K <- calc_kinship(pr.qc, type = "loco")
#heatmap(K[[1]])
K.overall <- calc_kinship(pr.qc, type = "overall")
#heatmap(K.overall)
kinship <- calc_kinship(pr.qc)
heatmap(kinship)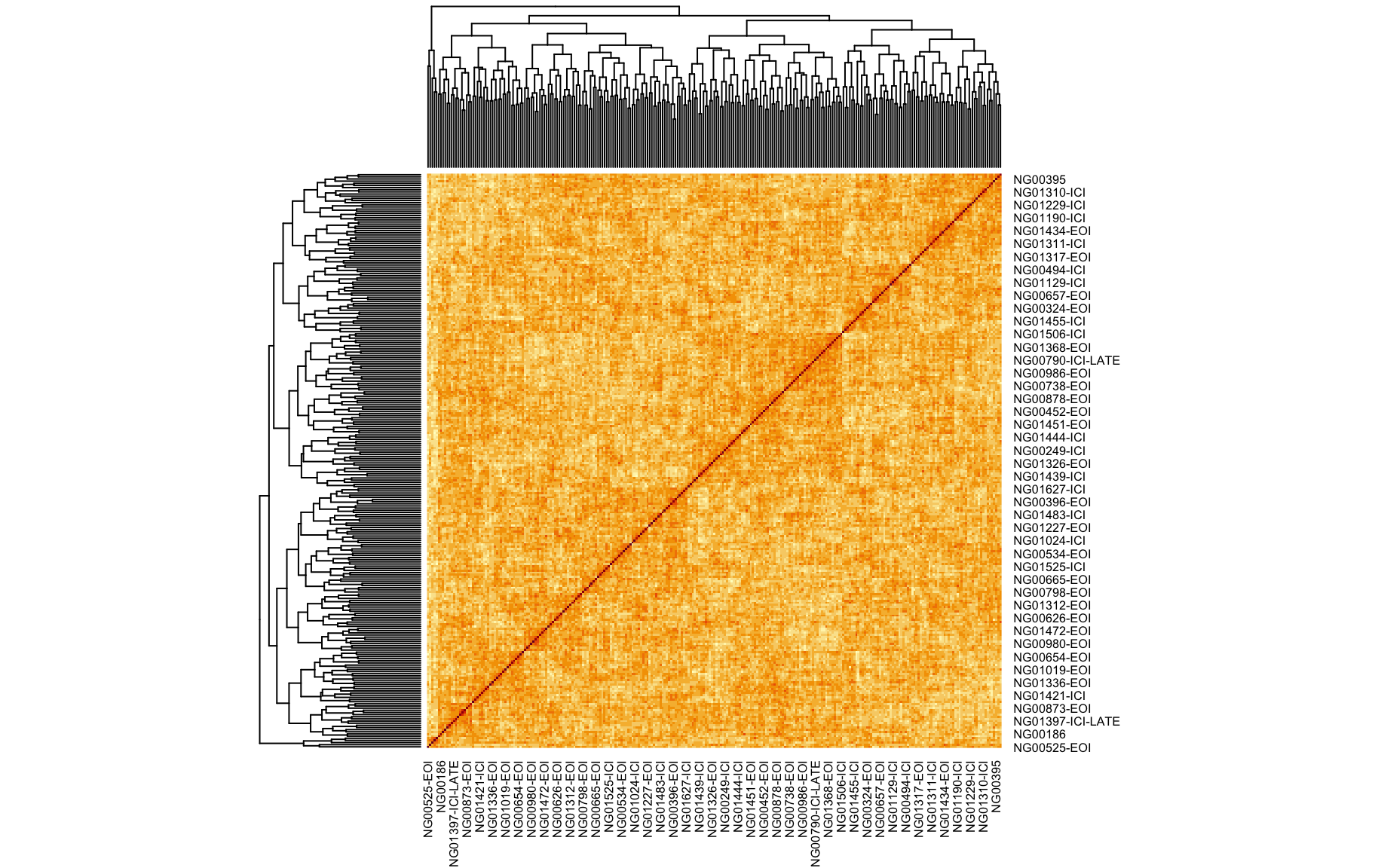
No additive covariates
Significance thresholds
operm <- scan1perm(pr.qc, covars["rz.age"], model="normal", n_perm=10, perm_Xsp=TRUE, chr_lengths=chr_lengths(gm$gmap))
summary_table<-data.frame(unclass(summary(operm, alpha=c(0.01, 0.05, 0.1))))
names(summary_table) <- c("autosomes","X")
summary_table$significance.level <- rownames(summary_table)
rownames(summary_table) <- NULL
summary_table[c(3,1:2)] %>%
kable(escape = F,align = c("ccc")) %>%
kable_styling("striped", full_width = T) %>%
column_spec(1, bold=TRUE)| significance.level | autosomes | X |
|---|---|---|
| 0.01 | 3.486372 | 4.774895 |
| 0.05 | 3.428501 | 4.647653 |
| 0.1 | 3.355982 | 4.481263 |
Genome-wide scan
plot_lod<-function(out,map){
for (i in 1:dim(out)[2]){
#par(mar=c(5.1, 6.1, 1.1, 1.1))
ymx <- maxlod(out) # overall maximum LOD score
plot(out, map, lodcolumn=i, col="slateblue", ylim=c(0, ymx+0.5))
#legend("topright", lwd=2, colnames(out)[i], bg="gray90")
title(main = paste0(colnames(out)[i], " - ICI vs. EOI (no covariates) [positions in cM]"))
add_threshold(map, summary(operm,alpha=0.1), col = 'purple')
add_threshold(map, summary(operm, alpha=0.05), col = 'red')
add_threshold(map, summary(operm, alpha=0.01), col = 'blue')
}
}
print("with no kinship")[1] “with no kinship”
outf <- scan1(pr.qc, covars["rz.age"], Xcovar=Xcovar, model="normal")
plot_lod(outf,gm$gmap)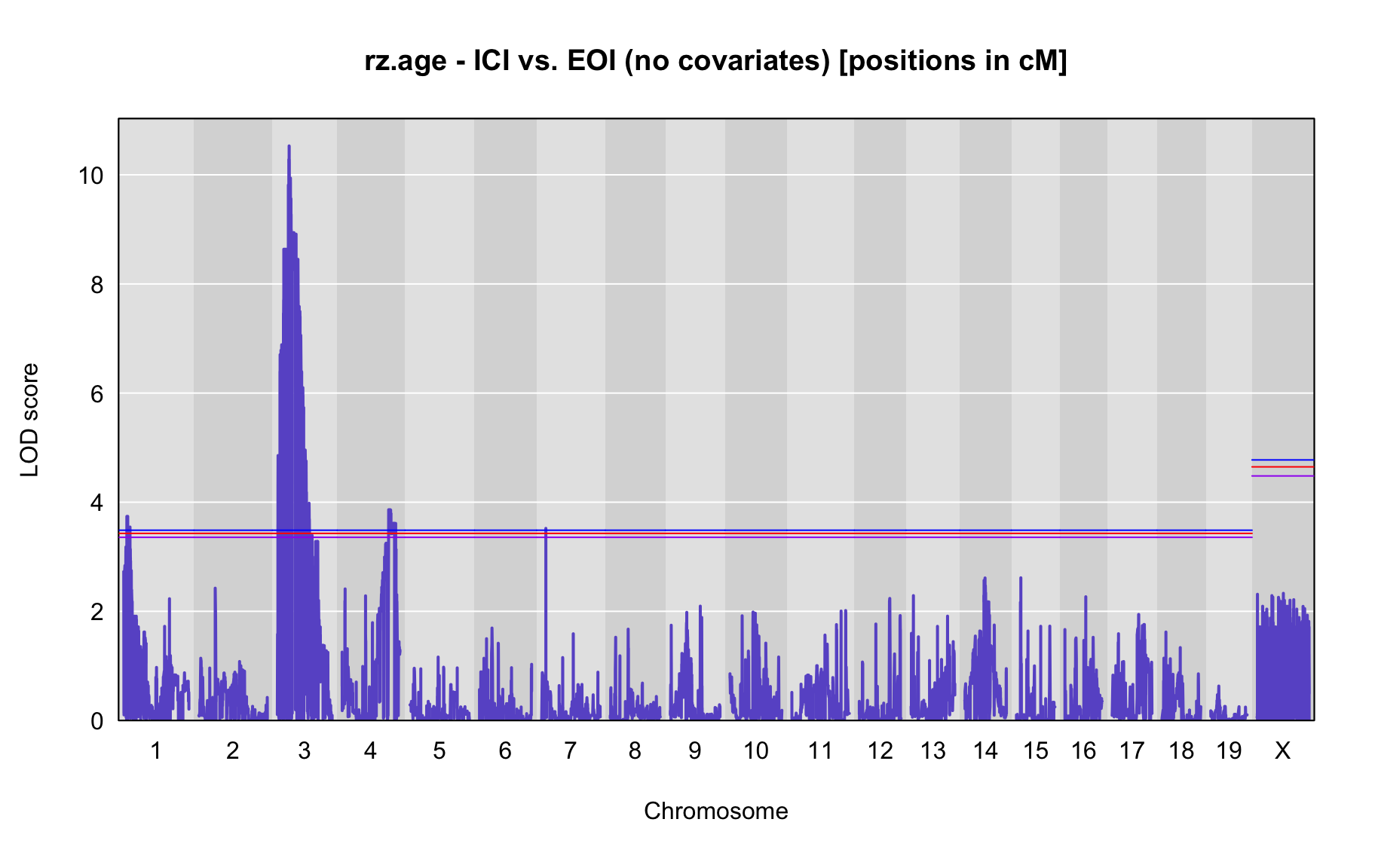
print("with normal kinship")[1] “with normal kinship”
outk <- scan1(pr.qc, covars["rz.age"], Xcovar=Xcovar, model="normal", kinship=kinship)
plot_lod(outk,gm$gmap)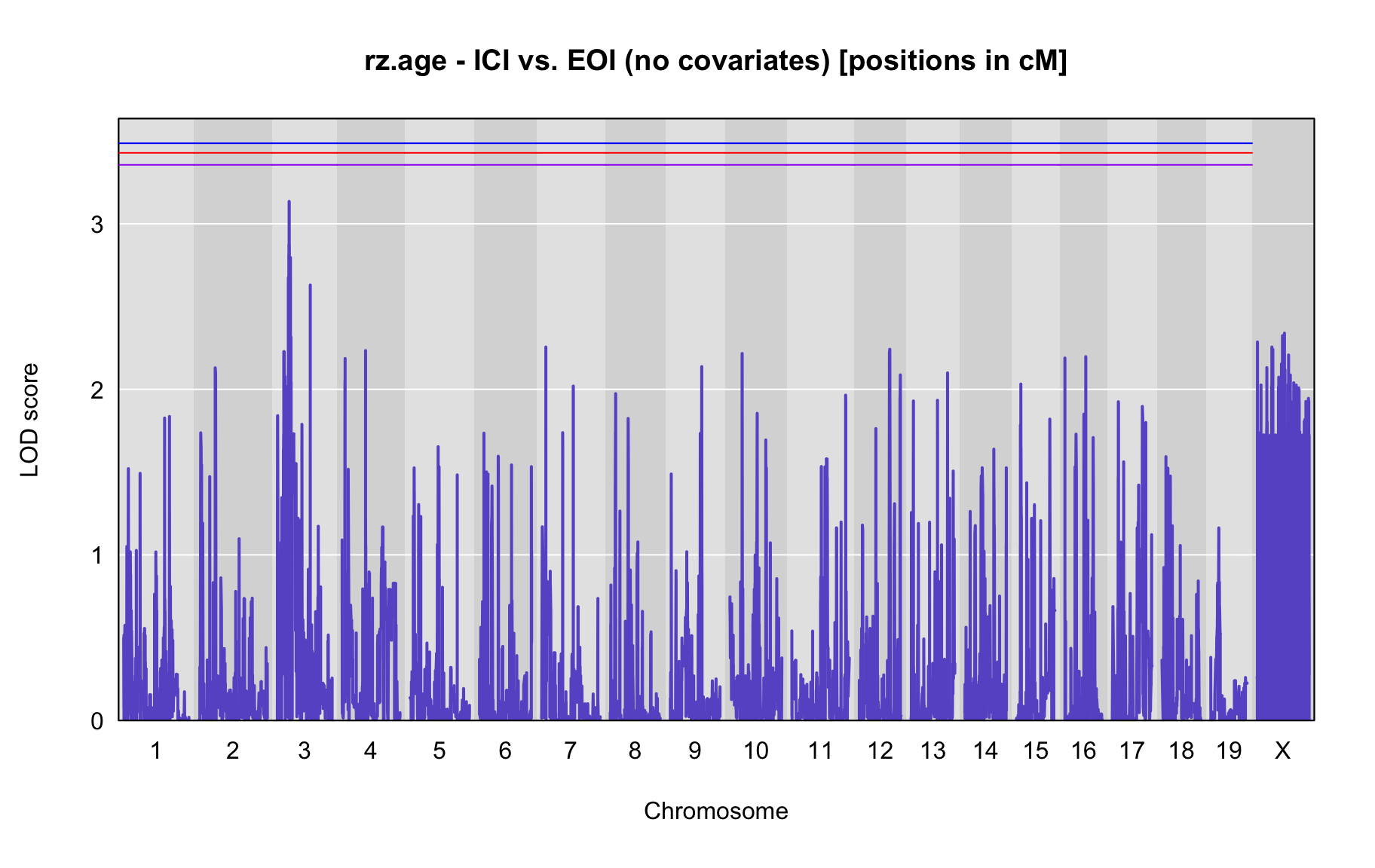
print("with loco kinship")[1] “with loco kinship”
out <- scan1(pr.qc, covars["rz.age"], Xcovar=Xcovar, model="normal", kinship=K)
plot_lod(out,gm$gmap)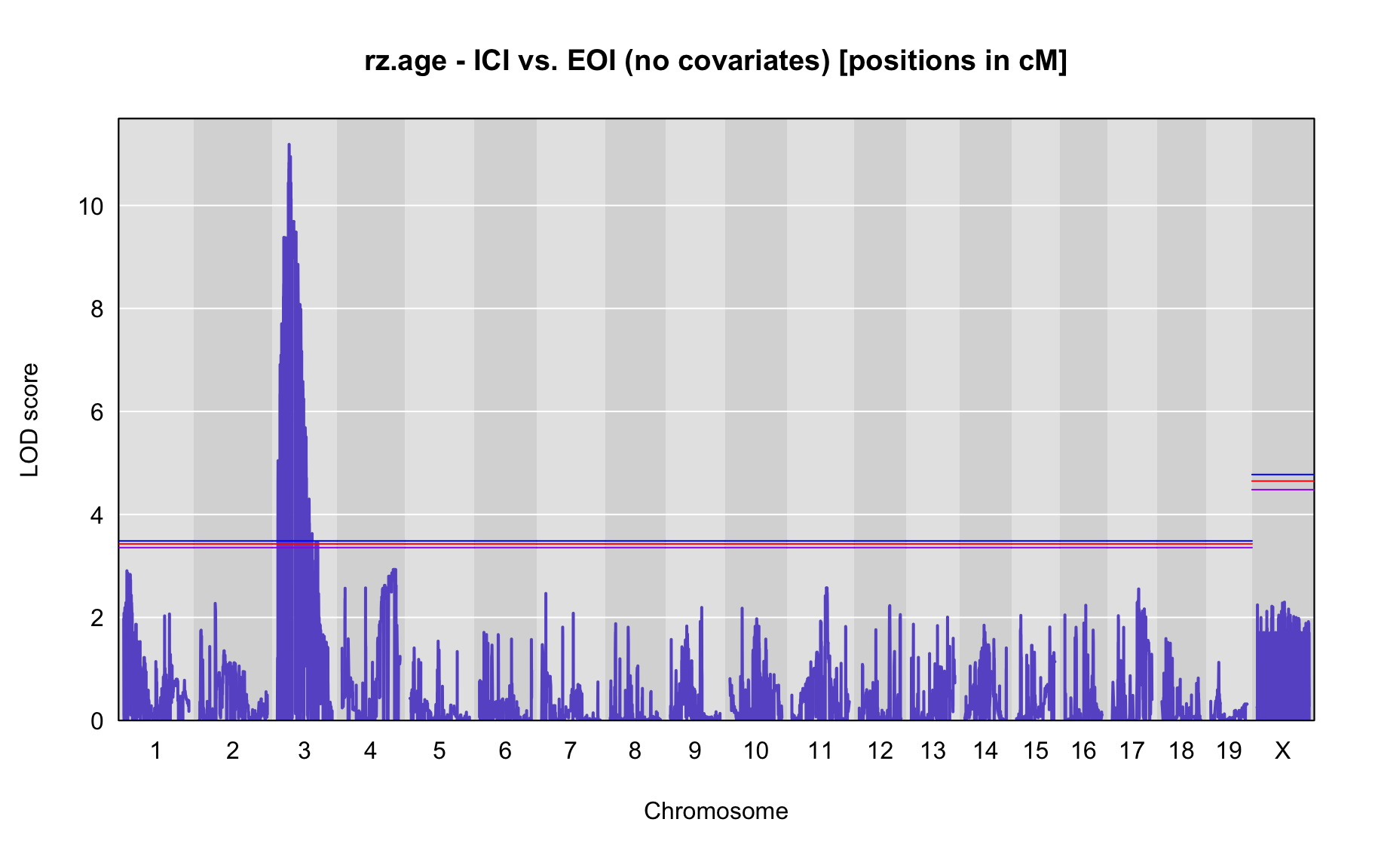
print(paste0("number of samples in analysis = ", as.data.frame(attributes(out)$sample_size)[1,]))[1] “number of samples in analysis = 267”
LOD peaks
Centimorgan (cM)
peaks<-find_peaks(out, gm$gmap, threshold=summary(operm,alpha=0.05)$A, thresholdX = summary(operm,alpha=0.05)$X, peakdrop=3, drop=1.5)
if(nrow(peaks) > 0){
peaks$marker <- find_marker(gm$gmap, chr=peaks$chr,pos=peaks$pos)
names(peaks)[2] <- c("phenotype")
peaks <- peaks[-1]
rownames(peaks) <- NULL
print(kable(peaks, escape = F, align = c("cccccccc"), "html")
%>% kable_styling("striped", full_width = T)%>%
column_spec(1, bold=TRUE)
)
#plot only peak chromosomes
plot_lod_chr<-function(out,map,chrom){
for (i in 1:dim(out)[2]){
#png(filename=paste0("/Users/chenm/Documents/qtl/Jai/",colnames(out)[i], "_lod.png"))
#par(mar=c(5.1, 6.1, 1.1, 1.1))
ymx <- maxlod(out) # overall maximum LOD score
plot(out, map, chr = chrom, lodcolumn=i, col="slateblue", ylim=c(0, ymx+0.5))
#legend("topright", lwd=2, colnames(out)[i], bg="gray90")
title(main = paste0(colnames(out)[i], " - chr", chrom, " - ICI vs. EOI (no covariates) [positions in cM]"))
add_threshold(map, summary(operm,alpha=0.1), col = 'purple')
add_threshold(map, summary(operm, alpha=0.05), col = 'red')
add_threshold(map, summary(operm, alpha=0.01), col = 'blue')
#for (j in 1: dim(summary_table)[1]){
# abline(h=summary_table[j, i],col="red")
# text(x=400, y =summary_table[j, i]+0.12, labels = paste("p=", row.names(summary_table)[j]))
#}
#dev.off()
}
}
} else {
print(paste0("There are no peaks that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}| phenotype | chr | pos | lod | ci_lo | ci_hi | marker |
|---|---|---|---|---|---|---|
| rz.age | 3 | 2.28000 | 5.043745 | 2.010 | 2.309 | JAX00514662 |
| rz.age | 3 | 5.56200 | 6.910605 | 5.079 | 5.822 | UNC4813186 |
| rz.age | 3 | 6.85700 | 7.094779 | 6.356 | 7.572 | UNC4836550 |
| rz.age | 3 | 7.69400 | 7.702394 | 7.603 | 10.190 | UNCHS008182 |
| rz.age | 3 | 10.81500 | 9.379513 | 10.282 | 10.970 | UNC4912039 |
| rz.age | 3 | 11.39100 | 9.379516 | 10.982 | 11.971 | UNC4924377 |
| rz.age | 3 | 12.29000 | 9.280016 | 12.254 | 12.989 | UNCHS008286 |
| rz.age | 3 | 13.56900 | 9.373639 | 13.052 | 15.109 | UNC4956618 |
| rz.age | 3 | 16.36100 | 9.286749 | 15.290 | 16.966 | UNC5006948 |
| rz.age | 3 | 18.74600 | 11.185850 | 17.482 | 19.077 | UNC5083168 |
| rz.age | 3 | 20.70900 | 10.948366 | 20.613 | 20.774 | UNCJPD001246 |
| rz.age | 3 | 21.11100 | 10.438840 | 20.784 | 21.549 | UNCHS008595 |
| rz.age | 3 | 21.68733 | 10.121892 | 21.580 | 22.187 | UNCHS008613 |
| rz.age | 3 | 23.13700 | 9.607665 | 22.489 | 23.199 | UNC5262472 |
| rz.age | 3 | 25.66500 | 9.687412 | 23.380 | 27.894 | UNC5300878 |
| rz.age | 3 | 29.00200 | 9.483694 | 28.262 | 29.026 | UNCHS008815 |
| rz.age | 3 | 29.06600 | 9.483566 | 29.027 | 30.167 | UNC5360055 |
| rz.age | 3 | 31.80500 | 8.855733 | 30.203 | 32.529 | UNC5454839 |
| rz.age | 3 | 32.82300 | 7.920116 | 32.607 | 33.376 | ICR1338 |
| rz.age | 3 | 33.71800 | 8.005633 | 33.391 | 33.741 | ICR2047 |
| rz.age | 3 | 34.83500 | 8.078228 | 33.825 | 35.168 | UNCJPD001341 |
| rz.age | 3 | 36.06800 | 7.983329 | 36.023 | 37.404 | UNC5645844 |
| rz.age | 3 | 37.53200 | 6.318534 | 37.408 | 37.640 | UNC5679155 |
| rz.age | 3 | 39.17200 | 6.580030 | 37.874 | 39.804 | JAX00529707 |
| rz.age | 3 | 40.06700 | 6.242010 | 40.024 | 40.390 | UNCHS009383 |
| rz.age | 3 | 42.34600 | 5.685666 | 40.519 | 44.496 | UNC5808246 |
| rz.age | 3 | 46.13600 | 4.089132 | 45.825 | 46.481 | UNCHS009693 |
| rz.age | 3 | 46.74000 | 4.175587 | 46.556 | 46.882 | sanger2623 |
| rz.age | 3 | 48.38000 | 4.299015 | 47.611 | 51.223 | UNC5945286 |
| rz.age | 3 | 52.87800 | 3.624985 | 51.460 | 56.964 | JAX00112499 |
| rz.age | 3 | 58.27200 | 3.449250 | 58.043 | 63.387 | UNC6227106 |
if(nrow(peaks) < 50 & nrow(peaks) > 0){
for(i in unique(peaks$chr)){
#for (i in 1:nrow(peaks)){
#plot_lod_chr(out,gm$gmap, peaks$chr[i])
plot_lod_chr(out,gm$gmap, i)
}
} else {
print(paste0("There are too many peaks (",nrow(peaks)," peaks) that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}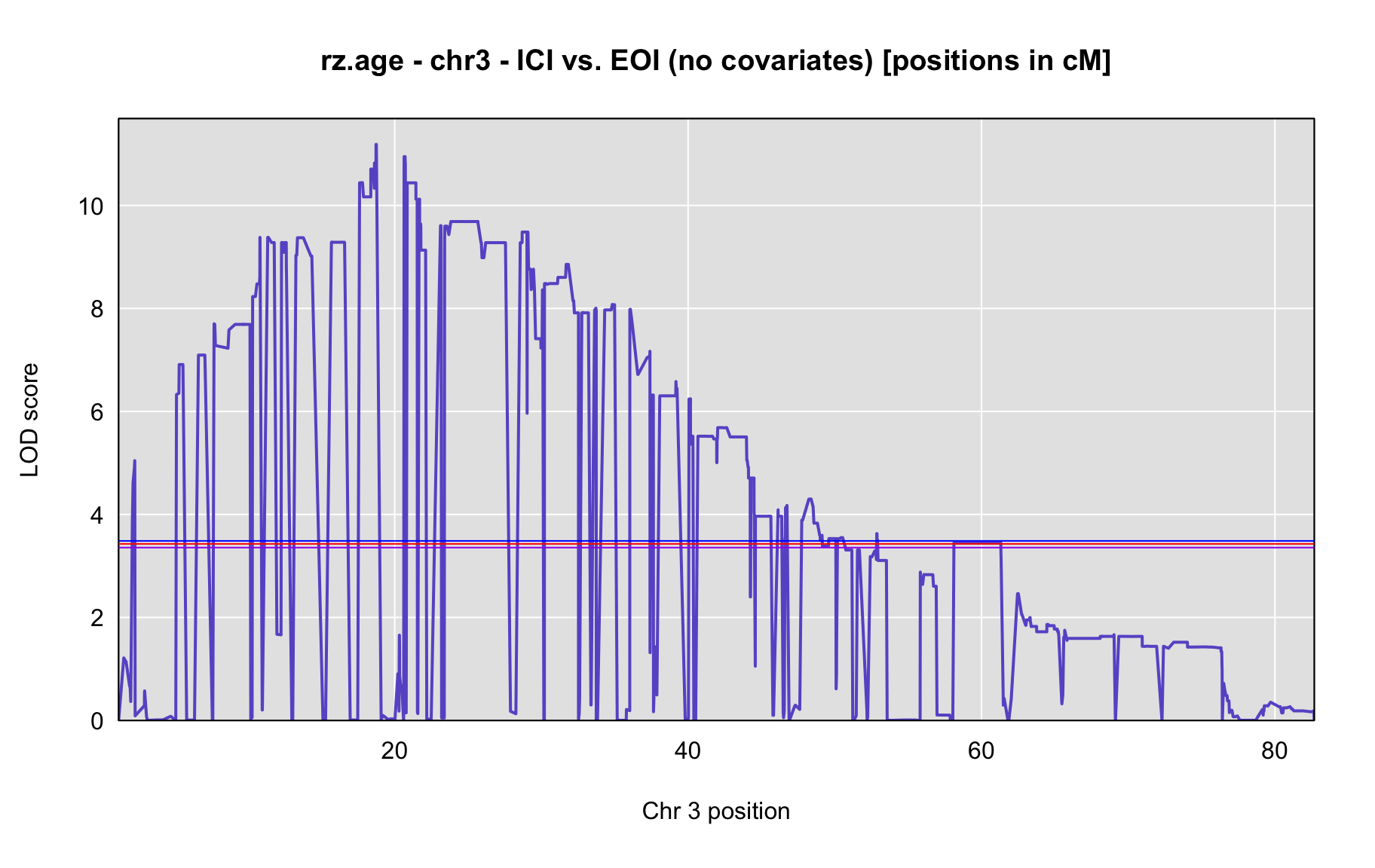
Megabase (MB)
peaks_mba<-find_peaks(out, gm$pmap, threshold=summary(operm,alpha=0.05)$A, thresholdX = summary(operm,alpha=0.05)$X, peakdrop=3, drop=1.5)
if(nrow(peaks) > 0){
peaks_mba$marker <- find_marker(gm$pmap, chr=peaks_mba$chr,pos=peaks_mba$pos)
names(peaks_mba)[2] <- c("phenotype")
peaks_mba <- peaks_mba[-1]
rownames(peaks_mba) <- NULL
print(kable(peaks_mba, escape = F, align = c("cccccccc"), "html")
%>% kable_styling("striped", full_width = T)%>%
column_spec(1, bold=TRUE)
)
plot_lod_chr_mb<-function(out,map,chrom){
for (i in 1:dim(out)[2]){
#png(filename=paste0("/Users/chenm/Documents/qtl/Jai/",colnames(out)[i], "_lod.png"))
#par(mar=c(5.1, 6.1, 1.1, 1.1))
ymx <- maxlod(out) # overall maximum LOD score
plot(out, map, chr = chrom, lodcolumn=i, col="slateblue", ylim=c(0, ymx+0.5))
#legend("topright", lwd=2, colnames(out)[i], bg="gray90")
title(main = paste0(colnames(out)[i], " - chr", chrom, " - ICI vs. EOI (no covariates) [positions in cM]"))
add_threshold(map, summary(operm,alpha=0.1), col = 'purple')
add_threshold(map, summary(operm, alpha=0.05), col = 'red')
add_threshold(map, summary(operm, alpha=0.01), col = 'blue')
#for (j in 1: dim(summary_table)[1]){
# abline(h=summary_table[j, i],col="red")
# text(x=400, y =summary_table[j, i]+0.12, labels = paste("p=", row.names(summary_table)[j]))
#}
#dev.off()
}
}
} else {
print(paste0("There are no peaks that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}| phenotype | chr | pos | lod | ci_lo | ci_hi | marker |
|---|---|---|---|---|---|---|
| rz.age | 3 | 9.258869 | 5.043745 | 7.129186 | 9.402052 | JAX00514662 |
| rz.age | 3 | 19.486223 | 6.910605 | 18.314568 | 19.729641 | UNC4813186 |
| rz.age | 3 | 21.048602 | 7.094779 | 20.305499 | 22.035726 | UNC4836550 |
| rz.age | 3 | 22.709183 | 7.702394 | 22.177798 | 25.286239 | UNCHS008182 |
| rz.age | 3 | 27.036217 | 9.379513 | 25.426933 | 27.557911 | UNCHS008242 |
| rz.age | 3 | 28.019689 | 9.379516 | 27.583922 | 28.411523 | UNC4924377 |
| rz.age | 3 | 28.700046 | 9.280016 | 28.647890 | 30.203812 | UNCHS008286 |
| rz.age | 3 | 30.369809 | 9.373639 | 30.221767 | 31.081568 | UNC4956618 |
| rz.age | 3 | 33.900624 | 9.286749 | 31.479710 | 34.955875 | UNC5006948 |
| rz.age | 3 | 38.948083 | 11.185850 | 36.524146 | 39.720506 | UNC5083168 |
| rz.age | 3 | 46.369552 | 10.948366 | 46.093688 | 46.773347 | UNCJPD001246 |
| rz.age | 3 | 47.991891 | 10.438840 | 47.267935 | 48.887661 | UNCHS008595 |
| rz.age | 3 | 49.609061 | 10.121892 | 48.950852 | 51.245920 | UNCHS008613 |
| rz.age | 3 | 52.227835 | 9.607665 | 51.504201 | 52.288687 | UNC5262472 |
| rz.age | 3 | 54.787025 | 9.687412 | 52.466515 | 57.567476 | UNC5300878 |
| rz.age | 3 | 59.186472 | 9.483694 | 58.021842 | 59.322589 | UNCHS008815 |
| rz.age | 3 | 59.677691 | 9.483566 | 59.333869 | 64.965863 | UNC5360055 |
| rz.age | 3 | 68.536022 | 8.855733 | 65.480179 | 70.648825 | UNC5454839 |
| rz.age | 3 | 73.269966 | 7.920116 | 72.338635 | 73.730105 | ICR1338 |
| rz.age | 3 | 75.230274 | 8.005633 | 73.742144 | 75.518981 | ICR5347 |
| rz.age | 3 | 78.195206 | 8.078228 | 75.789940 | 79.931821 | UNCJPD001341 |
| rz.age | 3 | 81.949482 | 7.983329 | 81.864954 | 83.879902 | UNC5645844 |
| rz.age | 3 | 84.296456 | 6.318534 | 83.892546 | 84.573833 | UNC5679155 |
| rz.age | 3 | 89.537778 | 6.580030 | 86.634703 | 90.662415 | JAX00529707r |
| rz.age | 3 | 91.300118 | 6.242010 | 90.812476 | 93.632371 | UNCHS009383 |
| rz.age | 3 | 97.884162 | 5.685666 | 93.882933 | 102.026335 | UNC5808246 |
| rz.age | 3 | 105.105036 | 4.089132 | 104.826451 | 106.170686 | UNCHS009693 |
| rz.age | 3 | 107.470377 | 4.175587 | 106.942181 | 108.045878 | sanger2623q |
| rz.age | 3 | 109.677215 | 4.299015 | 108.888517 | 117.883783 | UNC5945286 |
| rz.age | 3 | 121.181495 | 3.624985 | 118.208381 | 128.488205 | JAX00112499 |
| rz.age | 3 | 129.551327 | 3.449250 | 129.469583 | 136.918958 | UNC6227106 |
if(nrow(peaks_mba) < 50 & nrow(peaks_mba) > 0){
for(i in unique(peaks_mba$chr)){
#for (i in 1:nrow(peaks_mba)){
#plot_lod_chr_mb(out,gm$pmap, peaks_mba$chr[i])
plot_lod_chr_mb(out,gm$pmap,i)
}
} else {
print(paste0("There are too many peaks (",nrow(peaks_mba)," peaks) that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}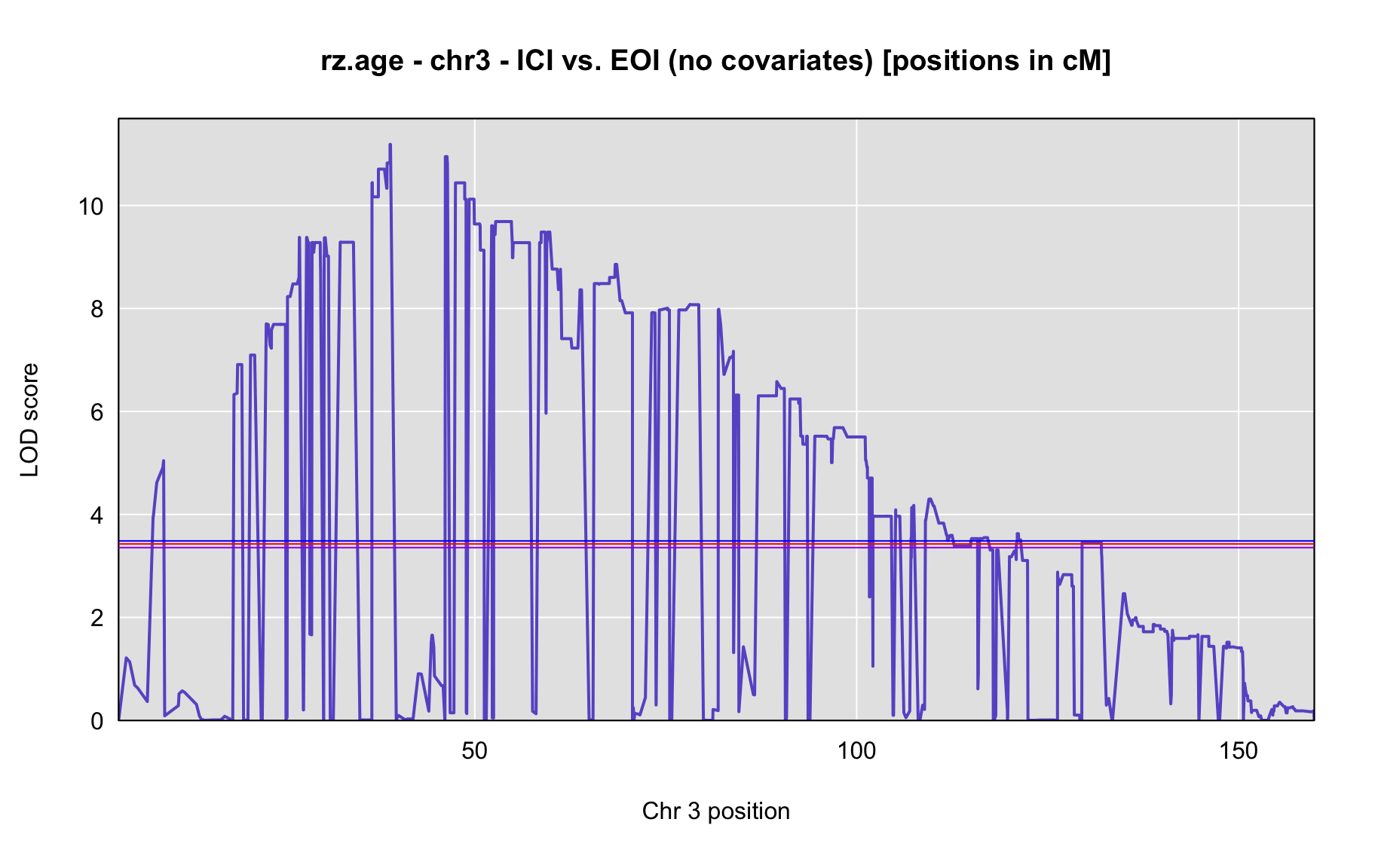
QTL effects
For each peak LOD location we give a list of gene
if(nrow(peaks) < 50 & nrow(peaks) > 0){
for (i in 1:nrow(peaks)){
#for (i in 1:1){
#Plot 1
g <- maxmarg(pr.qc, gm$gmap, chr=peaks$chr[i], pos=peaks$pos[i], return_char=TRUE)
#png(filename=paste0("/Users/chenm/Documents/qtl/Jai/","qtl_effect_", i, ".png"))
#par(mar=c(4.1, 4.1, 1.5, 0.6))
plot_pxg(g, covars[,peaks$phenotype[i]], ylab=peaks$phenotype[i], sort=FALSE)
title(main = paste0("chr: ", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$phenotype[i]," )"), line=0.2)
##dev.off()
chr = peaks$chr[i]
# Plot 2
pr_sub <- pull_genoprobint(pr.qc, gm$gmap, chr, c(peaks$ci_lo[i], peaks$ci_hi[i]))
#coeff <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]], addcovar = addcovar)
#coeff <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]], Xcovar=Xcovar)
#coeff <- scan1coef(pr.qc[,chr], gm$covar[peaks$lodcolumn[i]], model="binary")
#coeff_sub <- scan1coef(pr_sub[,chr], gm$covar[peaks$lodcolumn[i]], model="binary")
blup <- scan1blup(pr.qc[,chr], covars[peaks$phenotype[i]])
blup_sub <- scan1blup(pr_sub[,chr], covars[peaks$phenotype[i]])
write.csv(as.data.frame(blup_sub), paste0("data/ici.vs.eoi_",peaks$phenotype[i],"-no.covariates_blup_sub_chr-",chr,"_peak.marker-",peaks$marker[i],"_lod.drop-1.5_5.batches_52.csv"), quote=F)
#plot_coef(coeff,
# gm$gmap, columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$lodcolumn[i]," [scan1coeff; positions in cM] )")
# )
#plot_coef(coeff_sub,
# gm$gmap, columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$lodcolumn[i],"; 1.5 LOD drop interval [scan1coeff; positions in cM] ) ")
# )
plot_coef(blup,
gm$gmap, columns=1:2,
bgcolor="gray95", legend="bottomleft",
main = paste0("chr: ", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$phenotype[i]," [scan1blup; positions in cM] )")
)
plot_coef(blup_sub,
gm$gmap, columns=1:2,
bgcolor="gray95", legend="bottomleft",
main = paste0("chr: ", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$phenotype[i],"; 1.5 LOD drop interval [scan1blup; positions in cM] )")
)
# Plot 3
#c2effB <- scan1coef(pr.qc[,chr], gm$covar[peaks$lodcolumn[i]], model="binary", contrasts=cbind(a=c(-1, 0), d=c(0, -1)))
#c2effBb <- scan1blup(pr.qc[,chr], gm$covar[peaks$lodcolumn[i]], contrasts=cbind(a=c(-1, 0), d=c(0, -1)))
##c2effB <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]], addcovar = addcovar, contrasts=cbind(mu=c(1,1,1), a=c(-1, 0, 1), d=c(0, 1, 0)))
##c2effB <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]],Xcovar=Xcovar, contrasts=cbind(mu=c(1,1,1), a=c(-1, 0, 1), d=c(0, 1, 0)))
#plot(c2effB, gm$gmap[chr], columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i], "pos", peaks$pos[i], "(",peaks$lodcolumn[i],")")
# )
#plot(c2effBb, gm$gmap[chr], columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i], "pos", peaks$pos[i], "(",peaks$lodcolumn[i],")")
# )
##last_coef <- unclass(c2effB)[nrow(c2effB),2:3] # last two coefficients
##for(t in seq(along=last_coef))
## axis(side=4, at=last_coef[t], names(last_coef)[t], tick=FALSE)
#Table 1
chr = peaks_mba$chr[i]
start=as.numeric(peaks_mba$ci_lo[i])
end=as.numeric(peaks_mba$ci_hi[i])
genesgss = query_genes(chr, start, end)
write.csv(genesgss, file=paste0("data/ici.vs.eoi_",peaks$phenotype[i],"-no.covariates_genes_chr-",chr,"_peak.marker-",peaks$marker[i],"_lod.drop-1.5_5.batches_52.csv"), quote=F)
rownames(genesgss) <- NULL
genesgss$strand_old = genesgss$strand
genesgss$strand[genesgss$strand=="+"] <- "positive"
genesgss$strand[genesgss$strand=="-"] <- "negative"
#genesgss <-
#table <-
#genesgss[,c("chr","type","start","stop","strand","ID","Name","Dbxref","gene_id","mgi_type","description")] %>%
#kable(escape = F,align = c("ccccccccccc")) %>%
#kable_styling("striped", full_width = T) #%>%
#cat #%>%
#column_spec(1, bold=TRUE)
#
#print(kable(genesgss[,c("chr","type","start","stop","strand","ID","Name","Dbxref","gene_id","mgi_type","description")], escape = F,align = c("ccccccccccc")))
print(kable(genesgss[,c("chr","type","start","stop","strand","ID","Name","Dbxref","gene_id","mgi_type","description")], "html") %>% kable_styling("striped", full_width = T))
#table
}
} else {
print(paste0("There are too many peaks (",nrow(peaks_mba)," peaks) that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}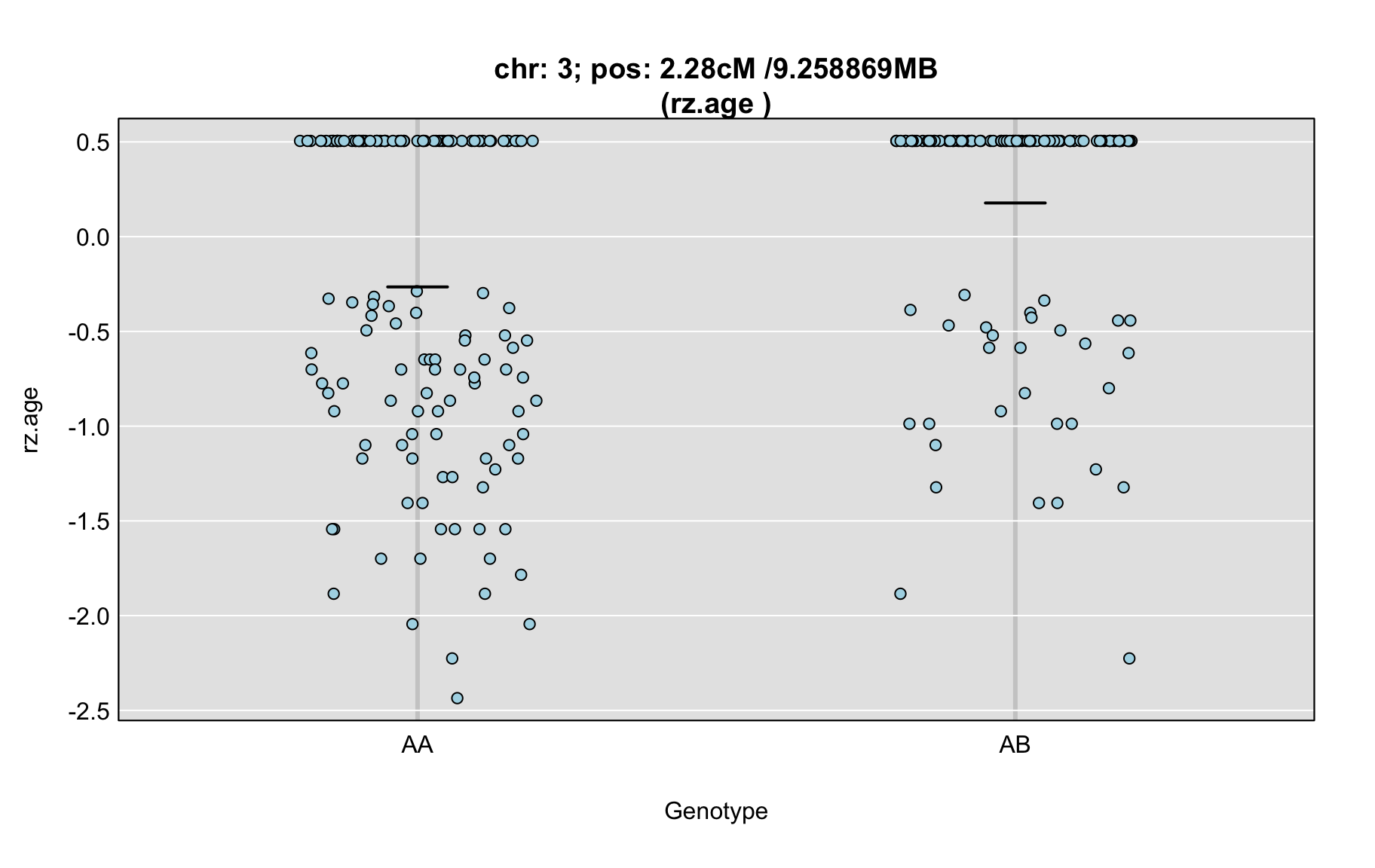
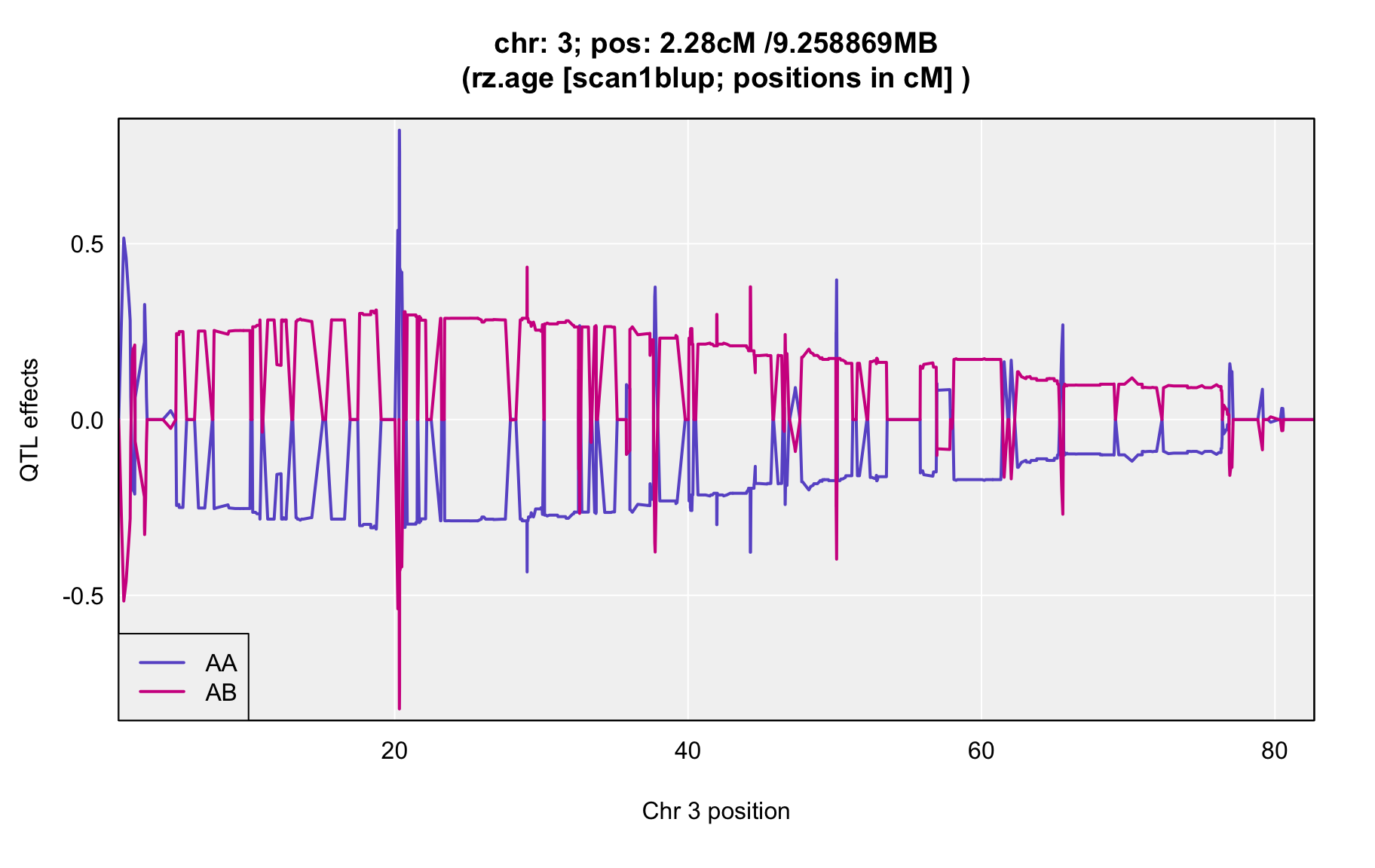
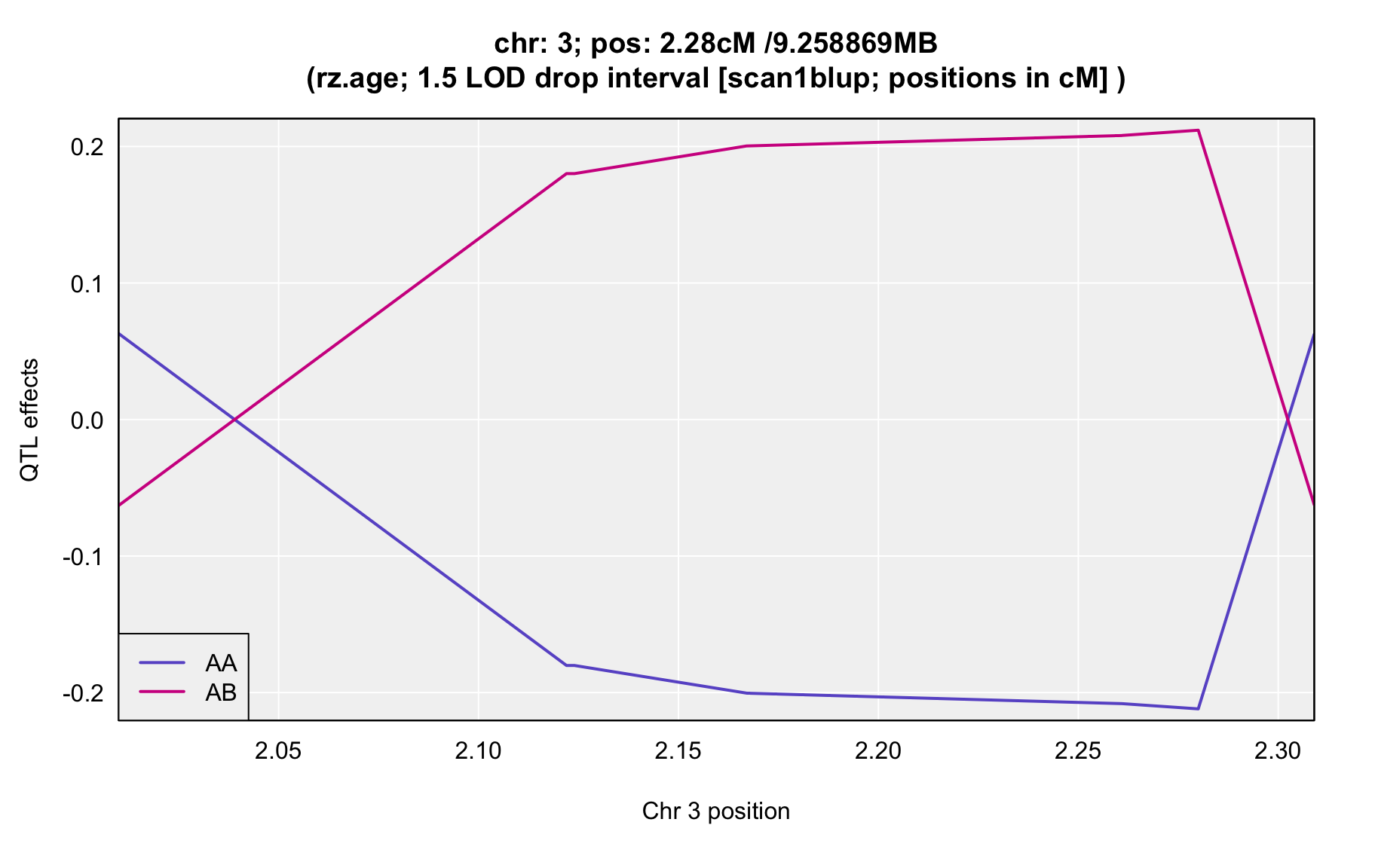
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | pseudogene | 7.192775 | 7.193210 | negative | MGI_C57BL6J_3647094 | Gm8823 | NCBI_Gene:667801,ENSEMBL:ENSMUSG00000104274 | MGI:3647094 | pseudogene | predicted gene 8823 |
| 3 | gene | 7.366604 | 7.445366 | positive | MGI_C57BL6J_104747 | Pkia | NCBI_Gene:18767,ENSEMBL:ENSMUSG00000027499 | MGI:104747 | protein coding gene | protein kinase inhibitor, alpha |
| 3 | gene | 7.408504 | 7.410589 | positive | MGI_C57BL6J_3027006 | C530038A18Rik | NA | NA | unclassified gene | RIKEN cDNA C530038A18 gene |
| 3 | pseudogene | 7.418340 | 7.418459 | negative | MGI_C57BL6J_5611146 | Gm37918 | ENSEMBL:ENSMUSG00000103274 | MGI:5611146 | pseudogene | predicted gene, 37918 |
| 3 | pseudogene | 7.454521 | 7.455539 | positive | MGI_C57BL6J_3647097 | Gm8826 | NCBI_Gene:667807,ENSEMBL:ENSMUSG00000102268 | MGI:3647097 | pseudogene | predicted gene 8826 |
| 3 | gene | 7.501904 | 7.502020 | negative | MGI_C57BL6J_5453369 | Gm23592 | ENSEMBL:ENSMUSG00000088484 | MGI:5453369 | snoRNA gene | predicted gene, 23592 |
| 3 | gene | 7.503415 | 7.553848 | positive | MGI_C57BL6J_1914556 | Zc2hc1a | NCBI_Gene:67306,ENSEMBL:ENSMUSG00000043542 | MGI:1914556 | protein coding gene | zinc finger, C2HC-type containing 1A |
| 3 | gene | 7.569994 | 7.613760 | negative | MGI_C57BL6J_96561 | Il7 | NCBI_Gene:16196,ENSEMBL:ENSMUSG00000040329 | MGI:96561 | protein coding gene | interleukin 7 |
| 3 | gene | 7.576267 | 7.690001 | positive | MGI_C57BL6J_4439609 | Gm16685 | NCBI_Gene:102634900,ENSEMBL:ENSMUSG00000097804 | MGI:4439609 | lncRNA gene | predicted gene, 16685 |
| 3 | pseudogene | 7.893335 | 7.893980 | positive | MGI_C57BL6J_5588906 | Gm29747 | NCBI_Gene:101055938,ENSEMBL:ENSMUSG00000103399 | MGI:5588906 | pseudogene | predicted gene, 29747 |
| 3 | gene | 7.925303 | 7.954889 | negative | MGI_C57BL6J_1922726 | 1700010I02Rik | NCBI_Gene:75476,ENSEMBL:ENSMUSG00000100010 | MGI:1922726 | lncRNA gene | RIKEN cDNA 1700010I02 gene |
| 3 | gene | 8.019669 | 8.022831 | negative | MGI_C57BL6J_3642332 | 5330432E05Rik | NCBI_Gene:383820 | MGI:3642332 | unclassified gene | RIKEN cDNA 5330432E05 gene |
| 3 | pseudogene | 8.214330 | 8.216183 | positive | MGI_C57BL6J_3646860 | Gm7103 | NCBI_Gene:633072,ENSEMBL:ENSMUSG00000103725 | MGI:3646860 | pseudogene | predicted gene 7103 |
| 3 | gene | 8.245438 | 8.245537 | negative | MGI_C57BL6J_3629711 | Gt(pU21)108Imeg | NA | NA | unclassified gene | gene trap 108%2c Institute of Molecular Embryology and Genetics |
| 3 | pseudogene | 8.327322 | 8.329907 | positive | MGI_C57BL6J_3645248 | Gm5841 | NCBI_Gene:545500,ENSEMBL:ENSMUSG00000103921 | MGI:3645248 | pseudogene | predicted gene 5841 |
| 3 | gene | 8.429158 | 8.429264 | positive | MGI_C57BL6J_5454391 | Gm24614 | ENSEMBL:ENSMUSG00000065203 | MGI:5454391 | snRNA gene | predicted gene, 24614 |
| 3 | pseudogene | 8.461387 | 8.463098 | negative | MGI_C57BL6J_3704094 | Gm6194 | NCBI_Gene:620966 | MGI:3704094 | pseudogene | predicted gene 6194 |
| 3 | pseudogene | 8.462228 | 8.463035 | negative | MGI_C57BL6J_1919464 | 1700003N22Rik | ENSEMBL:ENSMUSG00000102463 | MGI:1919464 | pseudogene | RIKEN cDNA 1700003N22 gene |
| 3 | gene | 8.494369 | 8.499902 | negative | MGI_C57BL6J_5624878 | Gm41993 | NCBI_Gene:105246751 | MGI:5624878 | lncRNA gene | predicted gene, 41993 |
| 3 | gene | 8.509360 | 8.561606 | positive | MGI_C57BL6J_98241 | Stmn2 | NCBI_Gene:20257,ENSEMBL:ENSMUSG00000027500 | MGI:98241 | protein coding gene | stathmin-like 2 |
| 3 | pseudogene | 8.615074 | 8.616522 | negative | MGI_C57BL6J_5011007 | Gm18822 | NCBI_Gene:100417782,ENSEMBL:ENSMUSG00000098047 | MGI:5011007 | pseudogene | predicted gene, 18822 |
| 3 | gene | 8.628051 | 8.632454 | positive | MGI_C57BL6J_5591655 | Gm32496 | NCBI_Gene:102635065,ENSEMBL:ENSMUSG00000104102 | MGI:5591655 | lncRNA gene | predicted gene, 32496 |
| 3 | gene | 8.663359 | 8.667256 | negative | MGI_C57BL6J_1341800 | Hey1 | NCBI_Gene:15213,ENSEMBL:ENSMUSG00000040289 | MGI:1341800 | protein coding gene | hairy/enhancer-of-split related with YRPW motif 1 |
| 3 | pseudogene | 8.673410 | 8.673722 | positive | MGI_C57BL6J_5611235 | Gm38007 | ENSEMBL:ENSMUSG00000103814 | MGI:5611235 | pseudogene | predicted gene, 38007 |
| 3 | gene | 8.679359 | 8.679477 | negative | MGI_C57BL6J_5453447 | Gm23670 | ENSEMBL:ENSMUSG00000088094 | MGI:5453447 | snoRNA gene | predicted gene, 23670 |
| 3 | gene | 8.679438 | 8.693098 | negative | MGI_C57BL6J_5591720 | Gm32561 | NCBI_Gene:102635149 | MGI:5591720 | lncRNA gene | predicted gene, 32561 |
| 3 | gene | 8.732188 | 8.736639 | negative | MGI_C57BL6J_2686488 | C230057A21Rik | NCBI_Gene:102635252,ENSEMBL:ENSMUSG00000103346 | MGI:2686488 | lncRNA gene | RIKEN cDNA C230057A21 gene |
| 3 | gene | 8.743677 | 8.746824 | positive | MGI_C57BL6J_1923855 | 1700042O13Rik | NA | NA | lncRNA gene | RIKEN cDNA 1700042O13 gene |
| 3 | gene | 8.743682 | 8.746824 | negative | MGI_C57BL6J_5610875 | Gm37647 | ENSEMBL:ENSMUSG00000103256 | MGI:5610875 | unclassified gene | predicted gene, 37647 |
| 3 | gene | 8.802146 | 8.923918 | negative | MGI_C57BL6J_1913480 | Mrps28 | NCBI_Gene:66230,ENSEMBL:ENSMUSG00000040269 | MGI:1913480 | protein coding gene | mitochondrial ribosomal protein S28 |
| 3 | gene | 8.866119 | 8.881665 | positive | MGI_C57BL6J_5624880 | Gm41995 | NCBI_Gene:105246753 | MGI:5624880 | lncRNA gene | predicted gene, 41995 |
| 3 | pseudogene | 8.881199 | 8.881643 | positive | MGI_C57BL6J_3705388 | Gm15467 | ENSEMBL:ENSMUSG00000081594 | MGI:3705388 | pseudogene | predicted gene 15467 |
| 3 | pseudogene | 8.881990 | 8.882945 | negative | MGI_C57BL6J_3705491 | Gm15466 | NCBI_Gene:100504173,ENSEMBL:ENSMUSG00000081657 | MGI:3705491 | pseudogene | predicted gene 15466 |
| 3 | gene | 8.925590 | 8.928038 | positive | MGI_C57BL6J_1918835 | 9130403I23Rik | NA | NA | unclassified gene | RIKEN cDNA 9130403I23 gene |
| 3 | gene | 8.925593 | 9.004771 | negative | MGI_C57BL6J_107749 | Tpd52 | NCBI_Gene:21985,ENSEMBL:ENSMUSG00000027506 | MGI:107749 | protein coding gene | tumor protein D52 |
| 3 | gene | 9.027570 | 9.029901 | positive | MGI_C57BL6J_5826469 | Gm46832 | NCBI_Gene:108168912 | MGI:5826469 | lncRNA gene | predicted gene, 46832 |
| 3 | pseudogene | 9.047171 | 9.047492 | negative | MGI_C57BL6J_5610844 | Gm37616 | ENSEMBL:ENSMUSG00000103461 | MGI:5610844 | pseudogene | predicted gene, 37616 |
| 3 | gene | 9.061964 | 9.071076 | positive | MGI_C57BL6J_1922399 | 4930539M17Rik | NCBI_Gene:75149,ENSEMBL:ENSMUSG00000103116 | MGI:1922399 | lncRNA gene | RIKEN cDNA 4930539M17 gene |
| 3 | pseudogene | 9.072767 | 9.073211 | positive | MGI_C57BL6J_3648010 | Gm8860 | ENSEMBL:ENSMUSG00000103011 | MGI:3648010 | pseudogene | predicted gene 8860 |
| 3 | gene | 9.157271 | 9.174711 | negative | MGI_C57BL6J_5624881 | Gm41996 | NCBI_Gene:105246754 | MGI:5624881 | lncRNA gene | predicted gene, 41996 |
| 3 | gene | 9.177692 | 9.182437 | negative | MGI_C57BL6J_5592177 | Gm33018 | NCBI_Gene:102635764 | MGI:5592177 | lncRNA gene | predicted gene, 33018 |
| 3 | gene | 9.250567 | 9.285333 | positive | MGI_C57BL6J_2139883 | Zbtb10 | NCBI_Gene:229055,ENSEMBL:ENSMUSG00000069114 | MGI:2139883 | protein coding gene | zinc finger and BTB domain containing 10 |
| 3 | gene | 9.302688 | 9.343835 | negative | MGI_C57BL6J_5624882 | Gm41997 | NCBI_Gene:105246755 | MGI:5624882 | lncRNA gene | predicted gene, 41997 |
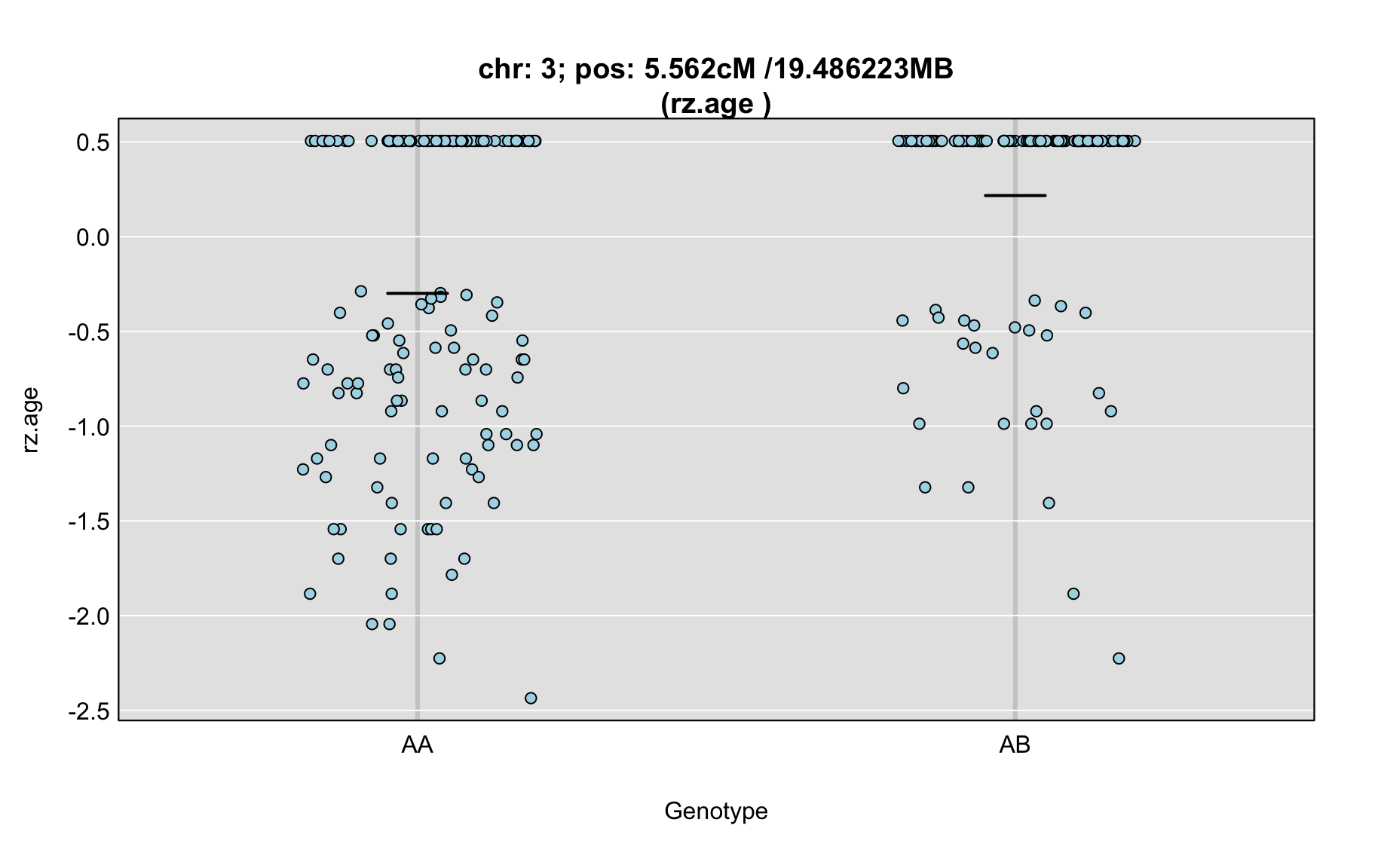
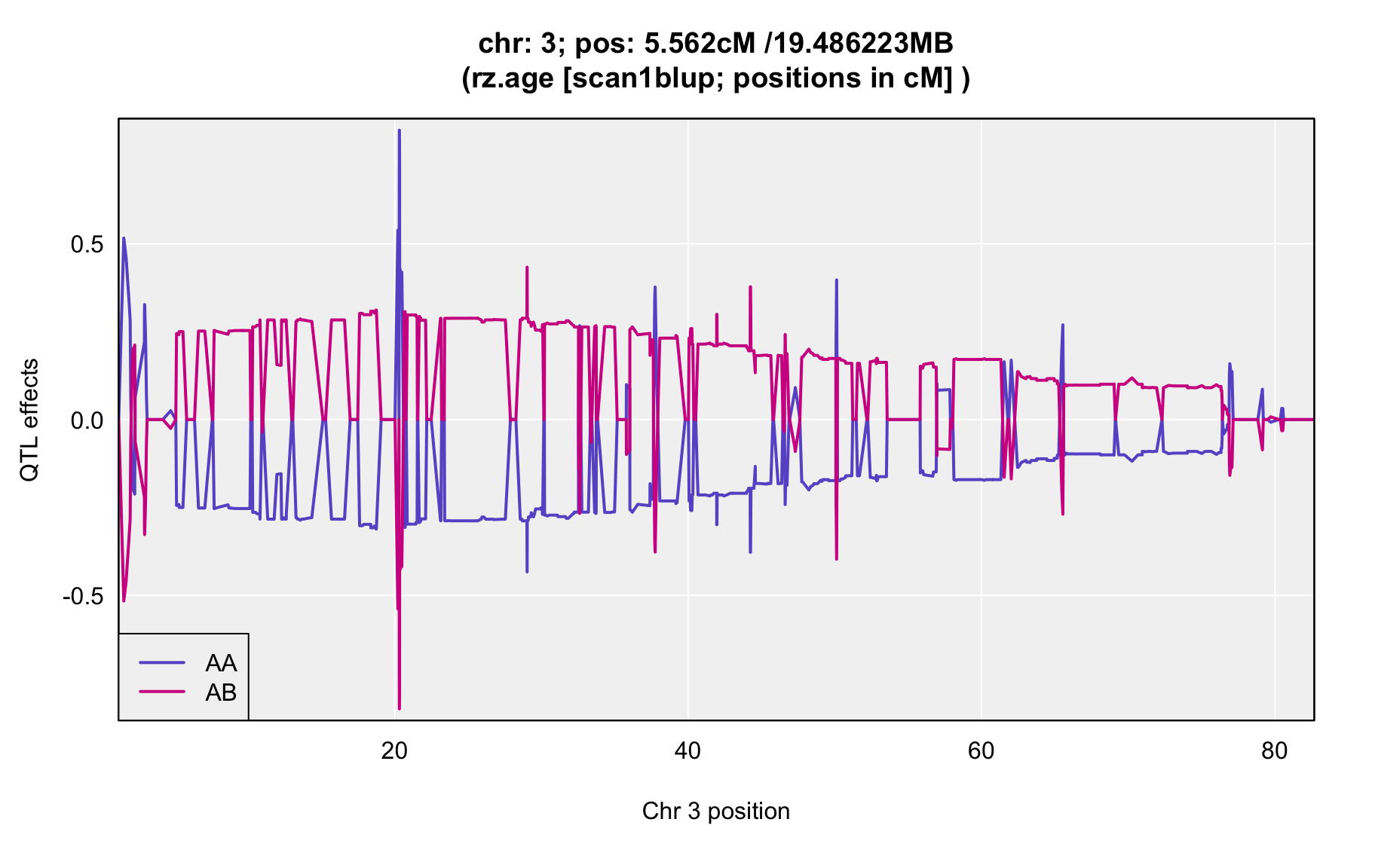
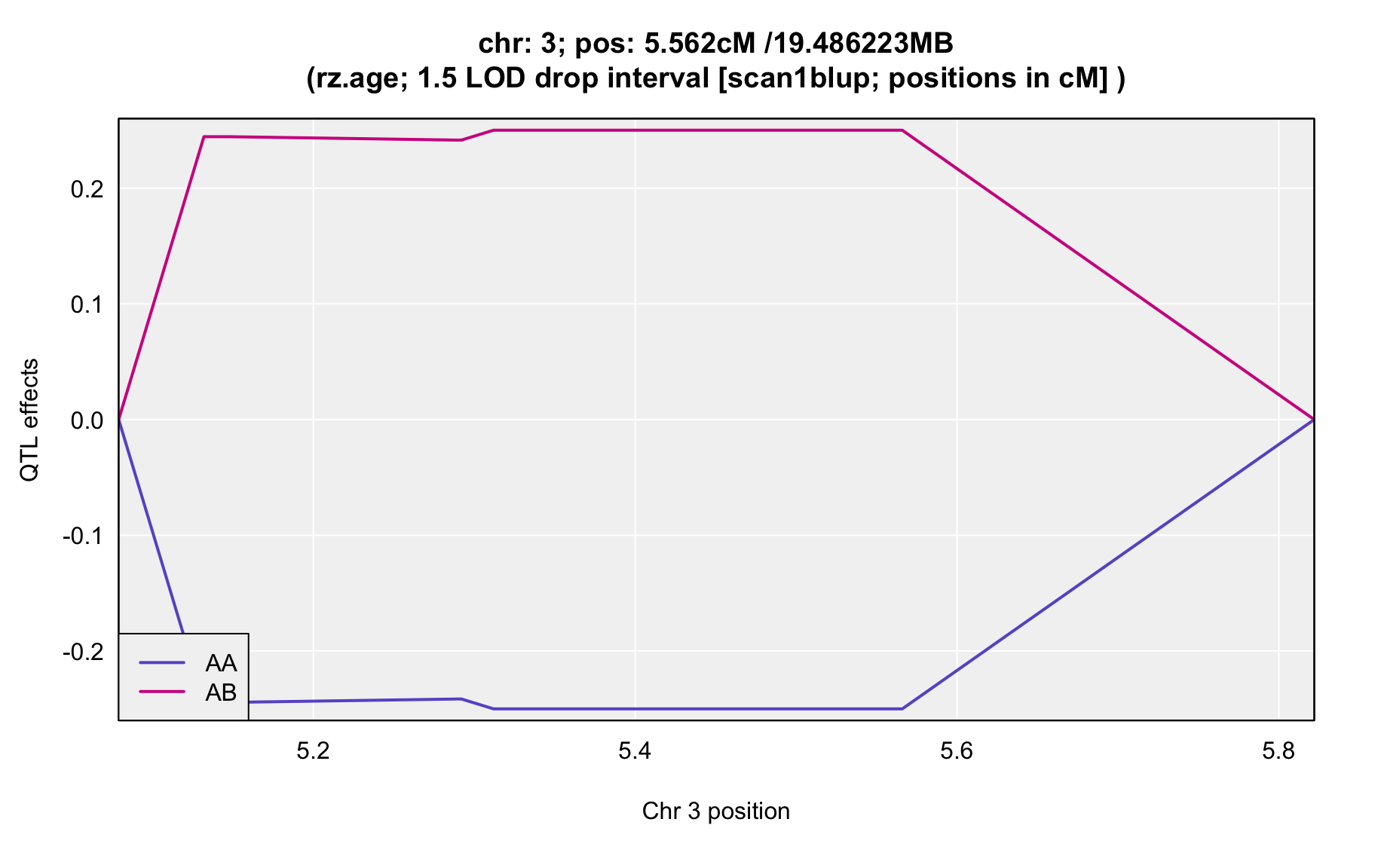
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 18.34167 | 18.46134 | negative | MGI_C57BL6J_5589826 | Gm30667 | NCBI_Gene:102632645,ENSEMBL:ENSMUSG00000102886 | MGI:5589826 | lncRNA gene | predicted gene, 30667 |
| 3 | gene | 18.46829 | 18.48048 | negative | MGI_C57BL6J_1921209 | 4930433B08Rik | NCBI_Gene:73959,ENSEMBL:ENSMUSG00000102699 | MGI:1921209 | lncRNA gene | RIKEN cDNA 4930433B08 gene |
| 3 | pseudogene | 18.49462 | 18.53429 | positive | MGI_C57BL6J_3644758 | Gm6369 | NCBI_Gene:622921,ENSEMBL:ENSMUSG00000040217 | MGI:3644758 | pseudogene | predicted gene 6369 |
| 3 | pseudogene | 18.56449 | 18.56614 | positive | MGI_C57BL6J_97558 | Pgk1-ps3 | NCBI_Gene:100416868,ENSEMBL:ENSMUSG00000105280 | MGI:97558 | pseudogene | phosphoglycerate kinase 1, pseudogene 3 |
| 3 | gene | 18.60947 | 18.61738 | negative | MGI_C57BL6J_5589882 | Gm30723 | NCBI_Gene:102632727 | MGI:5589882 | lncRNA gene | predicted gene, 30723 |
| 3 | pseudogene | 18.60981 | 18.61083 | negative | MGI_C57BL6J_5610545 | Gm37317 | ENSEMBL:ENSMUSG00000104091 | MGI:5610545 | pseudogene | predicted gene, 37317 |
| 3 | gene | 18.61955 | 18.64309 | negative | MGI_C57BL6J_5589948 | Gm30789 | NCBI_Gene:102632812 | MGI:5589948 | lncRNA gene | predicted gene, 30789 |
| 3 | gene | 18.62216 | 18.62614 | positive | MGI_C57BL6J_5663081 | Gm42944 | ENSEMBL:ENSMUSG00000104918 | MGI:5663081 | unclassified gene | predicted gene 42944 |
| 3 | pseudogene | 18.75308 | 18.75340 | positive | MGI_C57BL6J_5589500 | Gm30341 | NCBI_Gene:102632201,ENSEMBL:ENSMUSG00000103264 | MGI:5589500 | pseudogene | predicted gene, 30341 |
| 3 | gene | 19.13140 | 19.16309 | negative | MGI_C57BL6J_1921502 | Armc1 | NCBI_Gene:74252,ENSEMBL:ENSMUSG00000027599 | MGI:1921502 | protein coding gene | armadillo repeat containing 1 |
| 3 | gene | 19.14319 | 19.14390 | negative | MGI_C57BL6J_5611207 | Gm37979 | ENSEMBL:ENSMUSG00000103135 | MGI:5611207 | unclassified gene | predicted gene, 37979 |
| 3 | gene | 19.18731 | 19.22082 | positive | MGI_C57BL6J_1914722 | Mtfr1 | NCBI_Gene:67472,ENSEMBL:ENSMUSG00000027601 | MGI:1914722 | protein coding gene | mitochondrial fission regulator 1 |
| 3 | gene | 19.22311 | 19.31132 | negative | MGI_C57BL6J_1202402 | Pde7a | NCBI_Gene:18583,ENSEMBL:ENSMUSG00000069094 | MGI:1202402 | protein coding gene | phosphodiesterase 7A |
| 3 | gene | 19.22311 | 19.22369 | negative | MGI_C57BL6J_1921035 | 4930431H11Rik | NA | NA | unclassified gene | RIKEN cDNA 4930431H11 gene |
| 3 | gene | 19.22778 | 19.24099 | positive | MGI_C57BL6J_3801946 | Gm16093 | ENSEMBL:ENSMUSG00000087108 | MGI:3801946 | lncRNA gene | predicted gene 16093 |
| 3 | gene | 19.31166 | 19.31374 | positive | MGI_C57BL6J_1920375 | 3110037C07Rik | NA | NA | unclassified gene | RIKEN cDNA 3110037C07 gene |
| 3 | gene | 19.33077 | 19.33108 | positive | MGI_C57BL6J_5452059 | Gm22282 | ENSEMBL:ENSMUSG00000089149 | MGI:5452059 | unclassified non-coding RNA gene | predicted gene, 22282 |
| 3 | gene | 19.37418 | 19.38763 | positive | MGI_C57BL6J_5625073 | Gm42188 | NCBI_Gene:105247004 | MGI:5625073 | lncRNA gene | predicted gene, 42188 |
| 3 | gene | 19.50859 | 19.61086 | positive | MGI_C57BL6J_1913576 | Dnajc5b | NCBI_Gene:66326,ENSEMBL:ENSMUSG00000027606 | MGI:1913576 | protein coding gene | DnaJ heat shock protein family (Hsp40) member C5 beta |
| 3 | gene | 19.56025 | 19.56042 | positive | MGI_C57BL6J_5453107 | Gm23330 | ENSEMBL:ENSMUSG00000096211 | MGI:5453107 | snRNA gene | predicted gene, 23330 |
| 3 | pseudogene | 19.58479 | 19.58554 | positive | MGI_C57BL6J_3646012 | Gm6141 | NCBI_Gene:620290,ENSEMBL:ENSMUSG00000084339 | MGI:3646012 | pseudogene | predicted pseudogene 6141 |
| 3 | gene | 19.61037 | 19.62948 | negative | MGI_C57BL6J_1920674 | 1700064H15Rik | NCBI_Gene:102633032,ENSEMBL:ENSMUSG00000082766 | MGI:1920674 | protein coding gene | RIKEN cDNA 1700064H15 gene |
| 3 | gene | 19.62796 | 19.62803 | negative | MGI_C57BL6J_4413716 | n-TRacg3 | NCBI_Gene:102467637 | MGI:4413716 | tRNA gene | nuclear encoded tRNA arginine 3 (anticodon ACG) |
| 3 | gene | 19.62835 | 19.62845 | positive | MGI_C57BL6J_4414074 | n-TYgta1 | NCBI_Gene:102467442 | MGI:4414074 | tRNA gene | nuclear encoded tRNA tyrosine 1 (anticodon GTA) |
| 3 | gene | 19.62878 | 19.62887 | positive | MGI_C57BL6J_4414075 | n-TYgta2 | NCBI_Gene:102467395 | MGI:4414075 | tRNA gene | nuclear encoded tRNA tyrosine 2 (anticodon GTA) |
| 3 | gene | 19.62899 | 19.62907 | positive | MGI_C57BL6J_4413687 | n-TAagc14 | NCBI_Gene:102467233 | MGI:4413687 | tRNA gene | nuclear encoded tRNA alanine 14 (anticodon AGC) |
| 3 | gene | 19.64434 | 19.69242 | positive | MGI_C57BL6J_3036269 | Trim55 | NCBI_Gene:381485,ENSEMBL:ENSMUSG00000060913 | MGI:3036269 | protein coding gene | tripartite motif-containing 55 |
| 3 | gene | 19.68597 | 19.68657 | positive | MGI_C57BL6J_3642300 | BC002189 | ENSEMBL:ENSMUSG00000104465 | MGI:3642300 | lncRNA gene | cDNA sequence BC002189 |
| 3 | gene | 19.69340 | 19.69540 | negative | MGI_C57BL6J_88496 | Crh | NCBI_Gene:12918,ENSEMBL:ENSMUSG00000049796 | MGI:88496 | protein coding gene | corticotropin releasing hormone |
| 3 | gene | 19.69449 | 19.70735 | positive | MGI_C57BL6J_5625074 | Gm42189 | NCBI_Gene:105247005 | MGI:5625074 | lncRNA gene | predicted gene, 42189 |
| 3 | pseudogene | 19.70825 | 19.70932 | negative | MGI_C57BL6J_5011049 | Gm18864 | NCBI_Gene:100417853,ENSEMBL:ENSMUSG00000102261 | MGI:5011049 | pseudogene | predicted gene, 18864 |
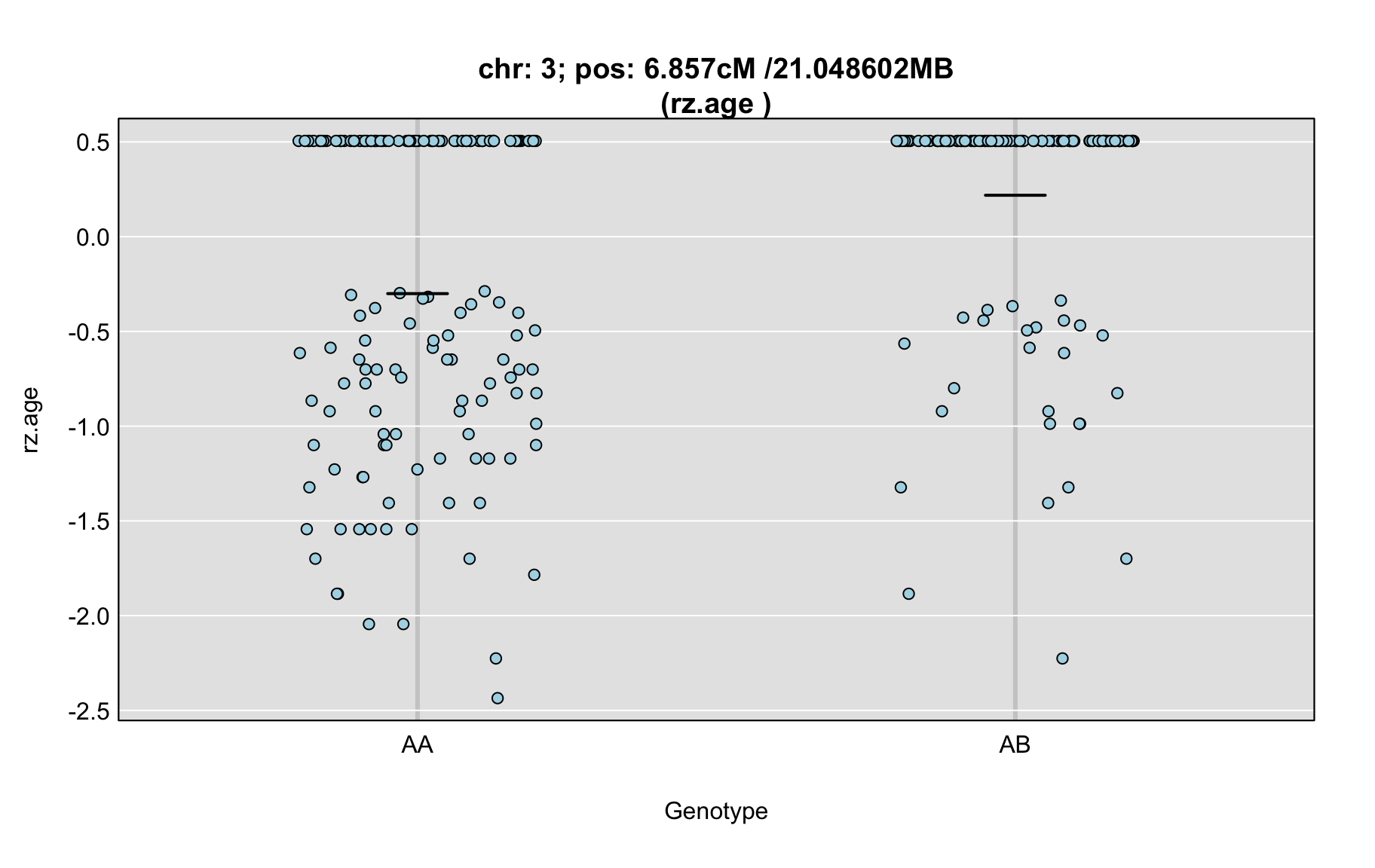
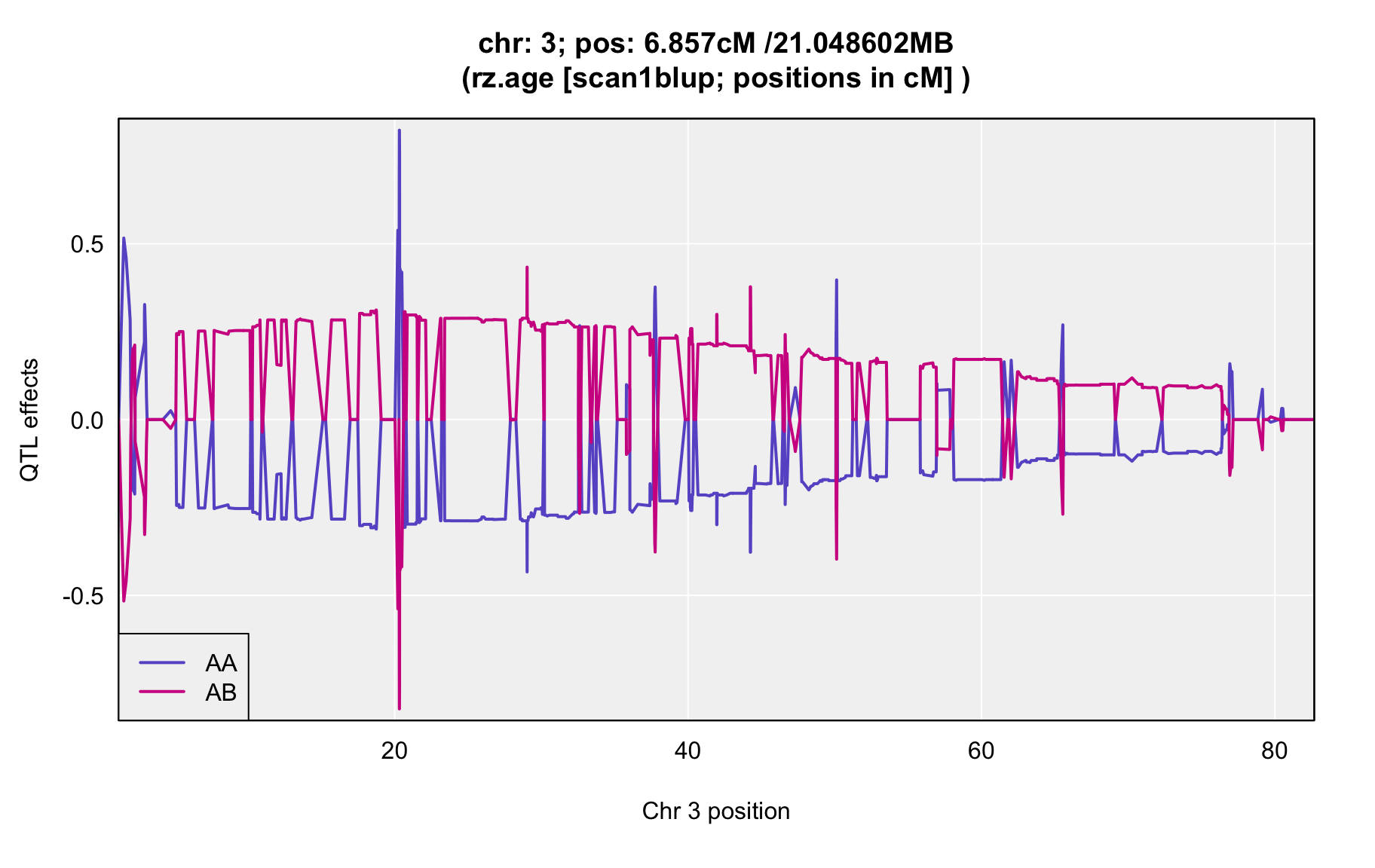
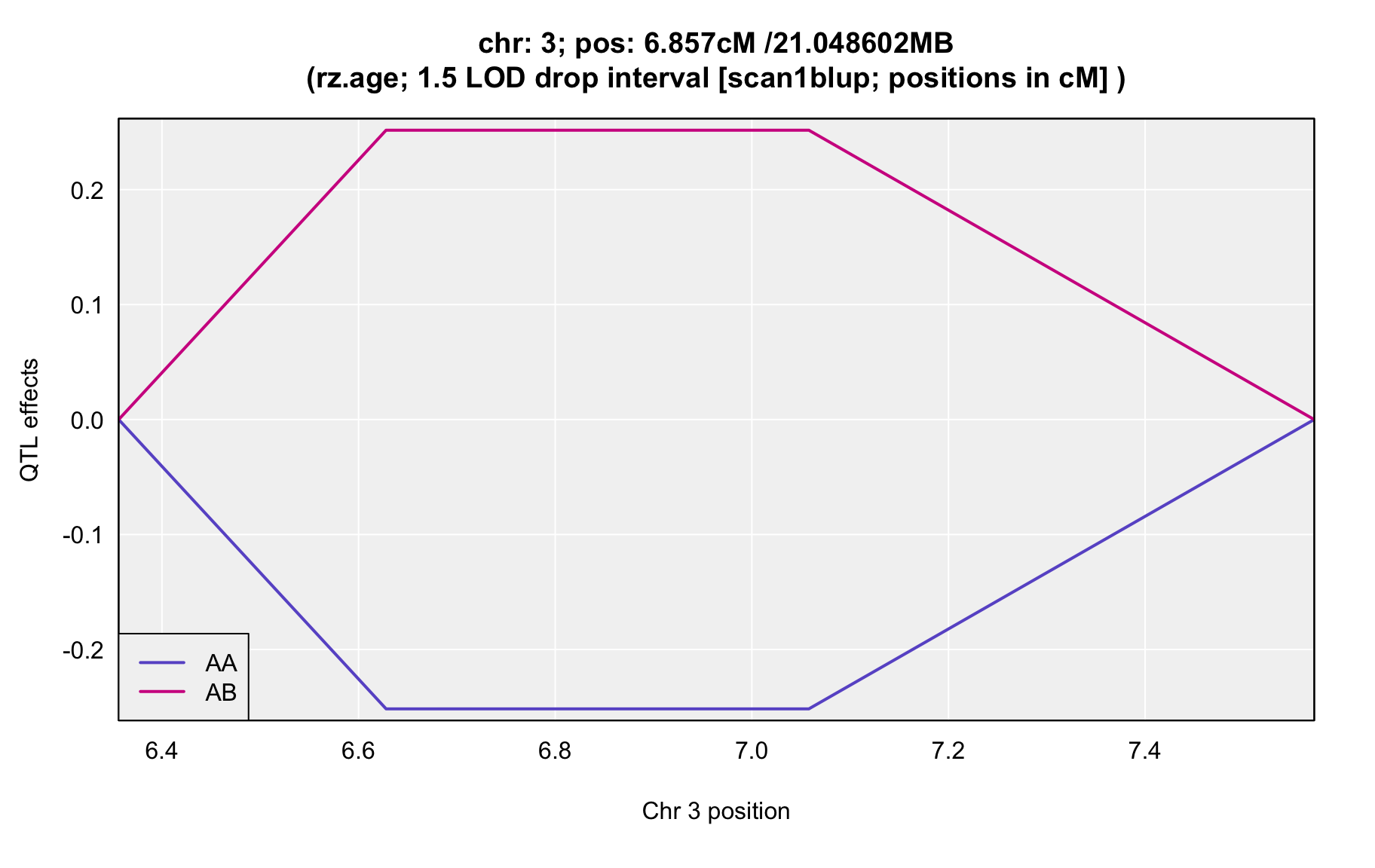
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 20.30841 | 20.30992 | positive | MGI_C57BL6J_5663662 | Gm43525 | ENSEMBL:ENSMUSG00000105208 | MGI:5663662 | lncRNA gene | predicted gene 43525 |
| 3 | gene | 20.31447 | 20.36718 | negative | MGI_C57BL6J_87965 | Agtr1b | NCBI_Gene:11608,ENSEMBL:ENSMUSG00000054988 | MGI:87965 | protein coding gene | angiotensin II receptor, type 1b |
| 3 | pseudogene | 20.39101 | 20.39198 | negative | MGI_C57BL6J_5010611 | Gm18426 | NCBI_Gene:100417153,ENSEMBL:ENSMUSG00000103573 | MGI:5010611 | pseudogene | predicted gene, 18426 |
| 3 | gene | 20.40250 | 20.50621 | negative | MGI_C57BL6J_5590625 | Gm31466 | NCBI_Gene:108168860,ENSEMBL:ENSMUSG00000102636 | MGI:5590625 | lncRNA gene | predicted gene, 31466 |
| 3 | gene | 20.40467 | 20.40491 | positive | MGI_C57BL6J_5662731 | Gm42594 | ENSEMBL:ENSMUSG00000106442 | MGI:5662731 | unclassified gene | predicted gene 42594 |
| 3 | pseudogene | 20.78247 | 20.78407 | negative | MGI_C57BL6J_5010676 | Gm18491 | NCBI_Gene:100417265,ENSEMBL:ENSMUSG00000103460 | MGI:5010676 | pseudogene | predicted gene, 18491 |
| 3 | pseudogene | 20.78794 | 20.78815 | positive | MGI_C57BL6J_5611551 | Gm38323 | ENSEMBL:ENSMUSG00000102986 | MGI:5611551 | pseudogene | predicted gene, 38323 |
| 3 | pseudogene | 20.80448 | 20.80609 | positive | MGI_C57BL6J_5010678 | Gm18493 | NCBI_Gene:100417268,ENSEMBL:ENSMUSG00000103533 | MGI:5010678 | pseudogene | predicted gene, 18493 |
| 3 | gene | 20.81639 | 20.82236 | negative | MGI_C57BL6J_5590711 | Gm31552 | NCBI_Gene:102633820 | MGI:5590711 | lncRNA gene | predicted gene, 31552 |
| 3 | gene | 20.81962 | 20.82369 | positive | MGI_C57BL6J_5610708 | Gm37480 | ENSEMBL:ENSMUSG00000102807 | MGI:5610708 | unclassified gene | predicted gene, 37480 |
| 3 | pseudogene | 21.04945 | 21.05053 | positive | MGI_C57BL6J_3647006 | Gm7488 | NCBI_Gene:665094,ENSEMBL:ENSMUSG00000039617 | MGI:3647006 | pseudogene | predicted gene 7488 |
| 3 | pseudogene | 21.11358 | 21.11400 | negative | MGI_C57BL6J_5663297 | Gm43160 | ENSEMBL:ENSMUSG00000106051 | MGI:5663297 | pseudogene | predicted gene 43160 |
| 3 | gene | 21.26837 | 21.26909 | negative | MGI_C57BL6J_5579843 | Gm29137 | ENSEMBL:ENSMUSG00000101897 | MGI:5579843 | lncRNA gene | predicted gene 29137 |
| 3 | gene | 21.42179 | 21.48966 | negative | MGI_C57BL6J_5590769 | Gm31610 | NCBI_Gene:102633894 | MGI:5590769 | lncRNA gene | predicted gene, 31610 |
| 3 | pseudogene | 21.60381 | 21.60551 | positive | MGI_C57BL6J_5663812 | Gm43675 | ENSEMBL:ENSMUSG00000106085 | MGI:5663812 | pseudogene | predicted gene 43675 |
| 3 | pseudogene | 21.64839 | 21.64918 | positive | MGI_C57BL6J_5010120 | Gm17935 | NCBI_Gene:100416137,ENSEMBL:ENSMUSG00000106460 | MGI:5010120 | pseudogene | predicted gene, 17935 |
| 3 | gene | 21.70282 | 21.71246 | positive | MGI_C57BL6J_5590852 | Gm31693 | NCBI_Gene:102634003,ENSEMBL:ENSMUSG00000105440 | MGI:5590852 | lncRNA gene | predicted gene, 31693 |
| 3 | gene | 21.75353 | 21.80080 | negative | MGI_C57BL6J_2442162 | 7530428D23Rik | NCBI_Gene:319506,ENSEMBL:ENSMUSG00000103441 | MGI:2442162 | lncRNA gene | RIKEN cDNA 7530428D23 gene |
| 3 | gene | 21.86417 | 21.96435 | negative | MGI_C57BL6J_5590972 | Gm31813 | NCBI_Gene:102634161 | MGI:5590972 | lncRNA gene | predicted gene, 31813 |
| 3 | gene | 21.99768 | 21.99847 | negative | MGI_C57BL6J_5663811 | Gm43674 | ENSEMBL:ENSMUSG00000106321 | MGI:5663811 | lncRNA gene | predicted gene 43674 |
| 3 | pseudogene | 22.02120 | 22.02163 | positive | MGI_C57BL6J_5663808 | Gm43671 | ENSEMBL:ENSMUSG00000106527 | MGI:5663808 | pseudogene | predicted gene 43671 |
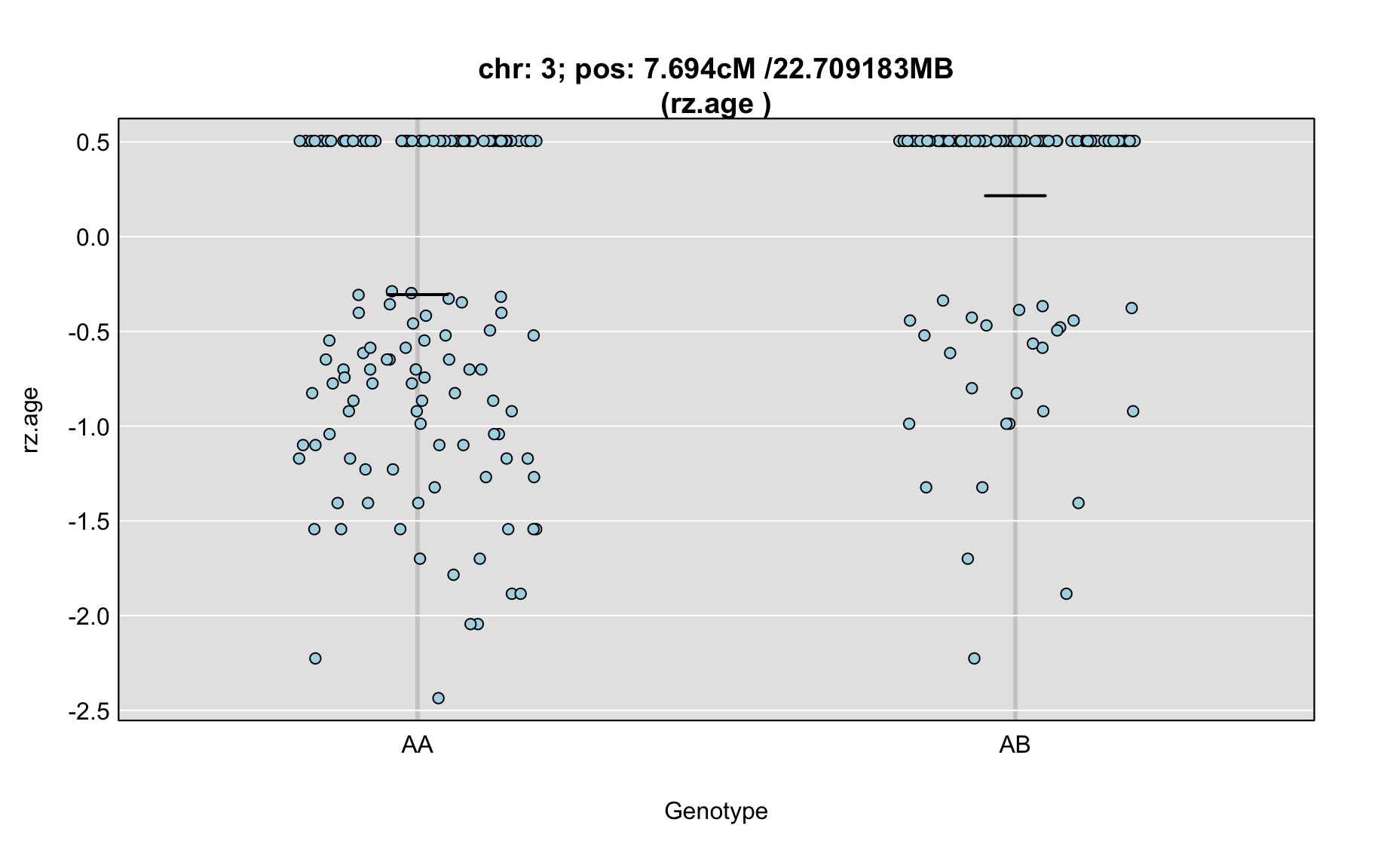
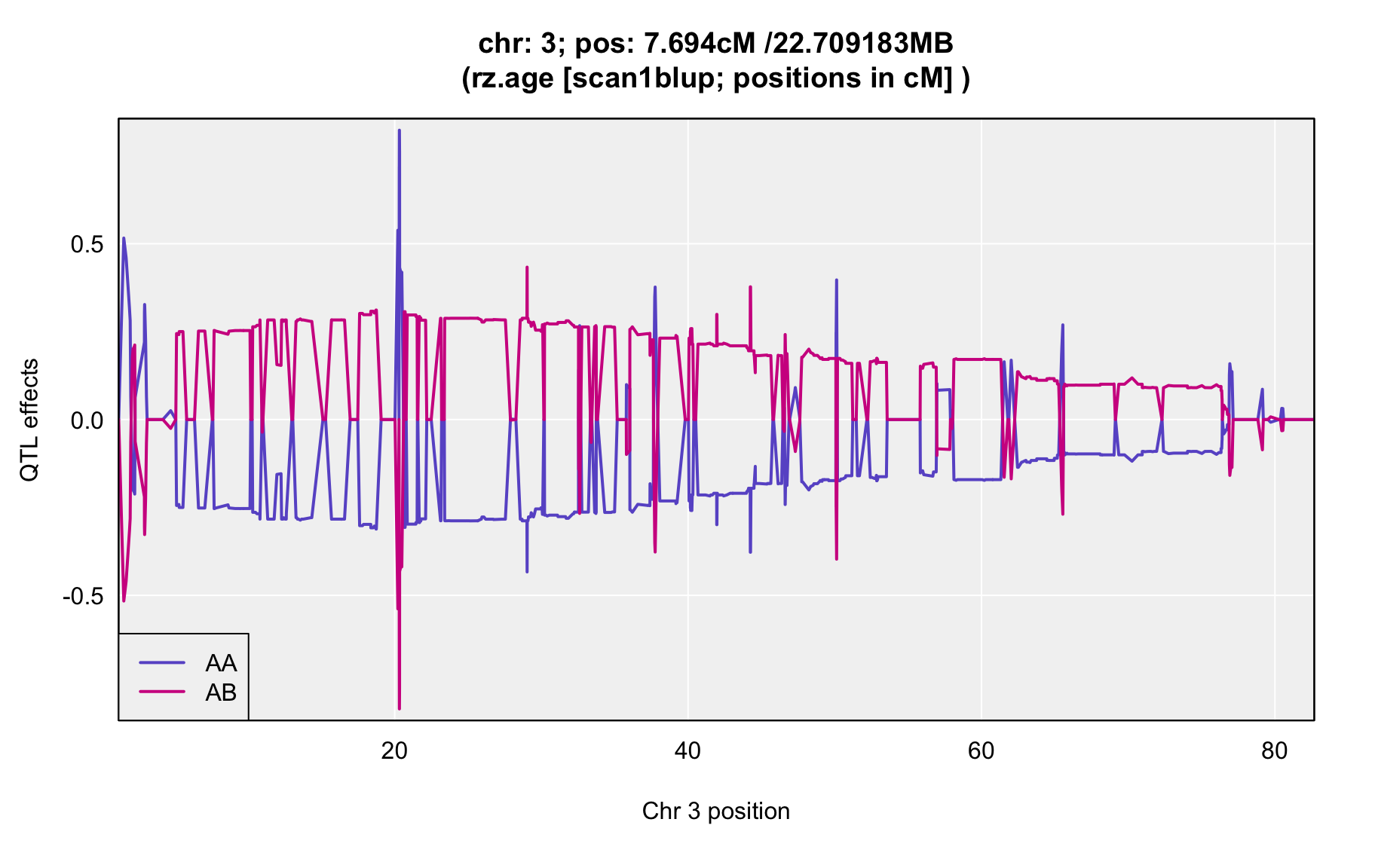
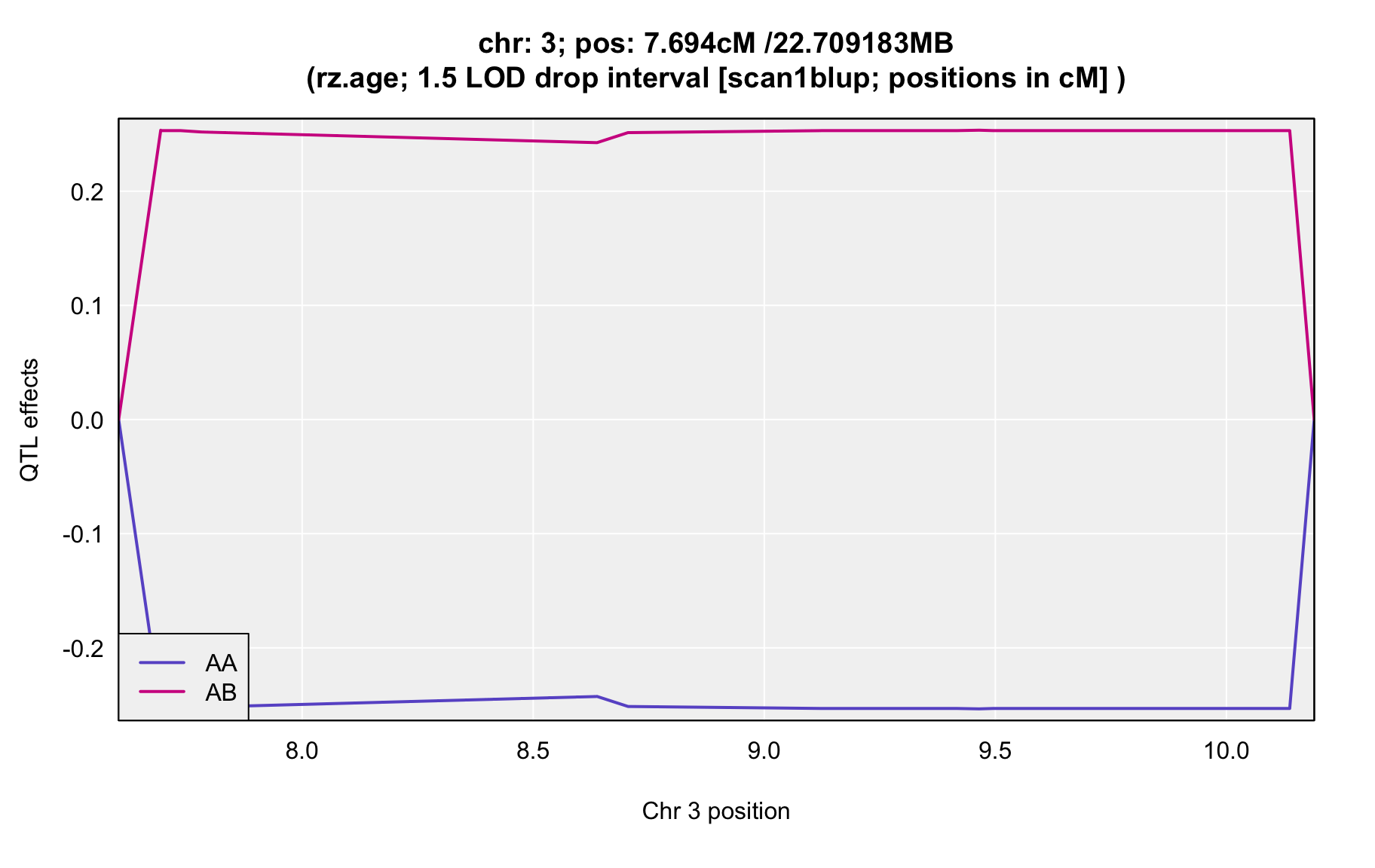
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 22.07659 | 22.21659 | positive | MGI_C57BL6J_2441730 | Tbl1xr1 | NCBI_Gene:81004,ENSEMBL:ENSMUSG00000027630 | MGI:2441730 | protein coding gene | transducin (beta)-like 1X-linked receptor 1 |
| 3 | gene | 22.25137 | 22.25166 | positive | MGI_C57BL6J_105104 | Rprl2 | NCBI_Gene:19784,ENSEMBL:ENSMUSG00000087775 | MGI:105104 | RNase P RNA gene | ribonuclease P RNA-like 2 |
| 3 | gene | 22.28508 | 22.29783 | negative | MGI_C57BL6J_5611552 | Gm38324 | ENSEMBL:ENSMUSG00000102987 | MGI:5611552 | lncRNA gene | predicted gene, 38324 |
| 3 | pseudogene | 22.34885 | 22.34961 | negative | MGI_C57BL6J_5610864 | Gm37636 | NCBI_Gene:108168850,ENSEMBL:ENSMUSG00000102866 | MGI:5610864 | pseudogene | predicted gene, 37636 |
| 3 | gene | 22.40992 | 22.41005 | negative | MGI_C57BL6J_5456131 | Gm26354 | ENSEMBL:ENSMUSG00000088846 | MGI:5456131 | snoRNA gene | predicted gene, 26354 |
| 3 | pseudogene | 22.52501 | 22.52624 | positive | MGI_C57BL6J_3780056 | Spin3-ps | NCBI_Gene:675593,ENSEMBL:ENSMUSG00000103801 | MGI:3780056 | pseudogene | spindlin family, member 3, pseudogene |
| 3 | gene | 22.81221 | 22.81246 | positive | MGI_C57BL6J_5611117 | Gm37889 | ENSEMBL:ENSMUSG00000102233 | MGI:5611117 | unclassified gene | predicted gene, 37889 |
| 3 | pseudogene | 23.07242 | 23.07492 | positive | MGI_C57BL6J_5610211 | Gm36983 | ENSEMBL:ENSMUSG00000103275 | MGI:5610211 | pseudogene | predicted gene, 36983 |
| 3 | gene | 23.08598 | 23.08895 | positive | MGI_C57BL6J_5610249 | Gm37021 | ENSEMBL:ENSMUSG00000103607 | MGI:5610249 | lncRNA gene | predicted gene, 37021 |
| 3 | pseudogene | 23.37494 | 23.37532 | negative | MGI_C57BL6J_5610720 | Gm37492 | ENSEMBL:ENSMUSG00000102214 | MGI:5610720 | pseudogene | predicted gene, 37492 |
| 3 | gene | 23.46105 | 23.52140 | positive | MGI_C57BL6J_1924114 | 4930412E21Rik | NCBI_Gene:100039548,ENSEMBL:ENSMUSG00000102203 | MGI:1924114 | lncRNA gene | RIKEN cDNA 4930412E21 gene |
| 3 | gene | 23.79810 | 25.14426 | negative | MGI_C57BL6J_2685867 | Naaladl2 | NCBI_Gene:635702,ENSEMBL:ENSMUSG00000102758 | MGI:2685867 | protein coding gene | N-acetylated alpha-linked acidic dipeptidase-like 2 |
| 3 | pseudogene | 23.93096 | 23.93200 | positive | MGI_C57BL6J_5011009 | Gm18824 | NCBI_Gene:100417784 | MGI:5011009 | pseudogene | predicted gene, 18824 |
| 3 | gene | 23.94002 | 23.94593 | positive | MGI_C57BL6J_5625076 | Gm42191 | NCBI_Gene:105247007 | MGI:5625076 | lncRNA gene | predicted gene, 42191 |
| 3 | gene | 24.16303 | 24.16311 | negative | MGI_C57BL6J_4413931 | n-TLcag1 | NCBI_Gene:102467553 | MGI:4413931 | tRNA gene | nuclear encoded tRNA leucine 1 (anticodon CAG) |
| 3 | pseudogene | 24.33304 | 24.33355 | positive | MGI_C57BL6J_3645137 | Gm7536 | NCBI_Gene:665189,ENSEMBL:ENSMUSG00000057036 | MGI:3645137 | pseudogene | predicted gene 7536 |
| 3 | gene | 24.48671 | 24.48749 | positive | MGI_C57BL6J_1920306 | 2900080J11Rik | NA | NA | unclassified gene | RIKEN cDNA 2900080J11 gene |
| 3 | gene | 24.56207 | 24.56218 | positive | MGI_C57BL6J_5454481 | Gm24704 | ENSEMBL:ENSMUSG00000070227 | MGI:5454481 | snRNA gene | predicted gene, 24704 |
| 3 | gene | 24.71872 | 24.71898 | positive | MGI_C57BL6J_5453839 | Gm24062 | ENSEMBL:ENSMUSG00000088115 | MGI:5453839 | unclassified non-coding RNA gene | predicted gene, 24062 |
| 3 | pseudogene | 25.00742 | 25.00771 | negative | MGI_C57BL6J_5662911 | Gm42774 | ENSEMBL:ENSMUSG00000105009 | MGI:5662911 | pseudogene | predicted gene 42774 |
| 3 | gene | 25.14407 | 25.14426 | positive | MGI_C57BL6J_5610364 | Gm37136 | ENSEMBL:ENSMUSG00000102876 | MGI:5610364 | unclassified gene | predicted gene, 37136 |
| 3 | pseudogene | 25.18480 | 25.18498 | negative | MGI_C57BL6J_5611230 | Gm38002 | ENSEMBL:ENSMUSG00000102855 | MGI:5611230 | pseudogene | predicted gene, 38002 |
| 3 | pseudogene | 25.21076 | 25.21273 | negative | MGI_C57BL6J_3704211 | Gm10259 | NCBI_Gene:100039592,ENSEMBL:ENSMUSG00000069083 | MGI:3704211 | pseudogene | predicted pseudogene 10259 |
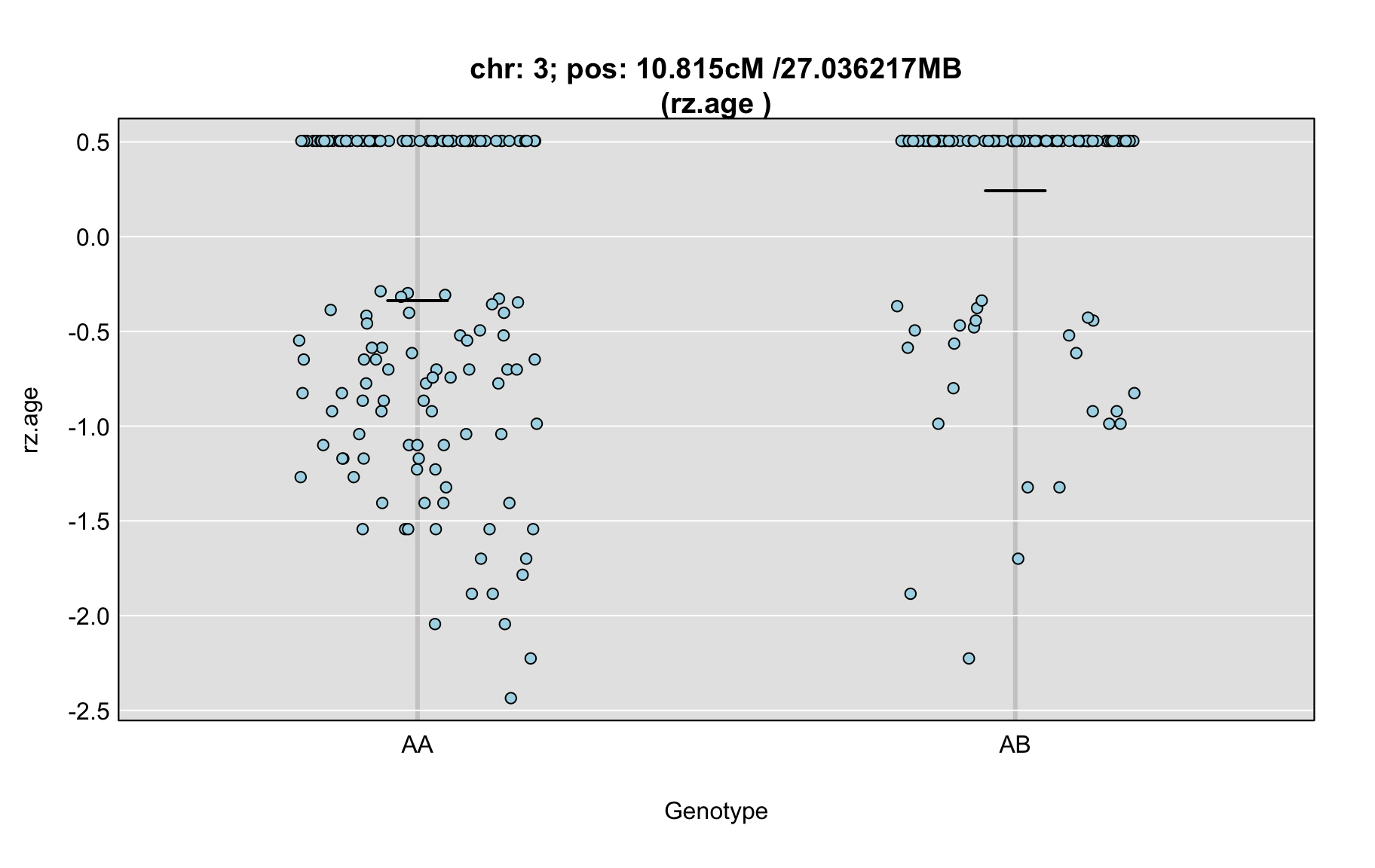
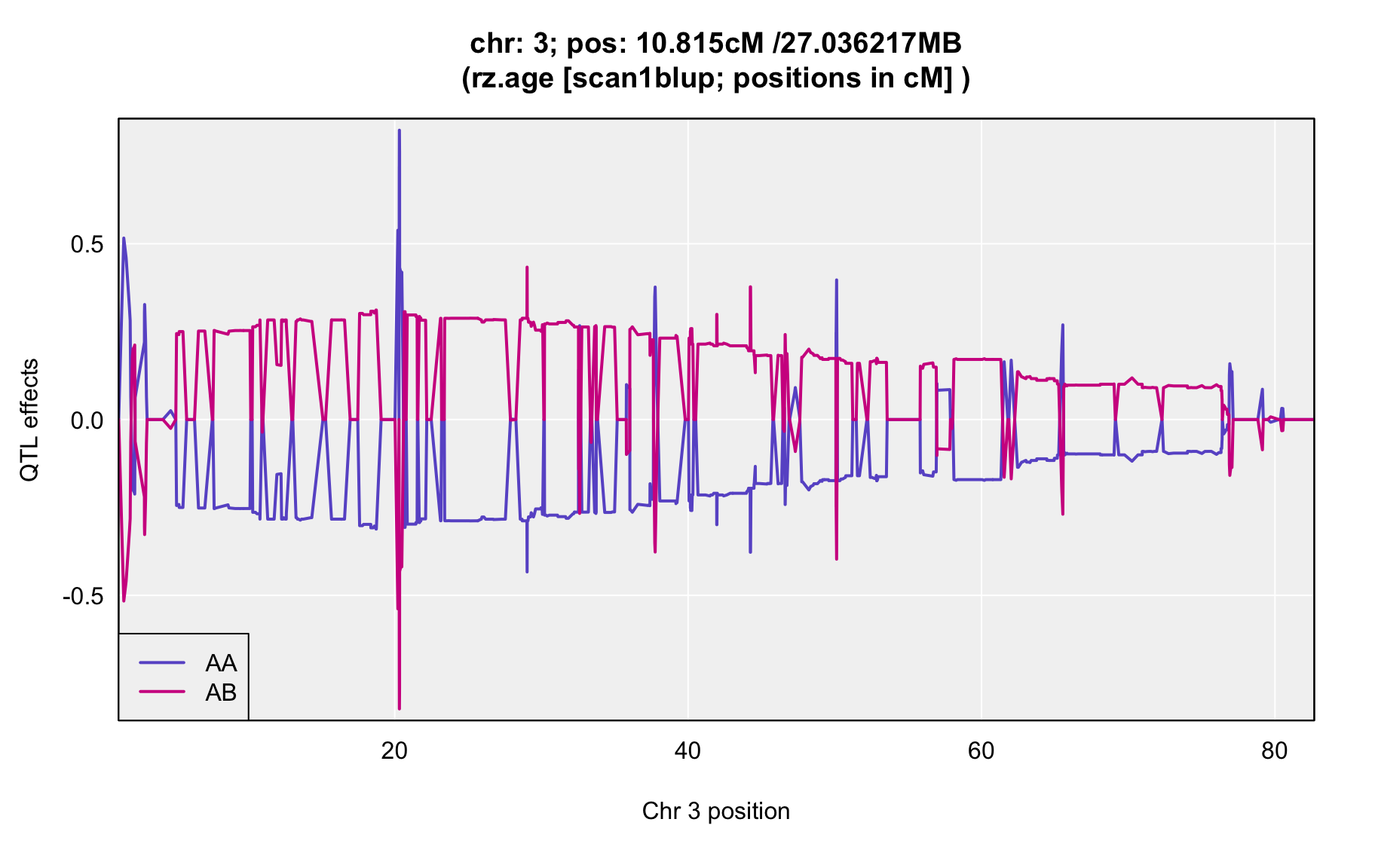
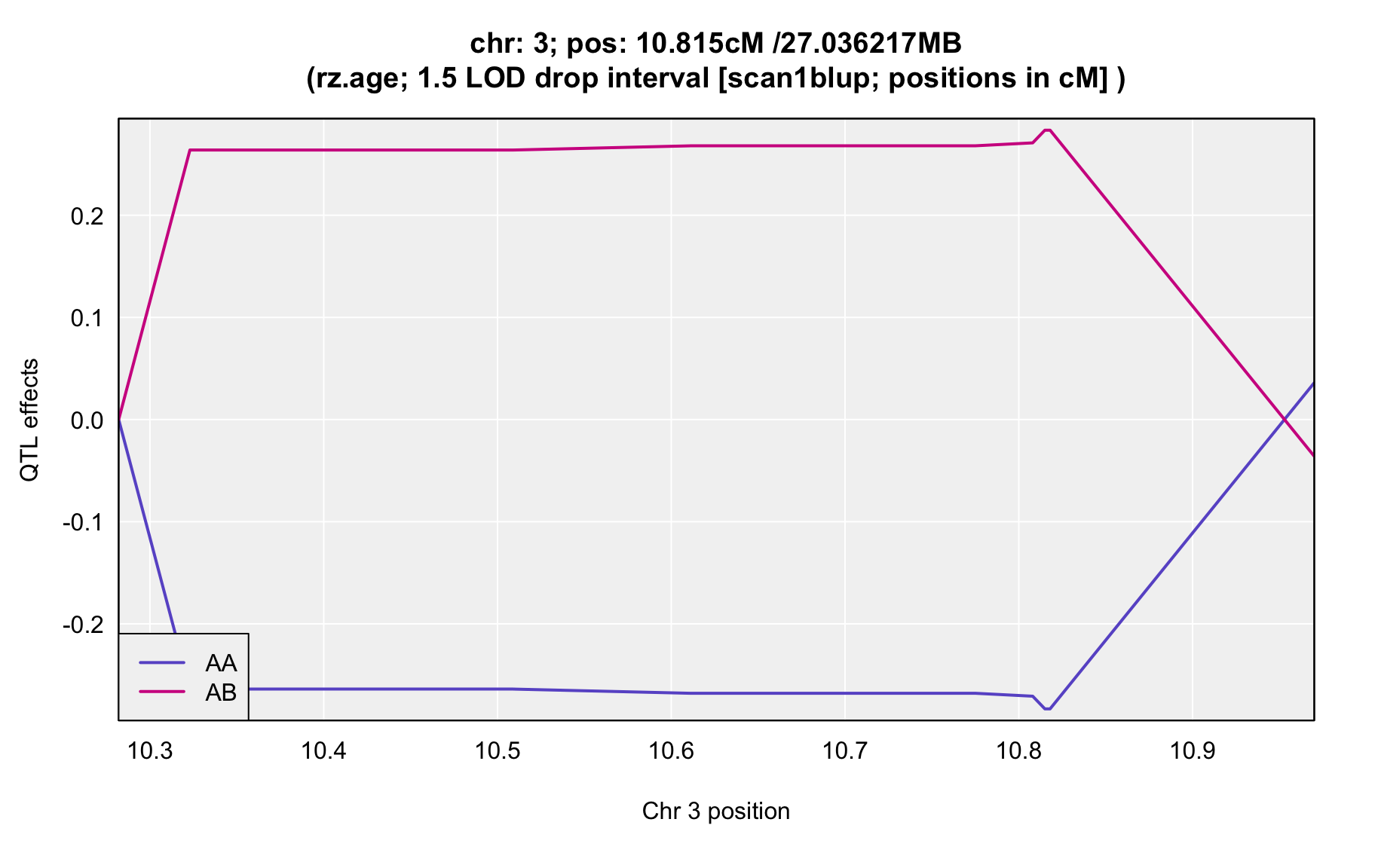
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 25.42621 | 26.33286 | negative | MGI_C57BL6J_2179435 | Nlgn1 | NCBI_Gene:192167,ENSEMBL:ENSMUSG00000063887 | MGI:2179435 | protein coding gene | neuroligin 1 |
| 3 | gene | 25.54602 | 25.54615 | positive | MGI_C57BL6J_5454027 | Gm24250 | ENSEMBL:ENSMUSG00000087945 | MGI:5454027 | rRNA gene | predicted gene, 24250 |
| 3 | gene | 25.64956 | 25.65077 | negative | MGI_C57BL6J_1920377 | 3110039C02Rik | NA | NA | unclassified gene | RIKEN cDNA 3110039C02 gene |
| 3 | gene | 25.69519 | 25.70030 | negative | MGI_C57BL6J_5610801 | Gm37573 | ENSEMBL:ENSMUSG00000102397 | MGI:5610801 | lncRNA gene | predicted gene, 37573 |
| 3 | pseudogene | 25.73123 | 25.73261 | positive | MGI_C57BL6J_3647151 | Gm6197 | NCBI_Gene:621017,ENSEMBL:ENSMUSG00000104295 | MGI:3647151 | pseudogene | predicted gene 6197 |
| 3 | gene | 25.86566 | 25.86752 | negative | MGI_C57BL6J_2443798 | A830010M09Rik | NA | NA | unclassified gene | RIKEN cDNA A830010M09 gene |
| 3 | gene | 26.33115 | 26.33688 | positive | MGI_C57BL6J_3698881 | A830092H15Rik | NCBI_Gene:100379095,ENSEMBL:ENSMUSG00000074664 | MGI:3698881 | lncRNA gene | RIKEN cDNA A830092H15 gene |
| 3 | pseudogene | 26.38044 | 26.38256 | positive | MGI_C57BL6J_3648363 | Gm4855 | NCBI_Gene:229152,ENSEMBL:ENSMUSG00000102274 | MGI:3648363 | pseudogene | predicted gene 4855 |
| 3 | gene | 26.62618 | 26.62631 | negative | MGI_C57BL6J_5455550 | Gm25773 | ENSEMBL:ENSMUSG00000080368 | MGI:5455550 | miRNA gene | predicted gene, 25773 |
| 3 | gene | 26.63762 | 26.98321 | positive | MGI_C57BL6J_1918112 | Spata16 | NCBI_Gene:70862,ENSEMBL:ENSMUSG00000039335 | MGI:1918112 | protein coding gene | spermatogenesis associated 16 |
| 3 | gene | 26.85020 | 26.87914 | negative | MGI_C57BL6J_5591617 | Gm32458 | NCBI_Gene:102635017 | MGI:5591617 | lncRNA gene | predicted gene, 32458 |
| 3 | pseudogene | 27.01276 | 27.01287 | positive | MGI_C57BL6J_5610887 | Gm37659 | ENSEMBL:ENSMUSG00000103712 | MGI:5610887 | pseudogene | predicted gene, 37659 |
| 3 | gene | 27.02642 | 27.02655 | negative | MGI_C57BL6J_5690856 | Gm44464 | ENSEMBL:ENSMUSG00000105670 | MGI:5690856 | miRNA gene | predicted gene, 44464 |
| 3 | pseudogene | 27.04674 | 27.04798 | negative | MGI_C57BL6J_3645515 | Gm7558 | NCBI_Gene:665258,ENSEMBL:ENSMUSG00000102908 | MGI:3645515 | pseudogene | predicted gene 7558 |
| 3 | gene | 27.09722 | 27.15391 | negative | MGI_C57BL6J_95281 | Ect2 | NCBI_Gene:13605,ENSEMBL:ENSMUSG00000027699 | MGI:95281 | protein coding gene | ect2 oncogene |
| 3 | gene | 27.10898 | 27.10909 | positive | MGI_C57BL6J_5454929 | Gm25152 | ENSEMBL:ENSMUSG00000087742 | MGI:5454929 | snoRNA gene | predicted gene, 25152 |
| 3 | gene | 27.15403 | 27.15702 | positive | MGI_C57BL6J_1923907 | 1700125G22Rik | NCBI_Gene:76657,ENSEMBL:ENSMUSG00000097584 | MGI:1923907 | lncRNA gene | RIKEN cDNA 1700125G22 gene |
| 3 | gene | 27.18296 | 27.28461 | positive | MGI_C57BL6J_2443191 | Nceh1 | NCBI_Gene:320024,ENSEMBL:ENSMUSG00000027698 | MGI:2443191 | protein coding gene | neutral cholesterol ester hydrolase 1 |
| 3 | gene | 27.26033 | 27.28446 | positive | MGI_C57BL6J_5625077 | Gm42192 | NCBI_Gene:105247008 | MGI:5625077 | lncRNA gene | predicted gene, 42192 |
| 3 | pseudogene | 27.27806 | 27.27845 | positive | MGI_C57BL6J_5610419 | Gm37191 | ENSEMBL:ENSMUSG00000102186 | MGI:5610419 | pseudogene | predicted gene, 37191 |
| 3 | pseudogene | 27.30841 | 27.30867 | negative | MGI_C57BL6J_5663482 | Gm43345 | ENSEMBL:ENSMUSG00000105806 | MGI:5663482 | pseudogene | predicted gene 43345 |
| 3 | gene | 27.31703 | 27.34243 | positive | MGI_C57BL6J_107414 | Tnfsf10 | NCBI_Gene:22035,ENSEMBL:ENSMUSG00000039304 | MGI:107414 | protein coding gene | tumor necrosis factor (ligand) superfamily, member 10 |
| 3 | gene | 27.37131 | 27.37923 | positive | MGI_C57BL6J_2441906 | Ghsr | NCBI_Gene:208188,ENSEMBL:ENSMUSG00000051136 | MGI:2441906 | protein coding gene | growth hormone secretagogue receptor |
| 3 | gene | 27.41511 | 27.41615 | negative | MGI_C57BL6J_1924904 | C430049A07Rik | NA | NA | unclassified gene | RIKEN cDNA C430049A07 gene |
| 3 | gene | 27.41616 | 27.71311 | negative | MGI_C57BL6J_1919257 | Fndc3b | NCBI_Gene:72007,ENSEMBL:ENSMUSG00000039286 | MGI:1919257 | protein coding gene | fibronectin type III domain containing 3B |
| 3 | gene | 27.44166 | 27.44411 | positive | MGI_C57BL6J_5625078 | Gm42193 | NCBI_Gene:105247009 | MGI:5625078 | lncRNA gene | predicted gene, 42193 |
| 3 | gene | 27.48228 | 27.48424 | positive | MGI_C57BL6J_5591897 | Gm32738 | NCBI_Gene:102635381 | MGI:5591897 | lncRNA gene | predicted gene, 32738 |
| 3 | gene | 27.54664 | 27.54787 | positive | MGI_C57BL6J_5663481 | Gm43344 | ENSEMBL:ENSMUSG00000106625 | MGI:5663481 | unclassified gene | predicted gene 43344 |
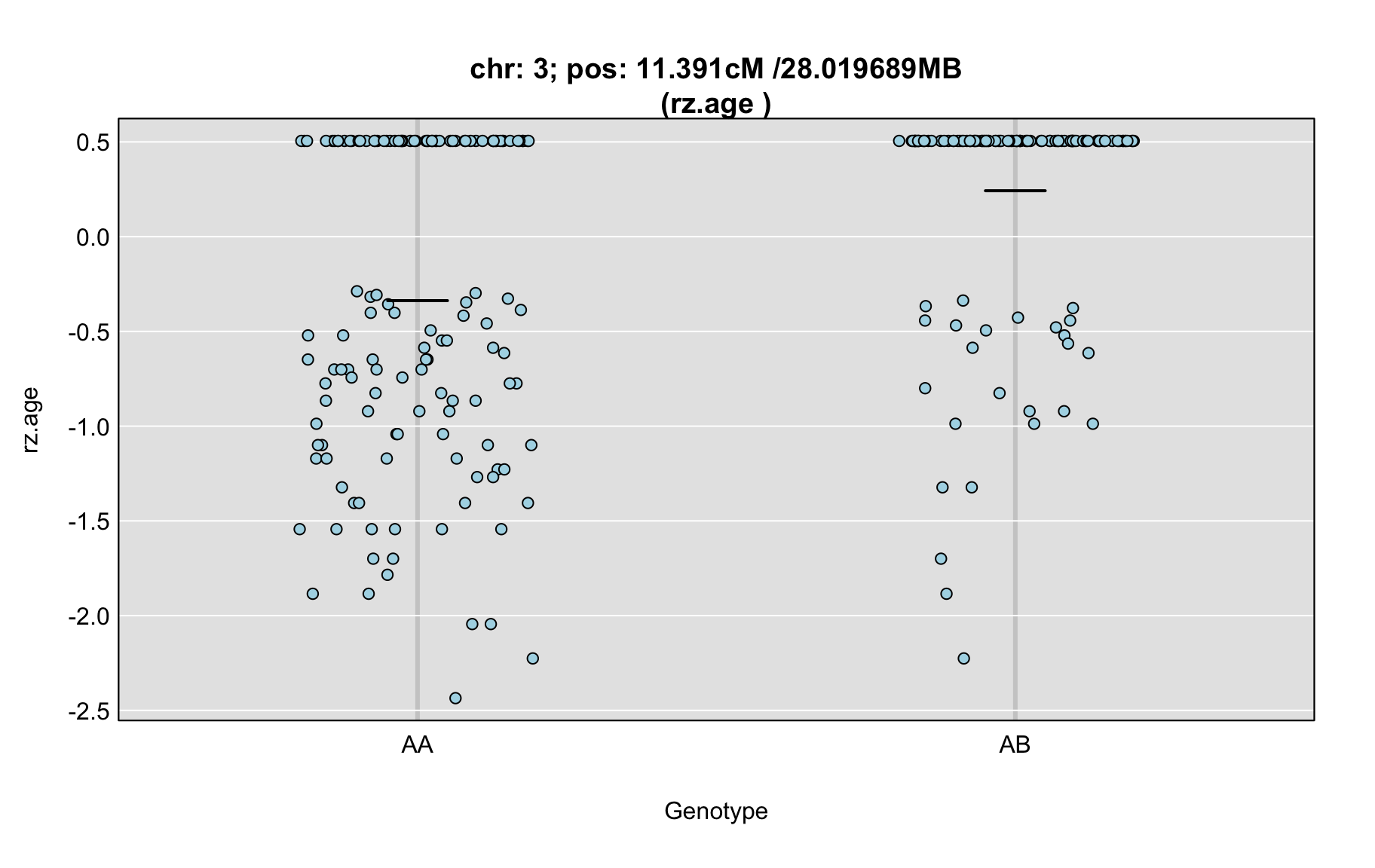
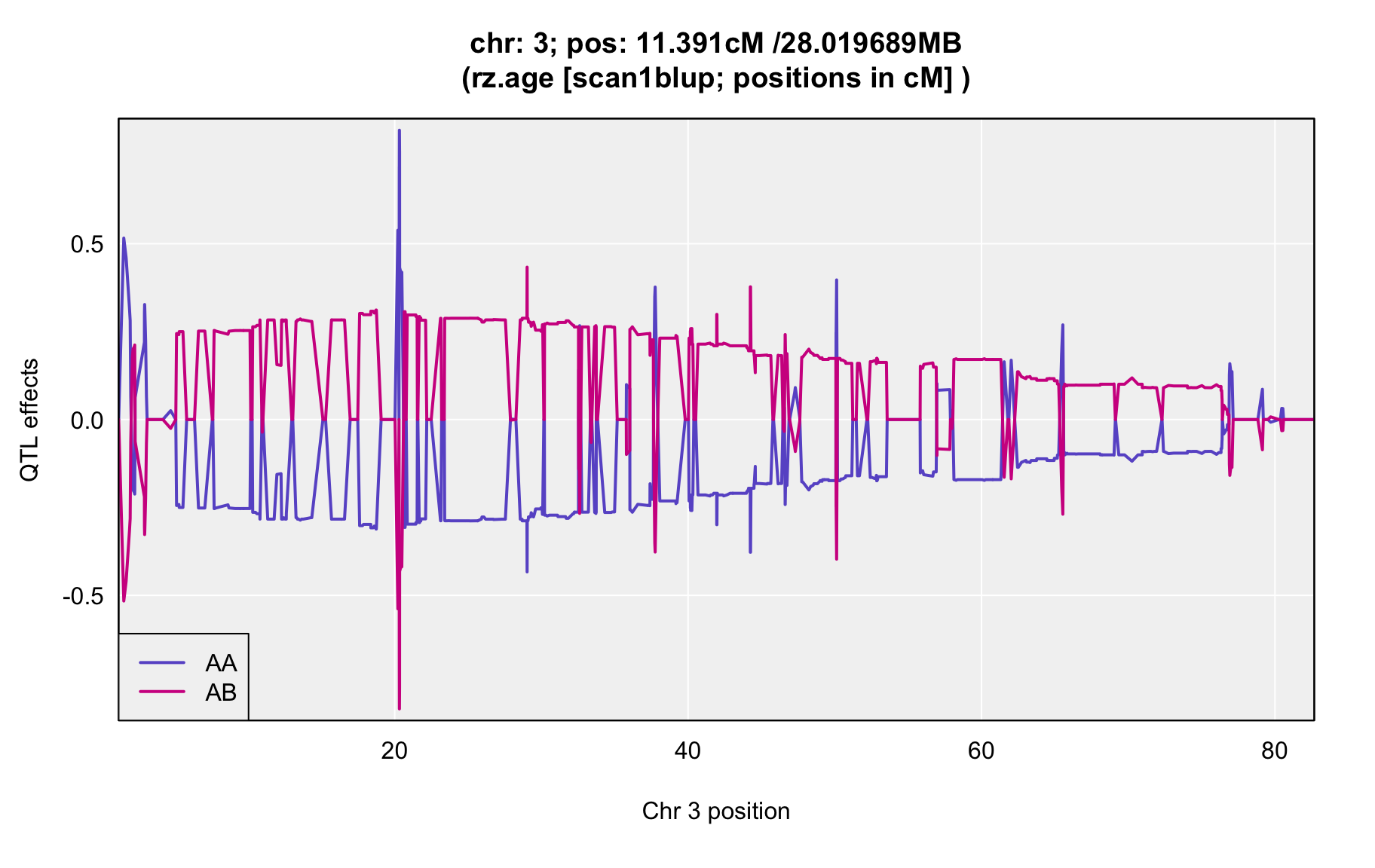
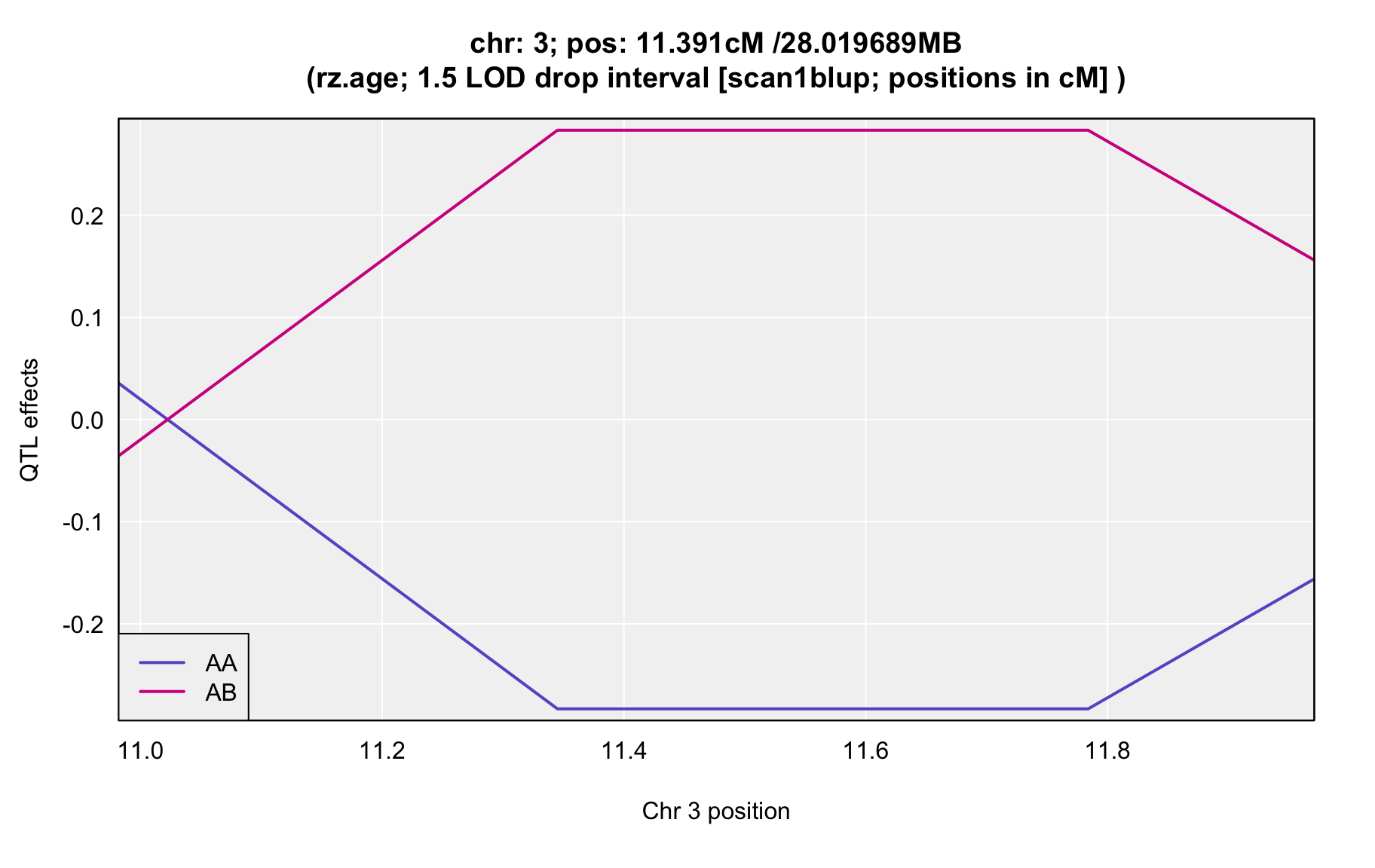
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 27.41616 | 27.71311 | negative | MGI_C57BL6J_1919257 | Fndc3b | NCBI_Gene:72007,ENSEMBL:ENSMUSG00000039286 | MGI:1919257 | protein coding gene | fibronectin type III domain containing 3B |
| 3 | gene | 27.58490 | 27.58499 | negative | MGI_C57BL6J_4834306 | Mir3092 | miRBase:MI0014085,NCBI_Gene:100526525,ENSEMBL:ENSMUSG00000092695 | MGI:4834306 | miRNA gene | microRNA 3092 |
| 3 | gene | 27.77786 | 27.84092 | positive | MGI_C57BL6J_5625079 | Gm42194 | NCBI_Gene:105247010 | MGI:5625079 | lncRNA gene | predicted gene, 42194 |
| 3 | gene | 27.79788 | 27.81538 | negative | MGI_C57BL6J_5591954 | Gm32795 | NCBI_Gene:102635463 | MGI:5591954 | lncRNA gene | predicted gene, 32795 |
| 3 | gene | 27.86606 | 27.89639 | negative | MGI_C57BL6J_2685410 | Tmem212 | NCBI_Gene:208613,ENSEMBL:ENSMUSG00000043164 | MGI:2685410 | protein coding gene | transmembrane protein 212 |
| 3 | gene | 27.86811 | 27.86827 | positive | MGI_C57BL6J_5455817 | Gm26040 | ENSEMBL:ENSMUSG00000088772 | MGI:5455817 | snoRNA gene | predicted gene, 26040 |
| 3 | gene | 27.89680 | 27.90023 | negative | MGI_C57BL6J_5625080 | Gm42195 | NCBI_Gene:105247011 | MGI:5625080 | lncRNA gene | predicted gene, 42195 |
| 3 | gene | 27.93841 | 28.13336 | positive | MGI_C57BL6J_109585 | Pld1 | NCBI_Gene:18805,ENSEMBL:ENSMUSG00000027695 | MGI:109585 | protein coding gene | phospholipase D1 |
| 3 | gene | 28.17651 | 28.21221 | positive | MGI_C57BL6J_5625081 | Gm42196 | NCBI_Gene:105247012,ENSEMBL:ENSMUSG00000109716 | MGI:5625081 | lncRNA gene | predicted gene, 42196 |
| 3 | pseudogene | 28.24552 | 28.24577 | positive | MGI_C57BL6J_5610326 | Gm37098 | ENSEMBL:ENSMUSG00000103729 | MGI:5610326 | pseudogene | predicted gene, 37098 |
| 3 | gene | 28.26321 | 28.67586 | positive | MGI_C57BL6J_1916264 | Tnik | NCBI_Gene:665113,ENSEMBL:ENSMUSG00000027692 | MGI:1916264 | protein coding gene | TRAF2 and NCK interacting kinase |
| 3 | gene | 28.26928 | 28.26983 | positive | MGI_C57BL6J_2139599 | AA589472 | NA | NA | unclassified gene | expressed sequence AA589472 |
| 3 | gene | 28.32436 | 28.32609 | positive | MGI_C57BL6J_5610765 | Gm37537 | ENSEMBL:ENSMUSG00000103420 | MGI:5610765 | unclassified gene | predicted gene, 37537 |
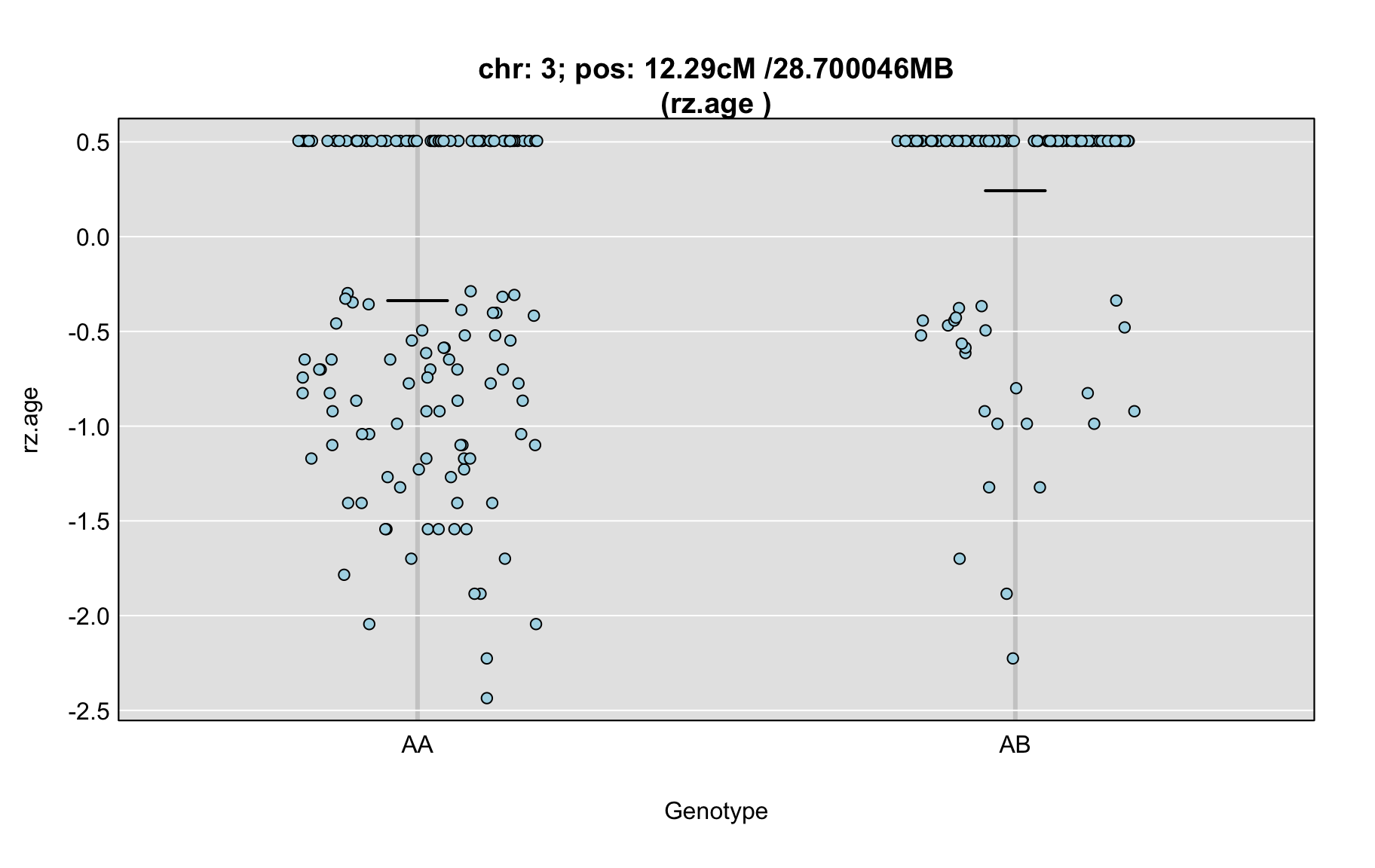
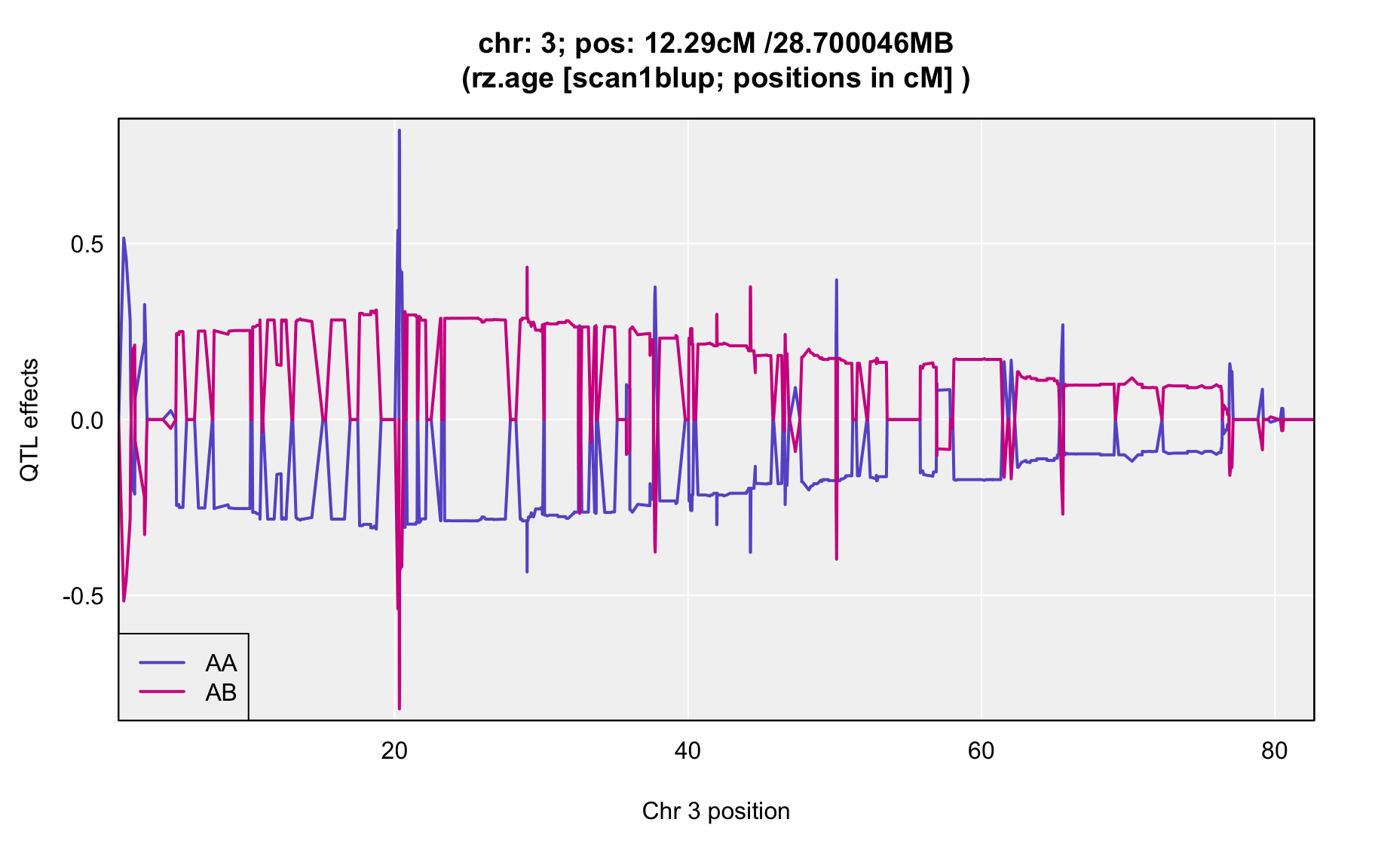
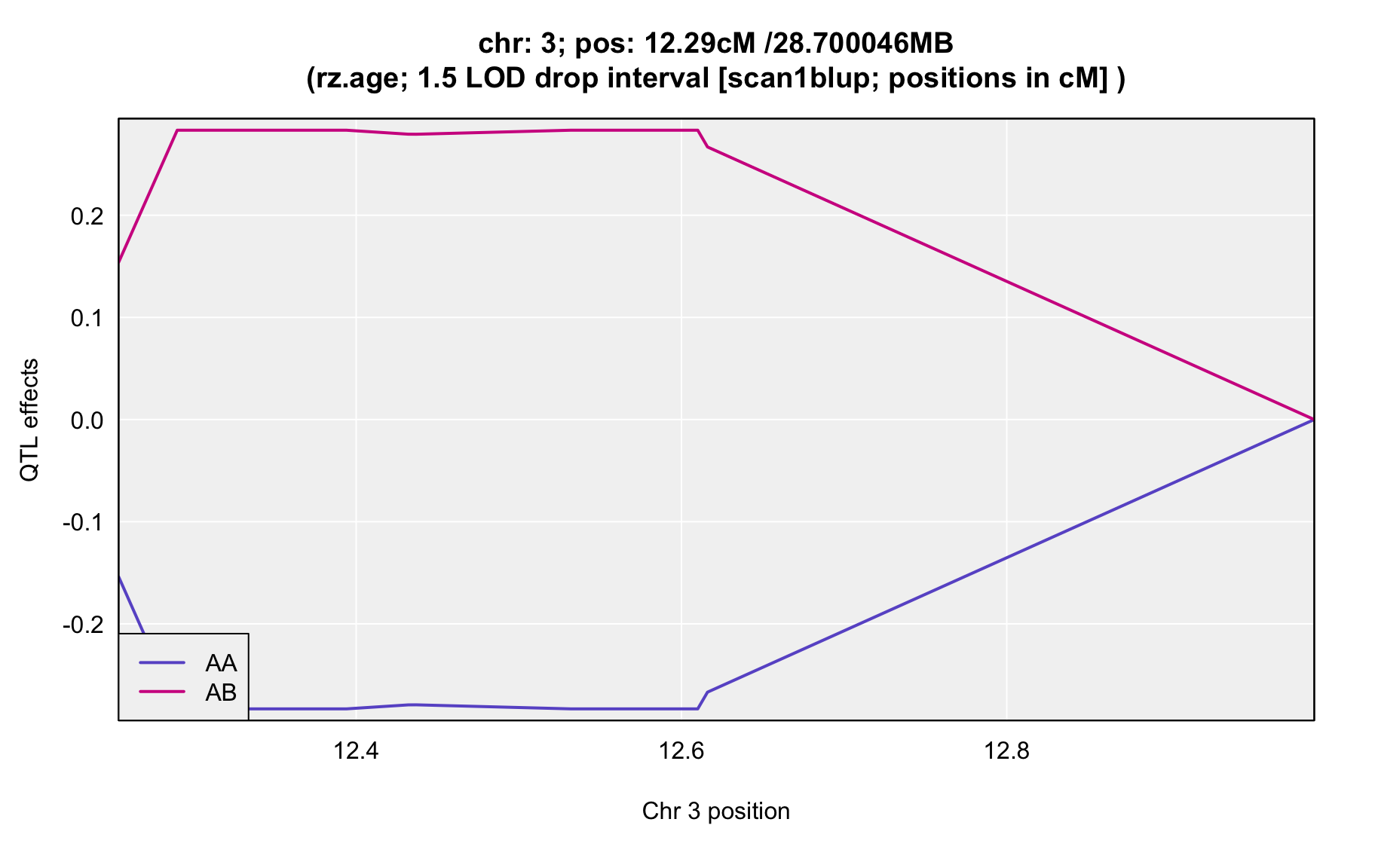
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 28.26321 | 28.67586 | positive | MGI_C57BL6J_1916264 | Tnik | NCBI_Gene:665113,ENSEMBL:ENSMUSG00000027692 | MGI:1916264 | protein coding gene | TRAF2 and NCK interacting kinase |
| 3 | gene | 28.69790 | 28.73136 | positive | MGI_C57BL6J_1095438 | Slc2a2 | NCBI_Gene:20526,ENSEMBL:ENSMUSG00000027690 | MGI:1095438 | protein coding gene | solute carrier family 2 (facilitated glucose transporter), member 2 |
| 3 | gene | 28.70267 | 28.78111 | negative | MGI_C57BL6J_1920861 | 1700112D23Rik | NCBI_Gene:73611,ENSEMBL:ENSMUSG00000091329 | MGI:1920861 | lncRNA gene | RIKEN cDNA 1700112D23 gene |
| 3 | pseudogene | 28.76353 | 28.76539 | negative | MGI_C57BL6J_3648080 | Gm6505 | NCBI_Gene:624446,ENSEMBL:ENSMUSG00000070522 | MGI:3648080 | pseudogene | predicted pseudogene 6505 |
| 3 | gene | 28.78127 | 28.79885 | positive | MGI_C57BL6J_1933735 | Eif5a2 | NCBI_Gene:208691,ENSEMBL:ENSMUSG00000050192 | MGI:1933735 | protein coding gene | eukaryotic translation initiation factor 5A2 |
| 3 | gene | 28.79064 | 28.80542 | negative | MGI_C57BL6J_5592109 | Gm32950 | NCBI_Gene:102635671,ENSEMBL:ENSMUSG00000103827 | MGI:5592109 | lncRNA gene | predicted gene, 32950 |
| 3 | gene | 28.80544 | 28.80742 | positive | MGI_C57BL6J_1915278 | Rpl22l1 | NCBI_Gene:68028,ENSEMBL:ENSMUSG00000039221 | MGI:1915278 | protein coding gene | ribosomal protein L22 like 1 |
| 3 | gene | 28.80720 | 28.81618 | negative | MGI_C57BL6J_5592210 | Gm33051 | NCBI_Gene:102635804,ENSEMBL:ENSMUSG00000104444 | MGI:5592210 | lncRNA gene | predicted gene, 33051 |
| 3 | gene | 28.85203 | 28.85426 | negative | MGI_C57BL6J_5611095 | Gm37867 | ENSEMBL:ENSMUSG00000104275 | MGI:5611095 | lncRNA gene | predicted gene, 37867 |
| 3 | gene | 28.87981 | 28.88346 | positive | MGI_C57BL6J_5611152 | Gm37924 | ENSEMBL:ENSMUSG00000102482 | MGI:5611152 | unclassified gene | predicted gene, 37924 |
| 3 | gene | 28.88346 | 28.89728 | negative | MGI_C57BL6J_5592261 | Gm33102 | NCBI_Gene:102635876 | MGI:5592261 | lncRNA gene | predicted gene, 33102 |
| 3 | gene | 28.89262 | 28.92672 | positive | MGI_C57BL6J_2686373 | Gm1527 | NCBI_Gene:385263,ENSEMBL:ENSMUSG00000074655 | MGI:2686373 | protein coding gene | predicted gene 1527 |
| 3 | gene | 29.04377 | 29.04555 | negative | MGI_C57BL6J_5592313 | Gm33154 | NCBI_Gene:102635945 | MGI:5592313 | lncRNA gene | predicted gene, 33154 |
| 3 | gene | 29.08072 | 29.69121 | positive | MGI_C57BL6J_1922990 | Egfem1 | NCBI_Gene:75740,ENSEMBL:ENSMUSG00000063600 | MGI:1922990 | protein coding gene | EGF-like and EMI domain containing 1 |
| 3 | gene | 29.17484 | 29.17693 | positive | MGI_C57BL6J_5611257 | Gm38029 | ENSEMBL:ENSMUSG00000103992 | MGI:5611257 | unclassified gene | predicted gene, 38029 |
| 3 | gene | 29.20934 | 29.21198 | positive | MGI_C57BL6J_2442122 | D230021J17Rik | NA | NA | unclassified gene | RIKEN cDNA D230021J17 gene |
| 3 | gene | 29.21989 | 29.22125 | positive | MGI_C57BL6J_5611193 | Gm37965 | ENSEMBL:ENSMUSG00000102579 | MGI:5611193 | unclassified gene | predicted gene, 37965 |
| 3 | gene | 29.26628 | 29.26684 | negative | MGI_C57BL6J_2139582 | AA516838 | NA | NA | unclassified gene | expressed sequence AA516838 |
| 3 | gene | 29.41682 | 29.41692 | positive | MGI_C57BL6J_3691605 | Mir551b | miRBase:MI0004131,NCBI_Gene:791072,ENSEMBL:ENSMUSG00000076120 | MGI:3691605 | miRNA gene | microRNA 551b |
| 3 | gene | 29.63674 | 29.63685 | positive | MGI_C57BL6J_5531333 | Mir21b | miRBase:MI0021906,NCBI_Gene:102466637,ENSEMBL:ENSMUSG00000098315 | MGI:5531333 | miRNA gene | microRNA 21b |
| 3 | gene | 29.66251 | 29.69656 | negative | MGI_C57BL6J_5625082 | Gm42197 | NCBI_Gene:105247013 | MGI:5625082 | lncRNA gene | predicted gene, 42197 |
| 3 | gene | 29.76404 | 29.89323 | negative | MGI_C57BL6J_5592365 | Gm33206 | NCBI_Gene:102636014,ENSEMBL:ENSMUSG00000102574 | MGI:5592365 | lncRNA gene | predicted gene, 33206 |
| 3 | pseudogene | 29.77128 | 29.77142 | positive | MGI_C57BL6J_5610785 | Gm37557 | ENSEMBL:ENSMUSG00000103095 | MGI:5610785 | pseudogene | predicted gene, 37557 |
| 3 | gene | 29.89101 | 29.92419 | positive | MGI_C57BL6J_5564803 | Mannr | NCBI_Gene:102636083,ENSEMBL:ENSMUSG00000102590 | MGI:5564803 | lncRNA gene | Mecom adjacent non-protein coding RNA |
| 3 | pseudogene | 29.92654 | 29.92697 | positive | MGI_C57BL6J_5610509 | Gm37281 | ENSEMBL:ENSMUSG00000102884 | MGI:5610509 | pseudogene | predicted gene, 37281 |
| 3 | gene | 29.95130 | 30.54801 | negative | MGI_C57BL6J_95457 | Mecom | NCBI_Gene:14013,ENSEMBL:ENSMUSG00000027684 | MGI:95457 | protein coding gene | MDS1 and EVI1 complex locus |
| 3 | gene | 30.01238 | 30.05481 | positive | MGI_C57BL6J_4439646 | Mecomos | ENSEMBL:ENSMUSG00000085702 | MGI:4439646 | antisense lncRNA gene | MDS1 and EVI1 complex locus, opposite strand |
| 3 | gene | 30.16025 | 30.16189 | negative | MGI_C57BL6J_5611294 | Gm38066 | ENSEMBL:ENSMUSG00000104486 | MGI:5611294 | unclassified gene | predicted gene, 38066 |
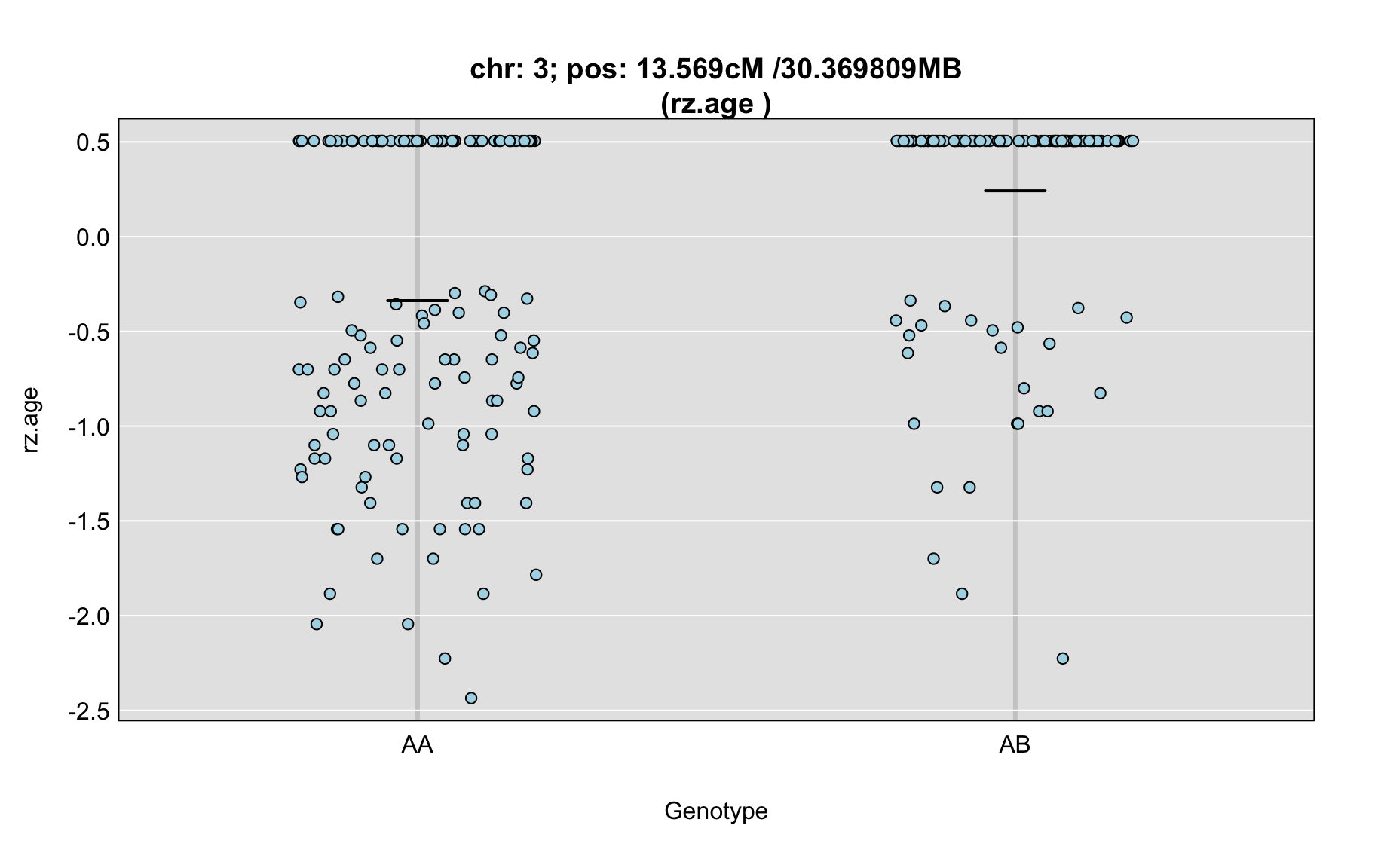
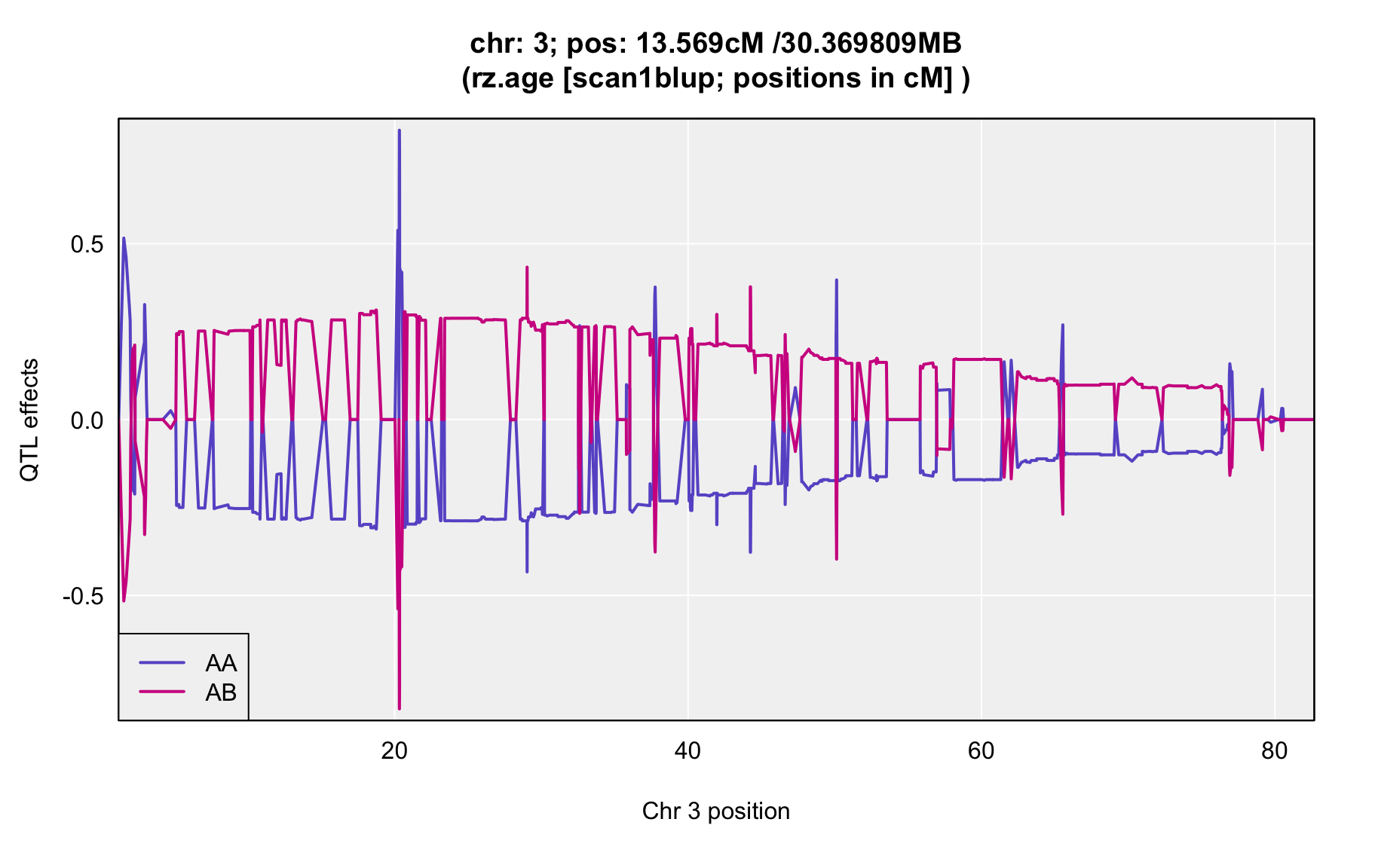
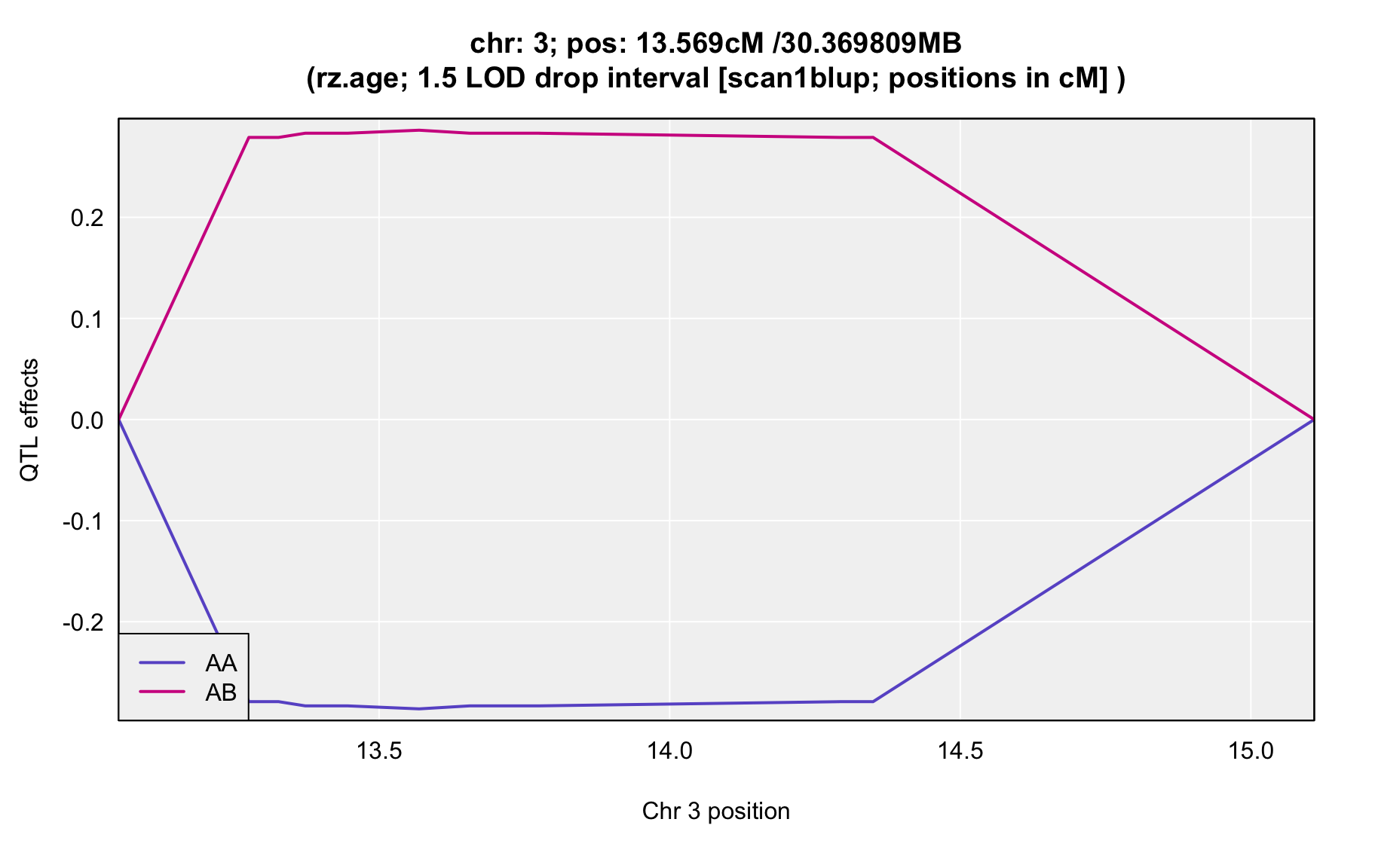
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 29.95130 | 30.54801 | negative | MGI_C57BL6J_95457 | Mecom | NCBI_Gene:14013,ENSEMBL:ENSMUSG00000027684 | MGI:95457 | protein coding gene | MDS1 and EVI1 complex locus |
| 3 | gene | 30.22557 | 30.22769 | negative | MGI_C57BL6J_5611425 | Gm38197 | ENSEMBL:ENSMUSG00000102460 | MGI:5611425 | unclassified gene | predicted gene, 38197 |
| 3 | gene | 30.26684 | 30.28019 | negative | MGI_C57BL6J_3648282 | Gm10258 | NCBI_Gene:545510,ENSEMBL:ENSMUSG00000069074 | MGI:3648282 | unclassified non-coding RNA gene | predicted gene 10258 |
| 3 | gene | 30.27232 | 30.27438 | negative | MGI_C57BL6J_5611590 | Gm38362 | ENSEMBL:ENSMUSG00000103870 | MGI:5611590 | unclassified gene | predicted gene, 38362 |
| 3 | gene | 30.34998 | 30.35132 | negative | MGI_C57BL6J_1924752 | 9530010C24Rik | NA | NA | unclassified gene | RIKEN cDNA 9530010C24 gene |
| 3 | gene | 30.48843 | 30.48929 | positive | MGI_C57BL6J_5610252 | Gm37024 | ENSEMBL:ENSMUSG00000104079 | MGI:5610252 | unclassified gene | predicted gene, 37024 |
| 3 | gene | 30.50172 | 30.50381 | negative | MGI_C57BL6J_5610880 | Gm37652 | ENSEMBL:ENSMUSG00000103738 | MGI:5610880 | unclassified gene | predicted gene, 37652 |
| 3 | gene | 30.55270 | 30.56424 | negative | MGI_C57BL6J_5611486 | Gm38258 | ENSEMBL:ENSMUSG00000102256 | MGI:5611486 | lncRNA gene | predicted gene, 38258 |
| 3 | pseudogene | 30.56166 | 30.56213 | negative | MGI_C57BL6J_5611381 | Gm38153 | ENSEMBL:ENSMUSG00000104243 | MGI:5611381 | pseudogene | predicted gene, 38153 |
| 3 | gene | 30.59484 | 30.59576 | negative | MGI_C57BL6J_1920499 | 1600017P15Rik | ENSEMBL:ENSMUSG00000102505,ENSEMBL:ENSMUSG00000065361 | MGI:1920499 | unclassified gene | RIKEN cDNA 1600017P15 gene |
| 3 | gene | 30.59707 | 30.59994 | negative | MGI_C57BL6J_1923902 | Actrt3 | NCBI_Gene:76652,ENSEMBL:ENSMUSG00000037737 | MGI:1923902 | protein coding gene | actin related protein T3 |
| 3 | gene | 30.60127 | 30.60134 | positive | MGI_C57BL6J_4414091 | n-TVaac8 | NCBI_Gene:102467402 | MGI:4414091 | tRNA gene | nuclear encoded tRNA valine 8 (anticodon AAC) |
| 3 | gene | 30.60206 | 30.61987 | positive | MGI_C57BL6J_1931415 | Mynn | NCBI_Gene:80732,ENSEMBL:ENSMUSG00000037730 | MGI:1931415 | protein coding gene | myoneurin |
| 3 | gene | 30.62427 | 30.64787 | negative | MGI_C57BL6J_1919077 | Lrrc34 | NCBI_Gene:71827,ENSEMBL:ENSMUSG00000027702 | MGI:1919077 | protein coding gene | leucine rich repeat containing 34 |
| 3 | gene | 30.64451 | 30.67243 | positive | MGI_C57BL6J_1915557 | Lrriq4 | NCBI_Gene:68307,ENSEMBL:ENSMUSG00000027703 | MGI:1915557 | protein coding gene | leucine-rich repeats and IQ motif containing 4 |
| 3 | gene | 30.67855 | 30.69984 | negative | MGI_C57BL6J_2443864 | Lrrc31 | NCBI_Gene:320352,ENSEMBL:ENSMUSG00000074653 | MGI:2443864 | protein coding gene | leucine rich repeat containing 31 |
| 3 | gene | 30.68594 | 30.69301 | positive | MGI_C57BL6J_3705271 | Gm15462 | NCBI_Gene:108168861,ENSEMBL:ENSMUSG00000086189 | MGI:3705271 | lncRNA gene | predicted gene 15462 |
| 3 | gene | 30.74629 | 30.76717 | positive | MGI_C57BL6J_1923203 | Samd7 | NCBI_Gene:75953,ENSEMBL:ENSMUSG00000051860 | MGI:1923203 | protein coding gene | sterile alpha motif domain containing 7 |
| 3 | gene | 30.75231 | 30.75665 | negative | MGI_C57BL6J_5592749 | Gm33590 | NCBI_Gene:102636556 | MGI:5592749 | lncRNA gene | predicted gene, 33590 |
| 3 | gene | 30.79257 | 30.79356 | negative | MGI_C57BL6J_1918530 | 4933429H19Rik | ENSEMBL:ENSMUSG00000097786 | MGI:1918530 | lncRNA gene | RIKEN cDNA 4933429H19 gene |
| 3 | gene | 30.79276 | 30.82126 | positive | MGI_C57BL6J_1916526 | Sec62 | NCBI_Gene:69276,ENSEMBL:ENSMUSG00000027706 | MGI:1916526 | protein coding gene | SEC62 homolog (S. cerevisiae) |
| 3 | pseudogene | 30.82857 | 30.82883 | negative | MGI_C57BL6J_5611308 | Gm38080 | ENSEMBL:ENSMUSG00000104361 | MGI:5611308 | pseudogene | predicted gene, 38080 |
| 3 | gene | 30.84154 | 30.85199 | negative | MGI_C57BL6J_5625083 | Gm42198 | NCBI_Gene:105247015 | MGI:5625083 | lncRNA gene | predicted gene, 42198 |
| 3 | gene | 30.85248 | 30.85586 | negative | MGI_C57BL6J_5625084 | Gm42199 | NCBI_Gene:105247016 | MGI:5625084 | lncRNA gene | predicted gene, 42199 |
| 3 | gene | 30.85587 | 30.89719 | positive | MGI_C57BL6J_1919112 | Gpr160 | NCBI_Gene:71862,ENSEMBL:ENSMUSG00000037661 | MGI:1919112 | protein coding gene | G protein-coupled receptor 160 |
| 3 | gene | 30.89929 | 30.96948 | negative | MGI_C57BL6J_2181434 | Phc3 | NCBI_Gene:241915,ENSEMBL:ENSMUSG00000037652 | MGI:2181434 | protein coding gene | polyhomeotic 3 |
| 3 | gene | 30.95733 | 30.95764 | negative | MGI_C57BL6J_1924590 | 9530022L04Rik | ENSEMBL:ENSMUSG00000102369 | MGI:1924590 | unclassified gene | RIKEN cDNA 9530022L04 gene |
| 3 | pseudogene | 30.97291 | 30.97511 | positive | MGI_C57BL6J_3781157 | Gm2979 | NCBI_Gene:100040809,ENSEMBL:ENSMUSG00000102741 | MGI:3781157 | pseudogene | predicted gene 2979 |
| 3 | gene | 30.99574 | 31.05296 | positive | MGI_C57BL6J_99260 | Prkci | NCBI_Gene:18759,ENSEMBL:ENSMUSG00000037643 | MGI:99260 | protein coding gene | protein kinase C, iota |
| 3 | gene | 31.03897 | 31.03903 | positive | MGI_C57BL6J_5531134 | Mir7008 | miRBase:MI0022857,NCBI_Gene:102465610,ENSEMBL:ENSMUSG00000098890 | MGI:5531134 | miRNA gene | microRNA 7008 |
| 3 | gene | 31.06130 | 31.06669 | positive | MGI_C57BL6J_5592800 | Gm33641 | NCBI_Gene:102636630 | MGI:5592800 | lncRNA gene | predicted gene, 33641 |
| 3 | gene | 31.06243 | 31.07994 | negative | MGI_C57BL6J_5592853 | Gm33694 | NCBI_Gene:102636694 | MGI:5592853 | lncRNA gene | predicted gene, 33694 |
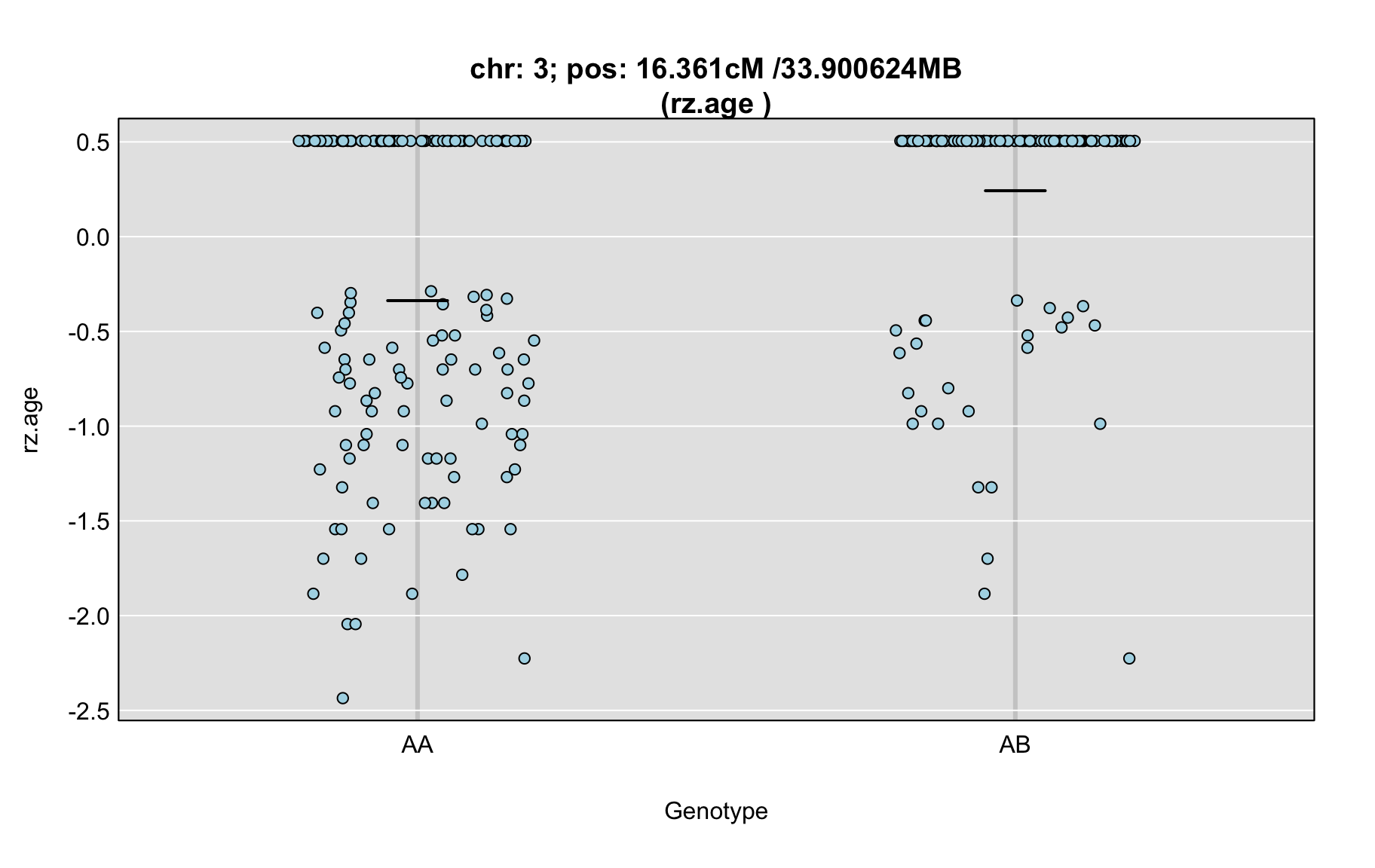

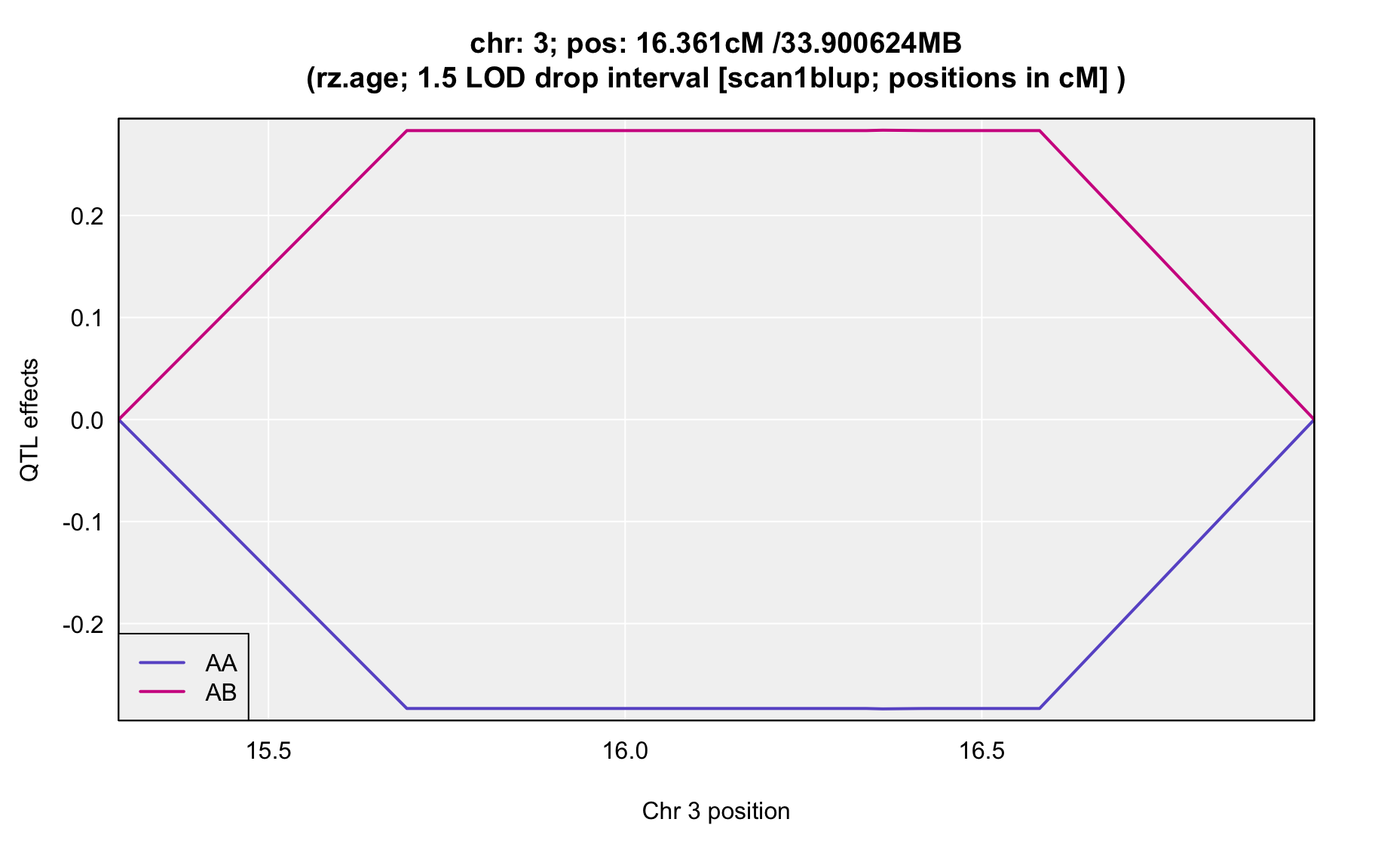
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | pseudogene | 31.64480 | 31.69058 | negative | MGI_C57BL6J_3779607 | Gm6558 | NCBI_Gene:625125,ENSEMBL:ENSMUSG00000103499 | MGI:3779607 | pseudogene | predicted gene 6558 |
| 3 | gene | 31.68642 | 31.69214 | positive | MGI_C57BL6J_5593045 | Gm33886 | NCBI_Gene:102636961 | MGI:5593045 | lncRNA gene | predicted gene, 33886 |
| 3 | gene | 31.89883 | 31.90212 | negative | MGI_C57BL6J_5593190 | Gm34031 | NCBI_Gene:102637147 | MGI:5593190 | lncRNA gene | predicted gene, 34031 |
| 3 | gene | 31.90213 | 32.20018 | positive | MGI_C57BL6J_1919663 | Kcnmb2 | NCBI_Gene:72413,ENSEMBL:ENSMUSG00000037610 | MGI:1919663 | protein coding gene | potassium large conductance calcium-activated channel, subfamily M, beta member 2 |
| 3 | gene | 31.95476 | 31.95780 | positive | MGI_C57BL6J_5611062 | Gm37834 | ENSEMBL:ENSMUSG00000103539 | MGI:5611062 | unclassified gene | predicted gene, 37834 |
| 3 | gene | 32.23753 | 32.32631 | negative | MGI_C57BL6J_5625086 | Gm42201 | NCBI_Gene:105247019 | MGI:5625086 | lncRNA gene | predicted gene, 42201 |
| 3 | pseudogene | 32.26033 | 32.26111 | positive | MGI_C57BL6J_5610998 | Gm37770 | NCBI_Gene:108168862,ENSEMBL:ENSMUSG00000103643 | MGI:5610998 | pseudogene | predicted gene, 37770 |
| 3 | gene | 32.33479 | 32.36601 | negative | MGI_C57BL6J_1195270 | Zmat3 | NCBI_Gene:22401,ENSEMBL:ENSMUSG00000027663 | MGI:1195270 | protein coding gene | zinc finger matrin type 3 |
| 3 | gene | 32.36549 | 32.36744 | positive | MGI_C57BL6J_1914826 | 4930429B21Rik | NCBI_Gene:67576 | MGI:1914826 | lncRNA gene | RIKEN cDNA 4930429B21 gene |
| 3 | gene | 32.37158 | 32.37299 | negative | MGI_C57BL6J_5610218 | Gm36990 | ENSEMBL:ENSMUSG00000102825 | MGI:5610218 | unclassified gene | predicted gene, 36990 |
| 3 | gene | 32.37898 | 32.38470 | negative | MGI_C57BL6J_5625087 | Gm42202 | NCBI_Gene:105247020 | MGI:5625087 | lncRNA gene | predicted gene, 42202 |
| 3 | gene | 32.39707 | 32.46849 | positive | MGI_C57BL6J_1206581 | Pik3ca | NCBI_Gene:18706,ENSEMBL:ENSMUSG00000027665 | MGI:1206581 | protein coding gene | phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha |
| 3 | gene | 32.45459 | 32.45576 | positive | MGI_C57BL6J_5610351 | Gm37123 | ENSEMBL:ENSMUSG00000103039 | MGI:5610351 | unclassified gene | predicted gene, 37123 |
| 3 | gene | 32.46778 | 32.50290 | negative | MGI_C57BL6J_3612244 | Kcnmb3 | NCBI_Gene:100502876,ENSEMBL:ENSMUSG00000091091 | MGI:3612244 | protein coding gene | potassium large conductance calcium-activated channel, subfamily M, beta member 3 |
| 3 | gene | 32.51026 | 32.52083 | positive | MGI_C57BL6J_1915028 | Zfp639 | NCBI_Gene:67778,ENSEMBL:ENSMUSG00000027667 | MGI:1915028 | protein coding gene | zinc finger protein 639 |
| 3 | gene | 32.51180 | 32.51234 | negative | MGI_C57BL6J_5610820 | Gm37592 | ENSEMBL:ENSMUSG00000104231 | MGI:5610820 | lncRNA gene | predicted gene, 37592 |
| 3 | gene | 32.52946 | 32.57924 | positive | MGI_C57BL6J_1914664 | Mfn1 | NCBI_Gene:67414,ENSEMBL:ENSMUSG00000027668 | MGI:1914664 | protein coding gene | mitofusin 1 |
| 3 | gene | 32.58033 | 32.61659 | negative | MGI_C57BL6J_104581 | Gnb4 | NCBI_Gene:14696,ENSEMBL:ENSMUSG00000027669 | MGI:104581 | protein coding gene | guanine nucleotide binding protein (G protein), beta 4 |
| 3 | gene | 32.61086 | 32.61351 | positive | MGI_C57BL6J_5610868 | Gm37640 | ENSEMBL:ENSMUSG00000102862 | MGI:5610868 | lncRNA gene | predicted gene, 37640 |
| 3 | gene | 32.70630 | 32.72697 | positive | MGI_C57BL6J_1861453 | Actl6a | NCBI_Gene:56456,ENSEMBL:ENSMUSG00000027671 | MGI:1861453 | protein coding gene | actin-like 6A |
| 3 | gene | 32.72178 | 32.72209 | negative | MGI_C57BL6J_5610581 | Gm37353 | ENSEMBL:ENSMUSG00000103590 | MGI:5610581 | unclassified gene | predicted gene, 37353 |
| 3 | gene | 32.72540 | 32.73761 | negative | MGI_C57BL6J_1921850 | Mrpl47 | NCBI_Gene:74600,ENSEMBL:ENSMUSG00000037531 | MGI:1921850 | protein coding gene | mitochondrial ribosomal protein L47 |
| 3 | gene | 32.73610 | 32.73761 | negative | MGI_C57BL6J_3704212 | Gm9859 | NA | NA | unclassified gene | predicted gene 9859 |
| 3 | gene | 32.73699 | 32.75157 | positive | MGI_C57BL6J_1913296 | Ndufb5 | NCBI_Gene:66046,ENSEMBL:ENSMUSG00000027673 | MGI:1913296 | protein coding gene | NADH:ubiquinone oxidoreductase subunit B5 |
| 3 | gene | 32.81702 | 32.93807 | positive | MGI_C57BL6J_1919857 | Usp13 | NCBI_Gene:72607,ENSEMBL:ENSMUSG00000056900 | MGI:1919857 | protein coding gene | ubiquitin specific peptidase 13 (isopeptidase T-3) |
| 3 | pseudogene | 32.88001 | 32.88061 | negative | MGI_C57BL6J_3783022 | Gm15574 | ENSEMBL:ENSMUSG00000081883 | MGI:3783022 | pseudogene | predicted gene 15574 |
| 3 | gene | 32.93931 | 32.98330 | positive | MGI_C57BL6J_5610687 | Gm37459 | ENSEMBL:ENSMUSG00000104283 | MGI:5610687 | lncRNA gene | predicted gene, 37459 |
| 3 | gene | 32.94793 | 33.14327 | negative | MGI_C57BL6J_1916672 | Pex5l | NCBI_Gene:58869,ENSEMBL:ENSMUSG00000027674 | MGI:1916672 | protein coding gene | peroxisomal biogenesis factor 5-like |
| 3 | gene | 33.01196 | 33.01211 | positive | MGI_C57BL6J_5452991 | Gm23214 | ENSEMBL:ENSMUSG00000084665 | MGI:5452991 | snRNA gene | predicted gene, 23214 |
| 3 | gene | 33.02478 | 33.08861 | positive | MGI_C57BL6J_1921124 | 4930419G24Rik | NCBI_Gene:73874,ENSEMBL:ENSMUSG00000087625 | MGI:1921124 | lncRNA gene | RIKEN cDNA 4930419G24 gene |
| 3 | gene | 33.03423 | 33.03705 | negative | MGI_C57BL6J_5610671 | Gm37443 | ENSEMBL:ENSMUSG00000102581 | MGI:5610671 | unclassified gene | predicted gene, 37443 |
| 3 | gene | 33.07392 | 33.07636 | negative | MGI_C57BL6J_5011630 | Gm19445 | ENSEMBL:ENSMUSG00000102498 | MGI:5011630 | unclassified gene | predicted gene, 19445 |
| 3 | pseudogene | 33.19080 | 33.19104 | positive | MGI_C57BL6J_5610356 | Gm37128 | ENSEMBL:ENSMUSG00000103896 | MGI:5610356 | pseudogene | predicted gene, 37128 |
| 3 | gene | 33.26072 | 33.26082 | negative | MGI_C57BL6J_5531356 | Gm27974 | ENSEMBL:ENSMUSG00000098622 | MGI:5531356 | miRNA gene | predicted gene, 27974 |
| 3 | gene | 33.28312 | 33.28565 | positive | MGI_C57BL6J_5610259 | Gm37031 | ENSEMBL:ENSMUSG00000104384 | MGI:5610259 | unclassified gene | predicted gene, 37031 |
| 3 | gene | 33.39306 | 33.39320 | negative | MGI_C57BL6J_5454557 | Gm24780 | ENSEMBL:ENSMUSG00000088260 | MGI:5454557 | snoRNA gene | predicted gene, 24780 |
| 3 | gene | 33.39338 | 33.39485 | negative | MGI_C57BL6J_5610626 | Gm37398 | ENSEMBL:ENSMUSG00000104203 | MGI:5610626 | unclassified gene | predicted gene, 37398 |
| 3 | gene | 33.54027 | 33.54342 | positive | MGI_C57BL6J_5593364 | Gm34205 | NCBI_Gene:102637377 | MGI:5593364 | lncRNA gene | predicted gene, 34205 |
| 3 | pseudogene | 33.62159 | 33.62245 | positive | MGI_C57BL6J_3649050 | Gm5845 | NCBI_Gene:545512,ENSEMBL:ENSMUSG00000103423 | MGI:3649050 | pseudogene | predicted gene 5845 |
| 3 | pseudogene | 33.68451 | 33.68487 | negative | MGI_C57BL6J_5662831 | Gm42694 | ENSEMBL:ENSMUSG00000104954 | MGI:5662831 | pseudogene | predicted gene 42694 |
| 3 | gene | 33.68696 | 33.69200 | negative | MGI_C57BL6J_5625088 | Gm42203 | NCBI_Gene:105247021 | MGI:5625088 | lncRNA gene | predicted gene, 42203 |
| 3 | gene | 33.70336 | 33.75962 | negative | MGI_C57BL6J_5663803 | Gm43666 | ENSEMBL:ENSMUSG00000104588 | MGI:5663803 | lncRNA gene | predicted gene 43666 |
| 3 | pseudogene | 33.73060 | 33.73353 | positive | MGI_C57BL6J_5010796 | Gm18611 | NCBI_Gene:100417436,ENSEMBL:ENSMUSG00000106231 | MGI:5010796 | pseudogene | predicted gene, 18611 |
| 3 | gene | 33.79983 | 33.81486 | positive | MGI_C57BL6J_1914370 | Ttc14 | NCBI_Gene:67120,ENSEMBL:ENSMUSG00000027677 | MGI:1914370 | protein coding gene | tetratricopeptide repeat domain 14 |
| 3 | gene | 33.81076 | 33.84432 | negative | MGI_C57BL6J_1289263 | Ccdc39 | NCBI_Gene:51938,ENSEMBL:ENSMUSG00000027676 | MGI:1289263 | protein coding gene | coiled-coil domain containing 39 |
| 3 | gene | 33.81726 | 33.81795 | positive | MGI_C57BL6J_5663277 | Gm43140 | ENSEMBL:ENSMUSG00000105286 | MGI:5663277 | lncRNA gene | predicted gene 43140 |
| 3 | gene | 33.86681 | 33.89758 | negative | MGI_C57BL6J_1921204 | 4930431L21Rik | ENSEMBL:ENSMUSG00000105475 | MGI:1921204 | lncRNA gene | RIKEN cDNA 4930431L21 gene |
| 3 | gene | 33.90684 | 33.90823 | positive | MGI_C57BL6J_5625089 | Gm42204 | NCBI_Gene:105247022 | MGI:5625089 | lncRNA gene | predicted gene, 42204 |
| 3 | gene | 33.95620 | 33.95715 | positive | MGI_C57BL6J_1922225 | 4930502C17Rik | ENSEMBL:ENSMUSG00000106584 | MGI:1922225 | unclassified gene | RIKEN cDNA 4930502C17 gene |
| 3 | gene | 33.95907 | 33.95920 | negative | MGI_C57BL6J_5452952 | Gm23175 | ENSEMBL:ENSMUSG00000077559 | MGI:5452952 | snoRNA gene | predicted gene, 23175 |
| 3 | gene | 33.97218 | 33.97545 | negative | MGI_C57BL6J_5826470 | Gm46833 | NCBI_Gene:108168913 | MGI:5826470 | lncRNA gene | predicted gene, 46833 |
| 3 | pseudogene | 34.00477 | 34.00551 | negative | MGI_C57BL6J_3642317 | Gm9791 | NCBI_Gene:100040232,ENSEMBL:ENSMUSG00000044434 | MGI:3642317 | pseudogene | predicted pseudogene 9791 |
| 3 | gene | 34.00880 | 34.01188 | negative | MGI_C57BL6J_5826435 | Gm46798 | NCBI_Gene:108168863 | MGI:5826435 | lncRNA gene | predicted gene, 46798 |
| 3 | gene | 34.01771 | 34.01988 | negative | MGI_C57BL6J_5663214 | Gm43077 | ENSEMBL:ENSMUSG00000105535 | MGI:5663214 | lncRNA gene | predicted gene 43077 |
| 3 | gene | 34.01994 | 34.07032 | positive | MGI_C57BL6J_104860 | Fxr1 | NCBI_Gene:14359,ENSEMBL:ENSMUSG00000027680 | MGI:104860 | protein coding gene | fragile X mental retardation gene 1, autosomal homolog |
| 3 | gene | 34.05434 | 34.05543 | negative | MGI_C57BL6J_5663804 | Gm43667 | ENSEMBL:ENSMUSG00000105135 | MGI:5663804 | unclassified gene | predicted gene 43667 |
| 3 | gene | 34.05602 | 34.08135 | negative | MGI_C57BL6J_1914963 | Dnajc19 | NCBI_Gene:67713,ENSEMBL:ENSMUSG00000027679 | MGI:1914963 | protein coding gene | DnaJ heat shock protein family (Hsp40) member C19 |
| 3 | gene | 34.07066 | 34.07600 | positive | MGI_C57BL6J_5663805 | Gm43668 | ENSEMBL:ENSMUSG00000105176 | MGI:5663805 | unclassified gene | predicted gene 43668 |
| 3 | gene | 34.10427 | 34.68262 | positive | MGI_C57BL6J_2444112 | Sox2ot | NCBI_Gene:320478,ENSEMBL:ENSMUSG00000105265 | MGI:2444112 | lncRNA gene | SOX2 overlapping transcript (non-protein coding) |
| 3 | pseudogene | 34.18331 | 34.18419 | positive | MGI_C57BL6J_3647478 | Gm5708 | NCBI_Gene:435727,ENSEMBL:ENSMUSG00000105739 | MGI:3647478 | pseudogene | predicted gene 5708 |
| 3 | gene | 34.28133 | 34.35199 | negative | MGI_C57BL6J_5621390 | Gm38505 | NCBI_Gene:102638008,ENSEMBL:ENSMUSG00000092196 | MGI:5621390 | lncRNA gene | predicted gene, 38505 |
| 3 | gene | 34.41544 | 34.41816 | positive | MGI_C57BL6J_5593758 | Gm34599 | NCBI_Gene:102637903,ENSEMBL:ENSMUSG00000105577 | MGI:5593758 | lncRNA gene | predicted gene, 34599 |
| 3 | gene | 34.42737 | 34.43510 | negative | MGI_C57BL6J_5141980 | Gm20515 | ENSEMBL:ENSMUSG00000092242 | MGI:5141980 | lncRNA gene | predicted gene 20515 |
| 3 | gene | 34.48143 | 34.48221 | negative | MGI_C57BL6J_5579841 | Gm29135 | ENSEMBL:ENSMUSG00000099437 | MGI:5579841 | lncRNA gene | predicted gene 29135 |
| 3 | gene | 34.56034 | 34.56055 | positive | MGI_C57BL6J_5530785 | Gm27403 | ENSEMBL:ENSMUSG00000098795 | MGI:5530785 | unclassified non-coding RNA gene | predicted gene, 27403 |
| 3 | gene | 34.56526 | 34.57220 | negative | MGI_C57BL6J_5662832 | Gm42695 | ENSEMBL:ENSMUSG00000106525 | MGI:5662832 | lncRNA gene | predicted gene 42695 |
| 3 | gene | 34.59782 | 34.60488 | negative | MGI_C57BL6J_5625090 | Gm42205 | NCBI_Gene:105247023,ENSEMBL:ENSMUSG00000105222 | MGI:5625090 | lncRNA gene | predicted gene, 42205 |
| 3 | gene | 34.63846 | 34.63854 | positive | MGI_C57BL6J_3811407 | Mir1897 | miRBase:MI0008314,NCBI_Gene:100316679,ENSEMBL:ENSMUSG00000104797 | MGI:3811407 | miRNA gene | microRNA 1897 |
| 3 | gene | 34.64326 | 34.64610 | positive | MGI_C57BL6J_5662829 | Gm42692 | ENSEMBL:ENSMUSG00000104901 | MGI:5662829 | unclassified gene | predicted gene 42692 |
| 3 | gene | 34.64999 | 34.65246 | positive | MGI_C57BL6J_98364 | Sox2 | NCBI_Gene:20674,ENSEMBL:ENSMUSG00000074637 | MGI:98364 | protein coding gene | SRY (sex determining region Y)-box 2 |
| 3 | gene | 34.65318 | 34.65328 | positive | MGI_C57BL6J_5530993 | Gm27611 | ENSEMBL:ENSMUSG00000098254 | MGI:5530993 | unclassified non-coding RNA gene | predicted gene, 27611 |
| 3 | gene | 34.66366 | 34.66429 | negative | MGI_C57BL6J_5662830 | Gm42693 | ENSEMBL:ENSMUSG00000105284 | MGI:5662830 | unclassified gene | predicted gene 42693 |
| 3 | gene | 34.68263 | 34.68275 | positive | MGI_C57BL6J_5663343 | Gm43206 | ENSEMBL:ENSMUSG00000105101 | MGI:5663343 | unclassified gene | predicted gene 43206 |
| 3 | gene | 34.68938 | 34.69292 | positive | MGI_C57BL6J_5594100 | Gm34941 | NCBI_Gene:102638356 | MGI:5594100 | lncRNA gene | predicted gene, 34941 |
| 3 | gene | 34.69964 | 34.71666 | negative | MGI_C57BL6J_3781322 | Gm3143 | NCBI_Gene:100041108,ENSEMBL:ENSMUSG00000105153 | MGI:3781322 | lncRNA gene | predicted gene 3143 |
| 3 | gene | 34.77208 | 34.78235 | positive | MGI_C57BL6J_5621394 | Gm38509 | NCBI_Gene:102638436,ENSEMBL:ENSMUSG00000106519 | MGI:5621394 | lncRNA gene | predicted gene, 38509 |
| 3 | gene | 34.82656 | 34.87928 | positive | MGI_C57BL6J_5434743 | Gm21388 | NCBI_Gene:100861998,ENSEMBL:ENSMUSG00000114832 | MGI:5434743 | lncRNA gene | predicted gene, 21388 |
| 3 | gene | 34.92254 | 34.92265 | negative | MGI_C57BL6J_5531088 | Mir6378 | miRBase:MI0021909,NCBI_Gene:102466155,ENSEMBL:ENSMUSG00000098862 | MGI:5531088 | miRNA gene | microRNA 6378 |
| 3 | gene | 34.95023 | 34.95953 | positive | MGI_C57BL6J_5594414 | Gm35255 | NCBI_Gene:102638771 | MGI:5594414 | lncRNA gene | predicted gene, 35255 |
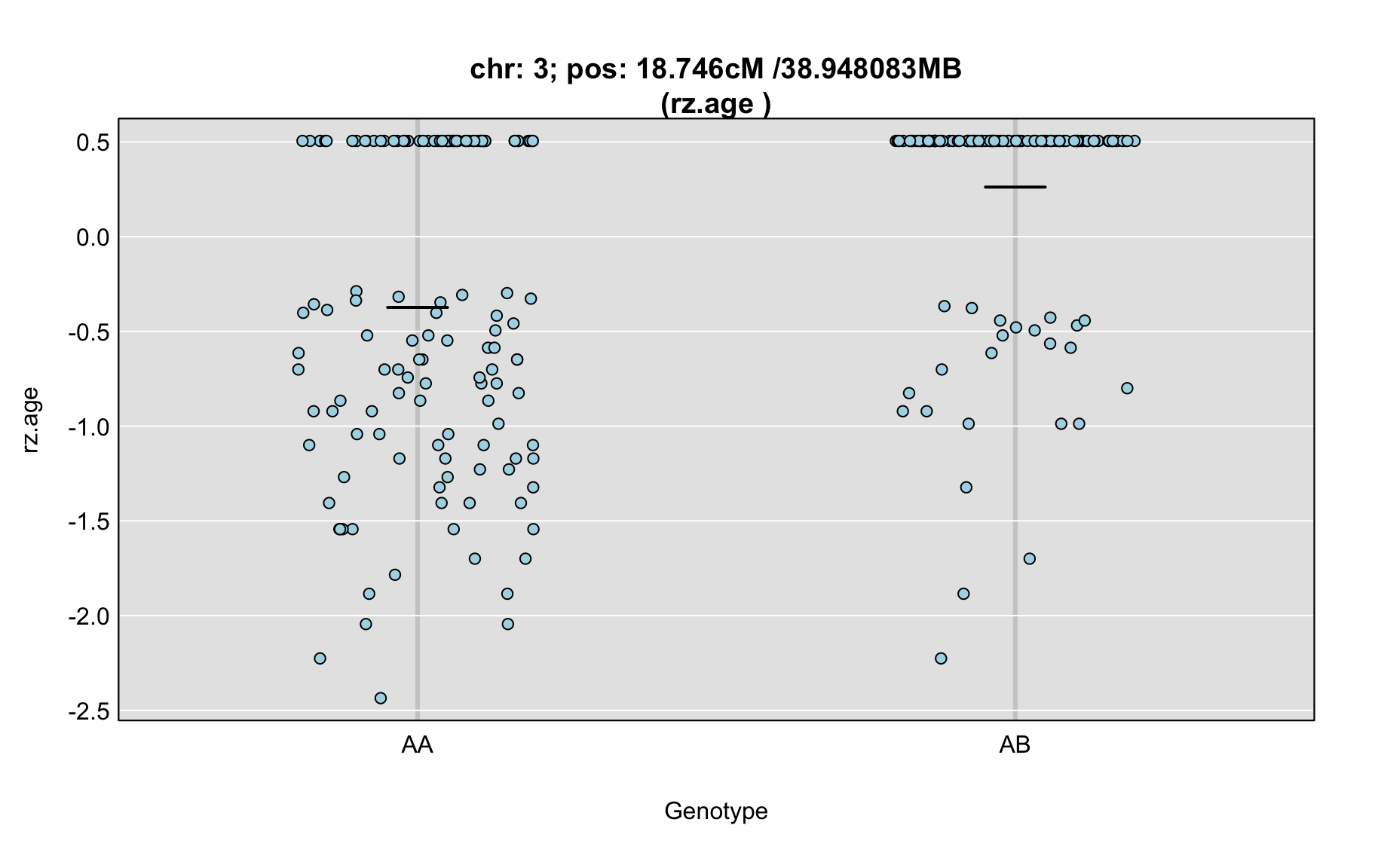
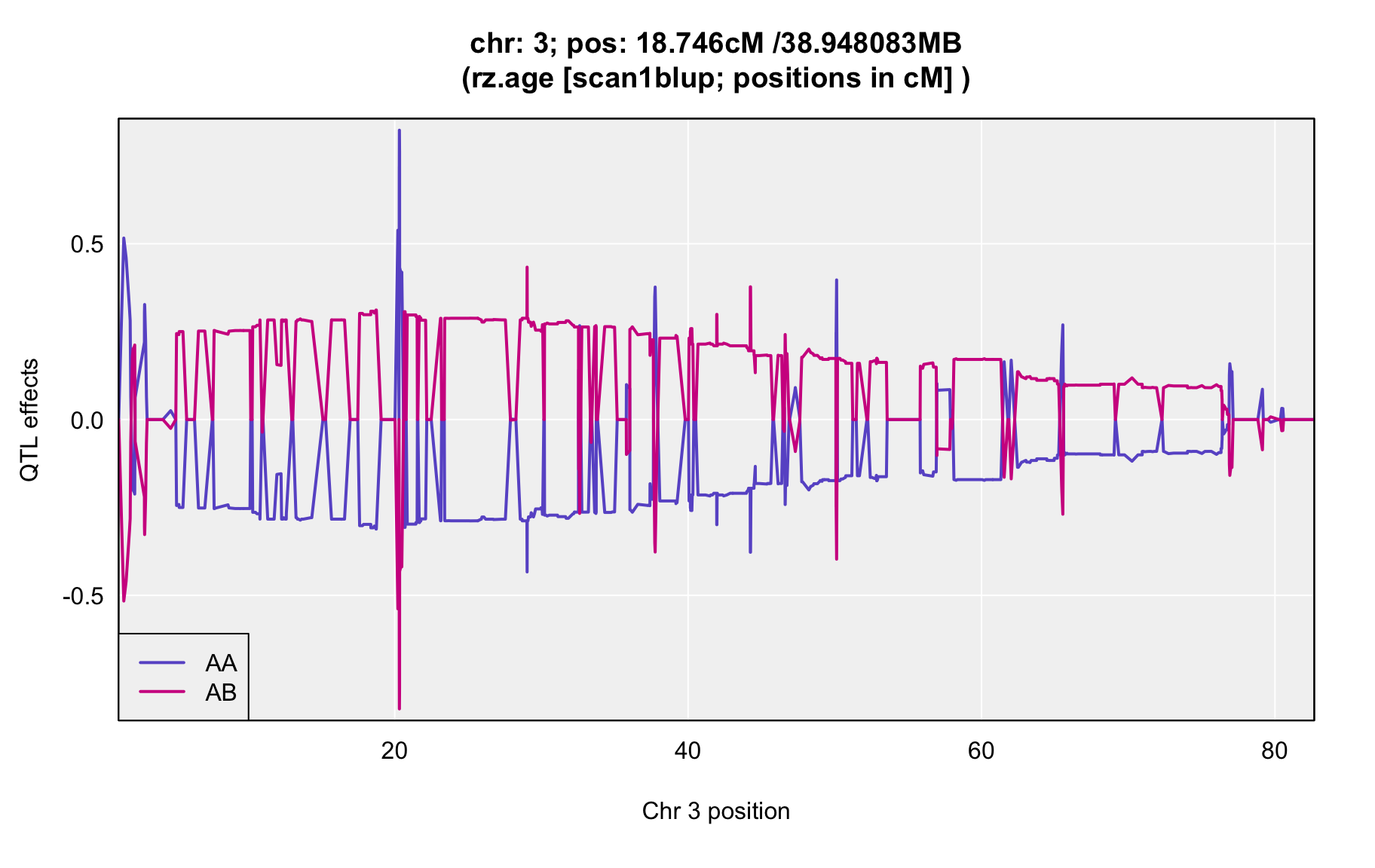
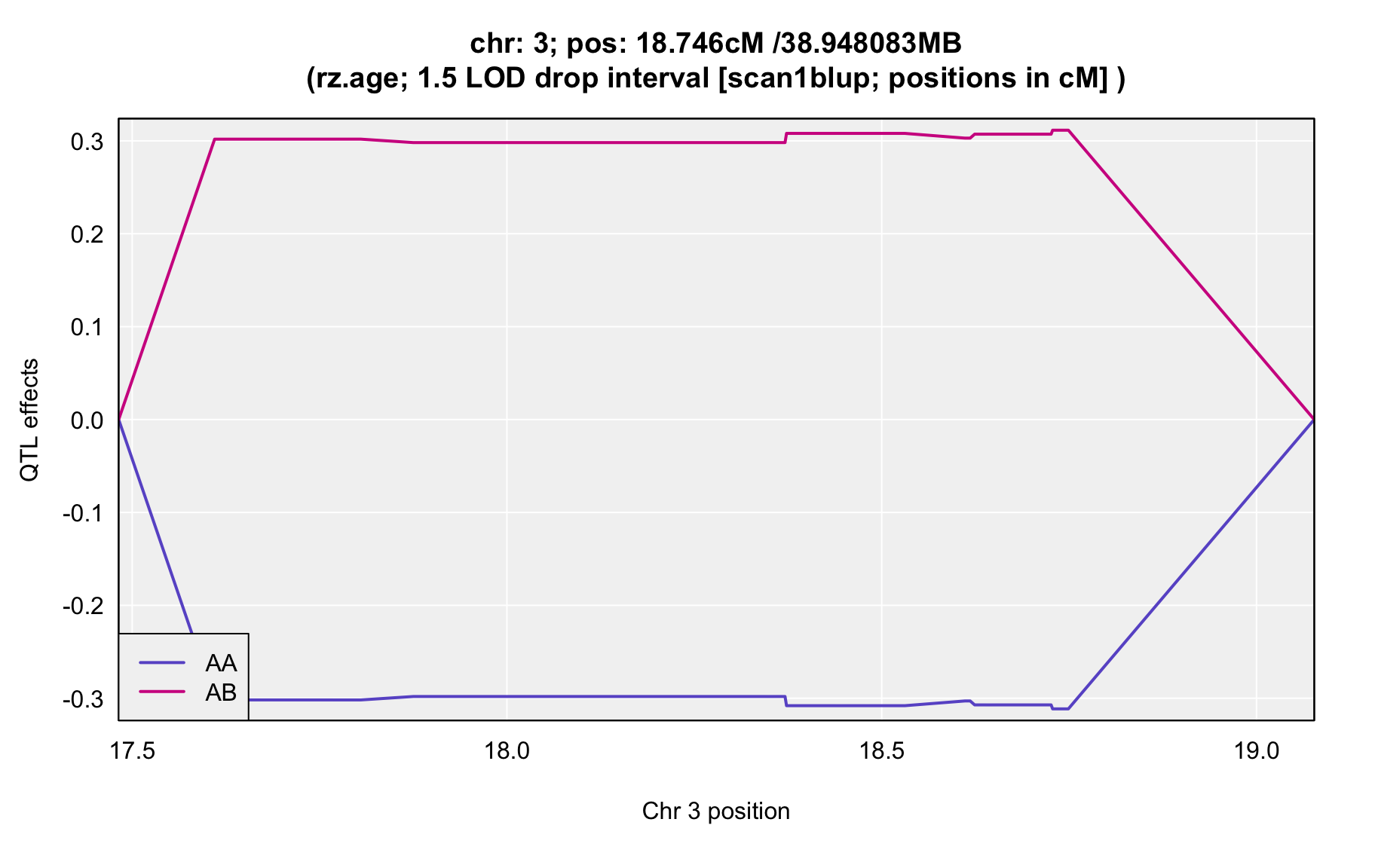
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 36.52922 | 36.53233 | negative | MGI_C57BL6J_5663674 | Gm43537 | ENSEMBL:ENSMUSG00000105022 | MGI:5663674 | unclassified gene | predicted gene 43537 |
| 3 | pseudogene | 36.53917 | 36.54010 | negative | MGI_C57BL6J_3650891 | Gm11550 | NCBI_Gene:665709,ENSEMBL:ENSMUSG00000083388 | MGI:3650891 | pseudogene | predicted gene 11550 |
| 3 | gene | 36.55261 | 36.56573 | positive | MGI_C57BL6J_1355319 | Exosc9 | NCBI_Gene:50911,ENSEMBL:ENSMUSG00000027714 | MGI:1355319 | protein coding gene | exosome component 9 |
| 3 | gene | 36.56486 | 36.57303 | negative | MGI_C57BL6J_108069 | Ccna2 | NCBI_Gene:12428,ENSEMBL:ENSMUSG00000027715 | MGI:108069 | protein coding gene | cyclin A2 |
| 3 | gene | 36.57314 | 36.61349 | negative | MGI_C57BL6J_1918742 | Bbs7 | NCBI_Gene:71492,ENSEMBL:ENSMUSG00000037325 | MGI:1918742 | protein coding gene | Bardet-Biedl syndrome 7 (human) |
| 3 | gene | 36.62048 | 36.69022 | negative | MGI_C57BL6J_109526 | Trpc3 | NCBI_Gene:22065,ENSEMBL:ENSMUSG00000027716 | MGI:109526 | protein coding gene | transient receptor potential cation channel, subfamily C, member 3 |
| 3 | gene | 36.65407 | 36.65501 | negative | MGI_C57BL6J_5595015 | Gm35856 | NCBI_Gene:102639576 | MGI:5595015 | lncRNA gene | predicted gene, 35856 |
| 3 | pseudogene | 36.69480 | 36.69505 | negative | MGI_C57BL6J_3651607 | Gm12377 | ENSEMBL:ENSMUSG00000082548 | MGI:3651607 | pseudogene | predicted gene 12377 |
| 3 | pseudogene | 36.82435 | 36.82506 | negative | MGI_C57BL6J_3651608 | Gm12376 | NCBI_Gene:100417988,ENSEMBL:ENSMUSG00000084027 | MGI:3651608 | pseudogene | predicted gene 12376 |
| 3 | gene | 36.86306 | 37.05303 | positive | MGI_C57BL6J_2444631 | 4932438A13Rik | NCBI_Gene:229227,ENSEMBL:ENSMUSG00000037270 | MGI:2444631 | protein coding gene | RIKEN cDNA 4932438A13 gene |
| 3 | gene | 37.05221 | 37.06003 | negative | MGI_C57BL6J_3650531 | Gm12531 | ENSEMBL:ENSMUSG00000086090 | MGI:3650531 | lncRNA gene | predicted gene 12531 |
| 3 | gene | 37.06309 | 37.12193 | positive | MGI_C57BL6J_103258 | Adad1 | NCBI_Gene:21744,ENSEMBL:ENSMUSG00000027719 | MGI:103258 | protein coding gene | adenosine deaminase domain containing 1 (testis specific) |
| 3 | pseudogene | 37.07756 | 37.07891 | positive | MGI_C57BL6J_3651489 | Gm12582 | NCBI_Gene:652987,ENSEMBL:ENSMUSG00000081705 | MGI:3651489 | pseudogene | predicted gene 12582 |
| 3 | pseudogene | 37.10808 | 37.10933 | positive | MGI_C57BL6J_3651485 | Gm12583 | NCBI_Gene:100416060,ENSEMBL:ENSMUSG00000082878 | MGI:3651485 | pseudogene | predicted gene 12583 |
| 3 | pseudogene | 37.11673 | 37.11784 | positive | MGI_C57BL6J_3651486 | Gm12584 | ENSEMBL:ENSMUSG00000083857 | MGI:3651486 | pseudogene | predicted gene 12584 |
| 3 | gene | 37.12052 | 37.12596 | negative | MGI_C57BL6J_96548 | Il2 | NCBI_Gene:16183,ENSEMBL:ENSMUSG00000027720 | MGI:96548 | protein coding gene | interleukin 2 |
| 3 | gene | 37.16972 | 37.17665 | negative | MGI_C57BL6J_5595159 | Gm36000 | NCBI_Gene:102639768 | MGI:5595159 | lncRNA gene | predicted gene, 36000 |
| 3 | gene | 37.18218 | 37.18860 | positive | MGI_C57BL6J_3650618 | Gm12532 | NCBI_Gene:102639848,ENSEMBL:ENSMUSG00000084754 | MGI:3650618 | lncRNA gene | predicted gene 12532 |
| 3 | gene | 37.22276 | 37.23264 | negative | MGI_C57BL6J_1890474 | Il21 | NCBI_Gene:60505,ENSEMBL:ENSMUSG00000027718 | MGI:1890474 | protein coding gene | interleukin 21 |
| 3 | gene | 37.22538 | 37.22711 | positive | MGI_C57BL6J_5595327 | Gm36168 | NCBI_Gene:102639985 | MGI:5595327 | lncRNA gene | predicted gene, 36168 |
| 3 | gene | 37.23133 | 37.23247 | positive | MGI_C57BL6J_3650303 | Gm12534 | ENSEMBL:ENSMUSG00000086517 | MGI:3650303 | lncRNA gene | predicted gene 12534 |
| 3 | gene | 37.23287 | 37.23334 | positive | MGI_C57BL6J_5663959 | Gm43822 | ENSEMBL:ENSMUSG00000104998 | MGI:5663959 | unclassified gene | predicted gene 43822 |
| 3 | gene | 37.25678 | 37.25689 | negative | MGI_C57BL6J_5453946 | Gm24169 | ENSEMBL:ENSMUSG00000094007 | MGI:5453946 | miRNA gene | predicted gene, 24169 |
| 3 | gene | 37.30775 | 37.31271 | negative | MGI_C57BL6J_2677454 | Cetn4 | NCBI_Gene:207175,ENSEMBL:ENSMUSG00000045031 | MGI:2677454 | protein coding gene | centrin 4 |
| 3 | gene | 37.31249 | 37.32145 | positive | MGI_C57BL6J_2686651 | Bbs12 | NCBI_Gene:241950,ENSEMBL:ENSMUSG00000051444 | MGI:2686651 | protein coding gene | Bardet-Biedl syndrome 12 (human) |
| 3 | gene | 37.31252 | 37.40487 | positive | MGI_C57BL6J_5663576 | Gm43439 | ENSEMBL:ENSMUSG00000106407 | MGI:5663576 | lncRNA gene | predicted gene 43439 |
| 3 | pseudogene | 37.31253 | 37.32919 | positive | MGI_C57BL6J_3651849 | Gm12540 | NCBI_Gene:102640067,ENSEMBL:ENSMUSG00000083353 | MGI:3651849 | pseudogene | predicted gene 12540 |
| 3 | gene | 37.33329 | 37.35103 | negative | MGI_C57BL6J_3649376 | Fgf2os | NCBI_Gene:329626,ENSEMBL:ENSMUSG00000054779 | MGI:3649376 | antisense lncRNA gene | fibroblast growth factor 2, opposite strand |
| 3 | gene | 37.33902 | 37.34152 | positive | MGI_C57BL6J_5663625 | Gm43488 | ENSEMBL:ENSMUSG00000105063 | MGI:5663625 | unclassified gene | predicted gene 43488 |
| 3 | gene | 37.34835 | 37.41011 | positive | MGI_C57BL6J_95516 | Fgf2 | NCBI_Gene:14173,ENSEMBL:ENSMUSG00000037225 | MGI:95516 | protein coding gene | fibroblast growth factor 2 |
| 3 | gene | 37.39221 | 37.39510 | negative | MGI_C57BL6J_5663060 | Gm42923 | ENSEMBL:ENSMUSG00000105134 | MGI:5663060 | unclassified gene | predicted gene 42923 |
| 3 | gene | 37.40491 | 37.42021 | negative | MGI_C57BL6J_2387618 | Nudt6 | NCBI_Gene:229228,ENSEMBL:ENSMUSG00000050174 | MGI:2387618 | protein coding gene | nudix (nucleoside diphosphate linked moiety X)-type motif 6 |
| 3 | gene | 37.41990 | 37.57910 | positive | MGI_C57BL6J_1927170 | Spata5 | NCBI_Gene:57815,ENSEMBL:ENSMUSG00000027722 | MGI:1927170 | protein coding gene | spermatogenesis associated 5 |
| 3 | gene | 37.48883 | 37.49479 | positive | MGI_C57BL6J_5663621 | Gm43484 | ENSEMBL:ENSMUSG00000106166 | MGI:5663621 | unclassified gene | predicted gene 43484 |
| 3 | gene | 37.54188 | 37.60239 | negative | MGI_C57BL6J_5595571 | Gm36412 | NCBI_Gene:102640314 | MGI:5595571 | lncRNA gene | predicted gene, 36412 |
| 3 | pseudogene | 37.55618 | 37.55650 | negative | MGI_C57BL6J_3652207 | Gm12564 | ENSEMBL:ENSMUSG00000080723 | MGI:3652207 | pseudogene | predicted gene 12564 |
| 3 | pseudogene | 37.59942 | 37.60007 | negative | MGI_C57BL6J_3652206 | Gm12565 | NCBI_Gene:665778,ENSEMBL:ENSMUSG00000081093 | MGI:3652206 | pseudogene | predicted gene 12565 |
| 3 | gene | 37.60989 | 37.61528 | positive | MGI_C57BL6J_1923197 | 4930594O21Rik | NCBI_Gene:75947,ENSEMBL:ENSMUSG00000085677 | MGI:1923197 | lncRNA gene | RIKEN cDNA 4930594O21 gene |
| 3 | gene | 37.63858 | 37.64066 | negative | MGI_C57BL6J_5663059 | Gm42922 | ENSEMBL:ENSMUSG00000104605 | MGI:5663059 | unclassified gene | predicted gene 42922 |
| 3 | gene | 37.63995 | 37.64460 | positive | MGI_C57BL6J_1345139 | Spry1 | NCBI_Gene:24063,ENSEMBL:ENSMUSG00000037211 | MGI:1345139 | protein coding gene | sprouty RTK signaling antagonist 1 |
| 3 | gene | 37.64461 | 37.64795 | positive | MGI_C57BL6J_5663957 | Gm43820 | ENSEMBL:ENSMUSG00000106062 | MGI:5663957 | unclassified gene | predicted gene 43820 |
| 3 | gene | 37.65268 | 37.65652 | negative | MGI_C57BL6J_5826436 | Gm46799 | NCBI_Gene:108168864 | MGI:5826436 | lncRNA gene | predicted gene, 46799 |
| 3 | gene | 37.69046 | 37.69486 | negative | MGI_C57BL6J_5625096 | Gm42211 | NCBI_Gene:105247029 | MGI:5625096 | lncRNA gene | predicted gene, 42211 |
| 3 | gene | 37.70802 | 37.71121 | positive | MGI_C57BL6J_5663551 | Gm43414 | ENSEMBL:ENSMUSG00000104935 | MGI:5663551 | unclassified gene | predicted gene 43414 |
| 3 | gene | 37.71419 | 37.72449 | negative | MGI_C57BL6J_3646006 | Gm5148 | NCBI_Gene:381438,ENSEMBL:ENSMUSG00000058174 | MGI:3646006 | protein coding gene | predicted gene 5148 |
| 3 | pseudogene | 37.71471 | 37.71522 | positive | MGI_C57BL6J_3705429 | Rps23-ps1 | NCBI_Gene:100038875,ENSEMBL:ENSMUSG00000100755 | MGI:3705429 | pseudogene | ribosomal protein S23, pseudogene 1 |
| 3 | gene | 37.72423 | 37.74132 | positive | MGI_C57BL6J_5595779 | Gm36620 | NCBI_Gene:102640592 | MGI:5595779 | lncRNA gene | predicted gene, 36620 |
| 3 | pseudogene | 37.73468 | 37.73514 | positive | MGI_C57BL6J_3643738 | Rpl31-ps15 | NCBI_Gene:626367,ENSEMBL:ENSMUSG00000097304 | MGI:3643738 | pseudogene | ribosomal protein L31, pseudogene 15 |
| 3 | gene | 37.75320 | 37.77440 | positive | MGI_C57BL6J_5625095 | Gm42210 | NCBI_Gene:105247028 | MGI:5625095 | lncRNA gene | predicted gene, 42210 |
| 3 | gene | 37.76104 | 37.76730 | positive | MGI_C57BL6J_5663058 | Gm42921 | ENSEMBL:ENSMUSG00000106441 | MGI:5663058 | lncRNA gene | predicted gene 42921 |
| 3 | gene | 37.80254 | 37.80265 | negative | MGI_C57BL6J_5456181 | Gm26404 | ENSEMBL:ENSMUSG00000089123 | MGI:5456181 | snoRNA gene | predicted gene, 26404 |
| 3 | gene | 37.82822 | 37.84829 | positive | MGI_C57BL6J_1925294 | 4930556A17Rik | NCBI_Gene:102640520 | MGI:1925294 | lncRNA gene | RIKEN cDNA 4930556A17 gene |
| 3 | gene | 37.83847 | 37.83957 | positive | MGI_C57BL6J_5662889 | Gm42752 | ENSEMBL:ENSMUSG00000106621 | MGI:5662889 | unclassified gene | predicted gene 42752 |
| 3 | gene | 37.84180 | 37.84307 | positive | MGI_C57BL6J_5662890 | Gm42753 | ENSEMBL:ENSMUSG00000105252 | MGI:5662890 | unclassified gene | predicted gene 42753 |
| 3 | gene | 37.89881 | 37.99273 | positive | MGI_C57BL6J_5434111 | Gm20755 | NCBI_Gene:626410,ENSEMBL:ENSMUSG00000106461 | MGI:5434111 | lncRNA gene | predicted gene, 20755 |
| 3 | pseudogene | 37.95961 | 37.96009 | negative | MGI_C57BL6J_3643378 | Gm7799 | ENSEMBL:ENSMUSG00000087457 | MGI:3643378 | pseudogene | predicted gene 7799 |
| 3 | gene | 37.98279 | 37.98335 | positive | MGI_C57BL6J_5662888 | Gm42751 | ENSEMBL:ENSMUSG00000105099 | MGI:5662888 | unclassified gene | predicted gene 42751 |
| 3 | gene | 37.99511 | 37.99524 | negative | MGI_C57BL6J_5452676 | Gm22899 | ENSEMBL:ENSMUSG00000077728 | MGI:5452676 | snoRNA gene | predicted gene, 22899 |
| 3 | gene | 38.09463 | 38.13019 | negative | MGI_C57BL6J_5663958 | Gm43821 | ENSEMBL:ENSMUSG00000105219 | MGI:5663958 | lncRNA gene | predicted gene 43821 |
| 3 | pseudogene | 38.10123 | 38.10798 | negative | MGI_C57BL6J_3643358 | Rpl21-ps10 | NCBI_Gene:665821,ENSEMBL:ENSMUSG00000105359 | MGI:3643358 | pseudogene | ribosomal protein L21, pseudogene 10 |
| 3 | gene | 38.15191 | 38.15915 | positive | MGI_C57BL6J_5595839 | Gm36680 | NCBI_Gene:102640666 | MGI:5595839 | lncRNA gene | predicted gene, 36680 |
| 3 | gene | 38.20315 | 38.20812 | negative | MGI_C57BL6J_1918592 | 5430434I15Rik | NCBI_Gene:71342,ENSEMBL:ENSMUSG00000106002 | MGI:1918592 | lncRNA gene | RIKEN cDNA 5430434I15 gene |
| 3 | pseudogene | 38.22345 | 38.22382 | positive | MGI_C57BL6J_3781143 | Gm2965 | NCBI_Gene:100040785,ENSEMBL:ENSMUSG00000105943 | MGI:3781143 | pseudogene | predicted gene 2965 |
| 3 | pseudogene | 38.25402 | 38.25480 | positive | MGI_C57BL6J_5010140 | Gm17955 | NCBI_Gene:100416169 | MGI:5010140 | pseudogene | predicted gene, 17955 |
| 3 | gene | 38.26784 | 38.26808 | negative | MGI_C57BL6J_5455351 | Gm25574 | ENSEMBL:ENSMUSG00000092986 | MGI:5455351 | unclassified non-coding RNA gene | predicted gene, 25574 |
| 3 | gene | 38.36615 | 38.36888 | negative | MGI_C57BL6J_5625098 | Gm42213 | NCBI_Gene:105247031 | MGI:5625098 | lncRNA gene | predicted gene, 42213 |
| 3 | gene | 38.36940 | 38.37364 | positive | MGI_C57BL6J_5663057 | Gm42920 | ENSEMBL:ENSMUSG00000106008 | MGI:5663057 | lncRNA gene | predicted gene 42920 |
| 3 | gene | 38.41897 | 38.42015 | negative | MGI_C57BL6J_1917958 | 3830422I06Rik | ENSEMBL:ENSMUSG00000106485 | MGI:1917958 | unclassified gene | RIKEN cDNA 3830422I06 gene |
| 3 | gene | 38.42655 | 38.44387 | negative | MGI_C57BL6J_5595982 | Gm36823 | NCBI_Gene:102640855,ENSEMBL:ENSMUSG00000105516 | MGI:5595982 | lncRNA gene | predicted gene, 36823 |
| 3 | pseudogene | 38.43895 | 38.43984 | negative | MGI_C57BL6J_3646285 | Gm7815 | NCBI_Gene:665838,ENSEMBL:ENSMUSG00000106201 | MGI:3646285 | pseudogene | predicted gene 7815 |
| 3 | gene | 38.44926 | 38.48486 | negative | MGI_C57BL6J_2139777 | Ankrd50 | NCBI_Gene:99696,ENSEMBL:ENSMUSG00000044864 | MGI:2139777 | protein coding gene | ankyrin repeat domain 50 |
| 3 | pseudogene | 38.55797 | 38.55886 | negative | MGI_C57BL6J_3779766 | Gm7824 | NCBI_Gene:665855,ENSEMBL:ENSMUSG00000105360 | MGI:3779766 | pseudogene | predicted gene 7824 |
| 3 | gene | 38.55811 | 38.55821 | negative | MGI_C57BL6J_5454239 | Gm24462 | ENSEMBL:ENSMUSG00000076385 | MGI:5454239 | miRNA gene | predicted gene, 24462 |
| 3 | gene | 38.61425 | 38.61556 | positive | MGI_C57BL6J_5663675 | Gm43538 | ENSEMBL:ENSMUSG00000106168 | MGI:5663675 | lncRNA gene | predicted gene 43538 |
| 3 | gene | 38.66361 | 38.70922 | positive | MGI_C57BL6J_5589038 | Gm29879 | NCBI_Gene:102631576 | MGI:5589038 | lncRNA gene | predicted gene, 29879 |
| 3 | gene | 38.76048 | 38.76195 | negative | MGI_C57BL6J_5589118 | Gm29959 | NCBI_Gene:102631683 | MGI:5589118 | lncRNA gene | predicted gene, 29959 |
| 3 | gene | 38.87457 | 38.88826 | negative | MGI_C57BL6J_2441892 | C230034O21Rik | NCBI_Gene:102632538,ENSEMBL:ENSMUSG00000086181 | MGI:2441892 | lncRNA gene | RIKEN cDNA C230034O21 gene |
| 3 | gene | 38.88563 | 39.01199 | positive | MGI_C57BL6J_3045256 | Fat4 | NCBI_Gene:329628,ENSEMBL:ENSMUSG00000046743 | MGI:3045256 | protein coding gene | FAT atypical cadherin 4 |
| 3 | gene | 39.00860 | 39.00968 | negative | MGI_C57BL6J_5663676 | Gm43539 | ENSEMBL:ENSMUSG00000105910 | MGI:5663676 | lncRNA gene | predicted gene 43539 |
| 3 | pseudogene | 39.12853 | 39.12891 | negative | MGI_C57BL6J_5663445 | Gm43308 | ENSEMBL:ENSMUSG00000104792 | MGI:5663445 | pseudogene | predicted gene 43308 |
| 3 | gene | 39.15083 | 39.15967 | negative | MGI_C57BL6J_5589170 | Gm30011 | NCBI_Gene:102631750 | MGI:5589170 | lncRNA gene | predicted gene, 30011 |
| 3 | pseudogene | 39.21756 | 39.21785 | negative | MGI_C57BL6J_5663145 | Gm43008 | ENSEMBL:ENSMUSG00000107333 | MGI:5663145 | pseudogene | predicted gene 43008 |
| 3 | gene | 39.27713 | 39.30593 | positive | MGI_C57BL6J_5625099 | Gm42214 | NCBI_Gene:105247032 | MGI:5625099 | lncRNA gene | predicted gene, 42214 |
| 3 | pseudogene | 39.35737 | 39.35913 | negative | MGI_C57BL6J_3704215 | Gm9845 | NCBI_Gene:100040853,ENSEMBL:ENSMUSG00000050533 | MGI:3704215 | pseudogene | predicted pseudogene 9845 |
| 3 | pseudogene | 39.44389 | 39.44435 | positive | MGI_C57BL6J_5295673 | Gm20566 | NCBI_Gene:100047050 | MGI:5295673 | pseudogene | predicted gene, 20566 |
| 3 | gene | 39.46078 | 39.59865 | positive | MGI_C57BL6J_5589222 | Gm30063 | NCBI_Gene:102631823 | MGI:5589222 | lncRNA gene | predicted gene, 30063 |
| 3 | gene | 39.52552 | 39.53415 | negative | MGI_C57BL6J_5625100 | Gm42215 | NCBI_Gene:105247033 | MGI:5625100 | lncRNA gene | predicted gene, 42215 |
| 3 | gene | 39.63596 | 39.63608 | negative | MGI_C57BL6J_4422060 | n-R5s195 | ENSEMBL:ENSMUSG00000093952 | MGI:4422060 | rRNA gene | nuclear encoded rRNA 5S 195 |
| 3 | gene | 39.63606 | 39.63878 | positive | MGI_C57BL6J_5662918 | Gm42781 | ENSEMBL:ENSMUSG00000106786 | MGI:5662918 | unclassified gene | predicted gene 42781 |
| 3 | gene | 39.66629 | 39.66645 | positive | MGI_C57BL6J_5453573 | Gm23796 | ENSEMBL:ENSMUSG00000065945 | MGI:5453573 | snRNA gene | predicted gene, 23796 |
| 3 | gene | 39.71315 | 39.72565 | positive | MGI_C57BL6J_5826471 | Gm46834 | NCBI_Gene:108168914 | MGI:5826471 | lncRNA gene | predicted gene, 46834 |
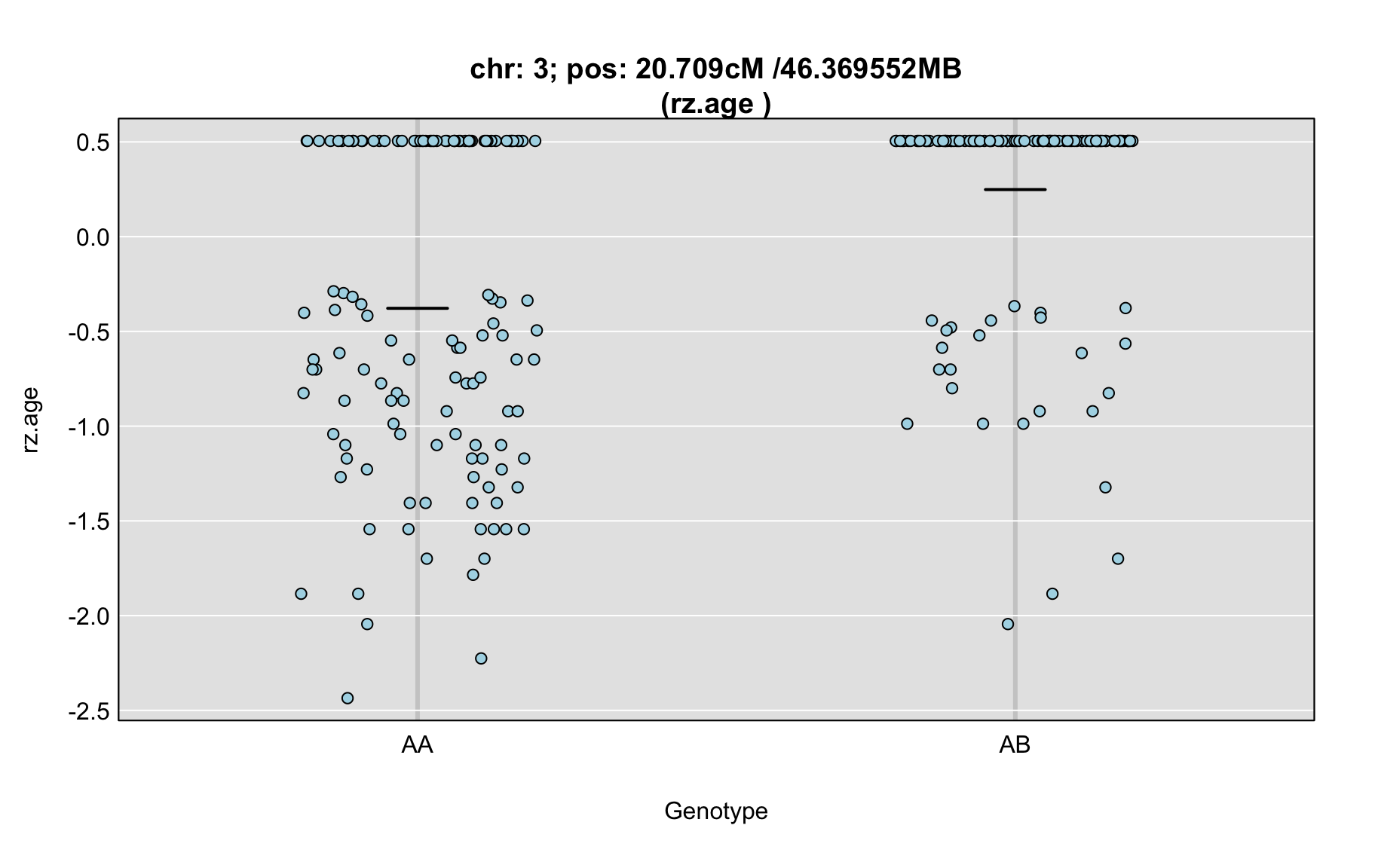
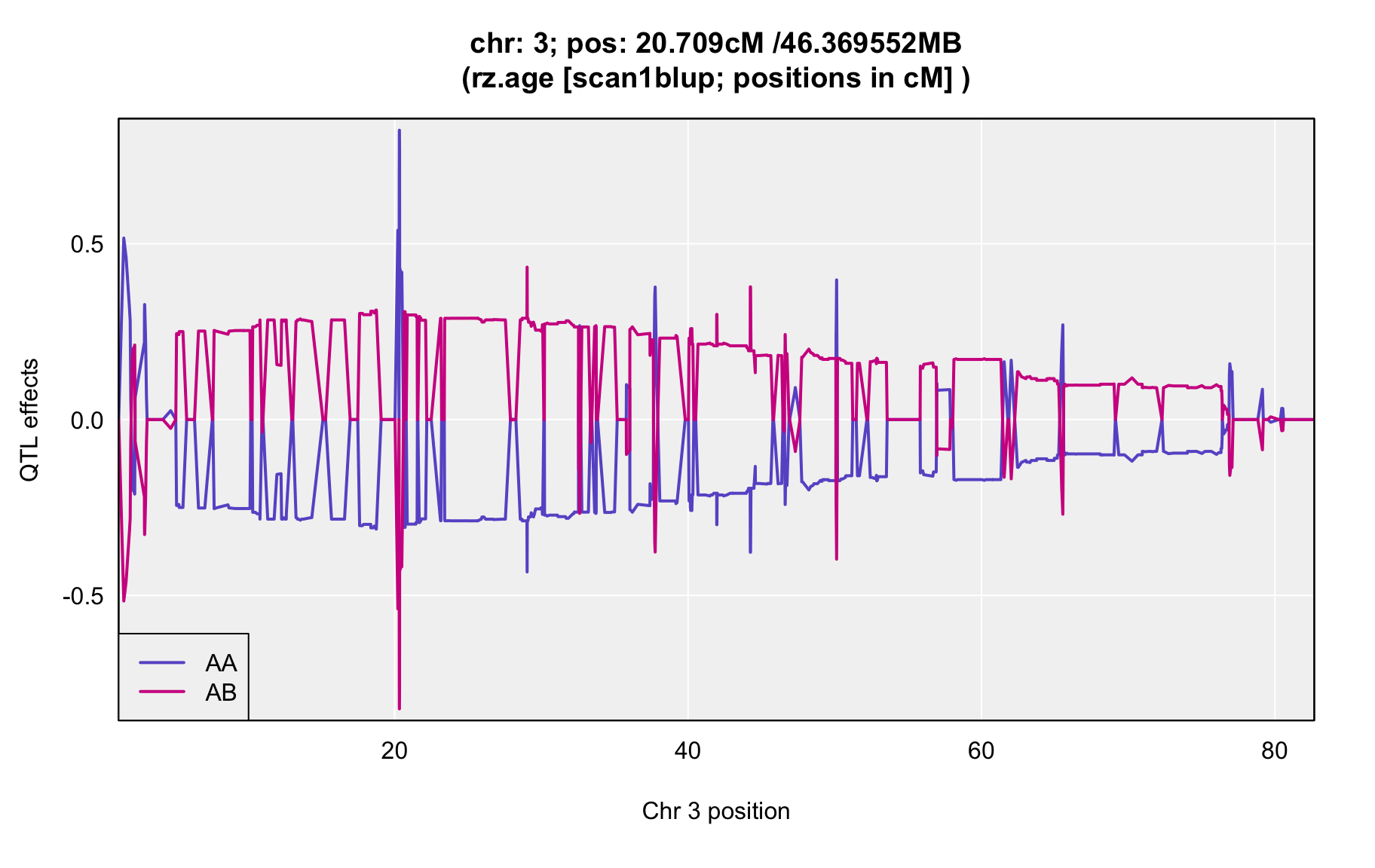
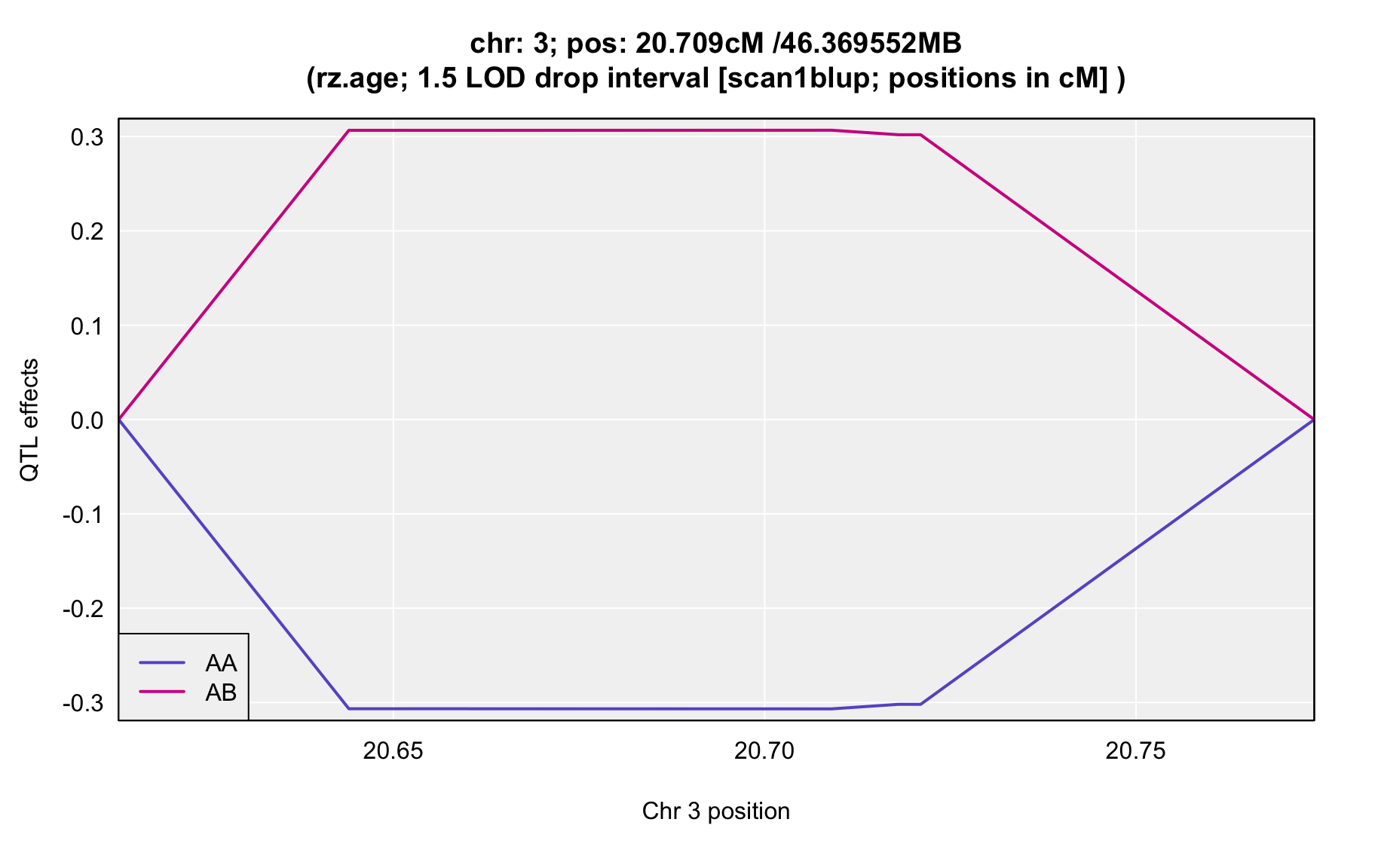
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 46.28983 | 46.28996 | positive | MGI_C57BL6J_5455858 | Gm26081 | ENSEMBL:ENSMUSG00000077256 | MGI:5455858 | snoRNA gene | predicted gene, 26081 |
| 3 | pseudogene | 46.41340 | 46.41412 | positive | MGI_C57BL6J_5011028 | Gm18843 | NCBI_Gene:100417811,ENSEMBL:ENSMUSG00000104511 | MGI:5011028 | pseudogene | predicted gene, 18843 |
| 3 | gene | 46.44085 | 46.44822 | negative | MGI_C57BL6J_3643087 | Pabpc4l | NCBI_Gene:241989,ENSEMBL:ENSMUSG00000090919 | MGI:3643087 | protein coding gene | poly(A) binding protein, cytoplasmic 4-like |
| 3 | pseudogene | 46.62112 | 46.62136 | positive | MGI_C57BL6J_5610673 | Gm37445 | ENSEMBL:ENSMUSG00000102589 | MGI:5610673 | pseudogene | predicted gene, 37445 |
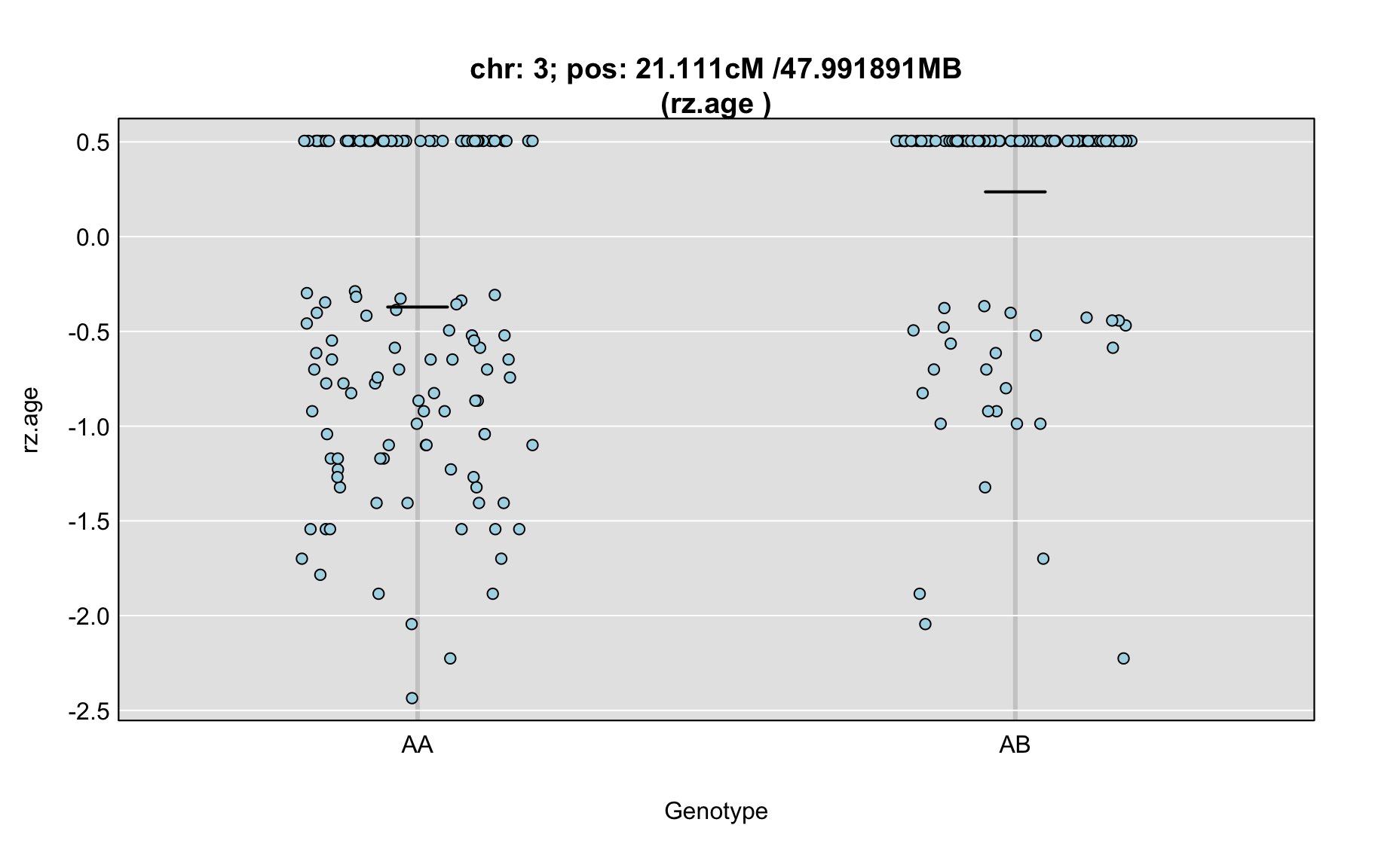
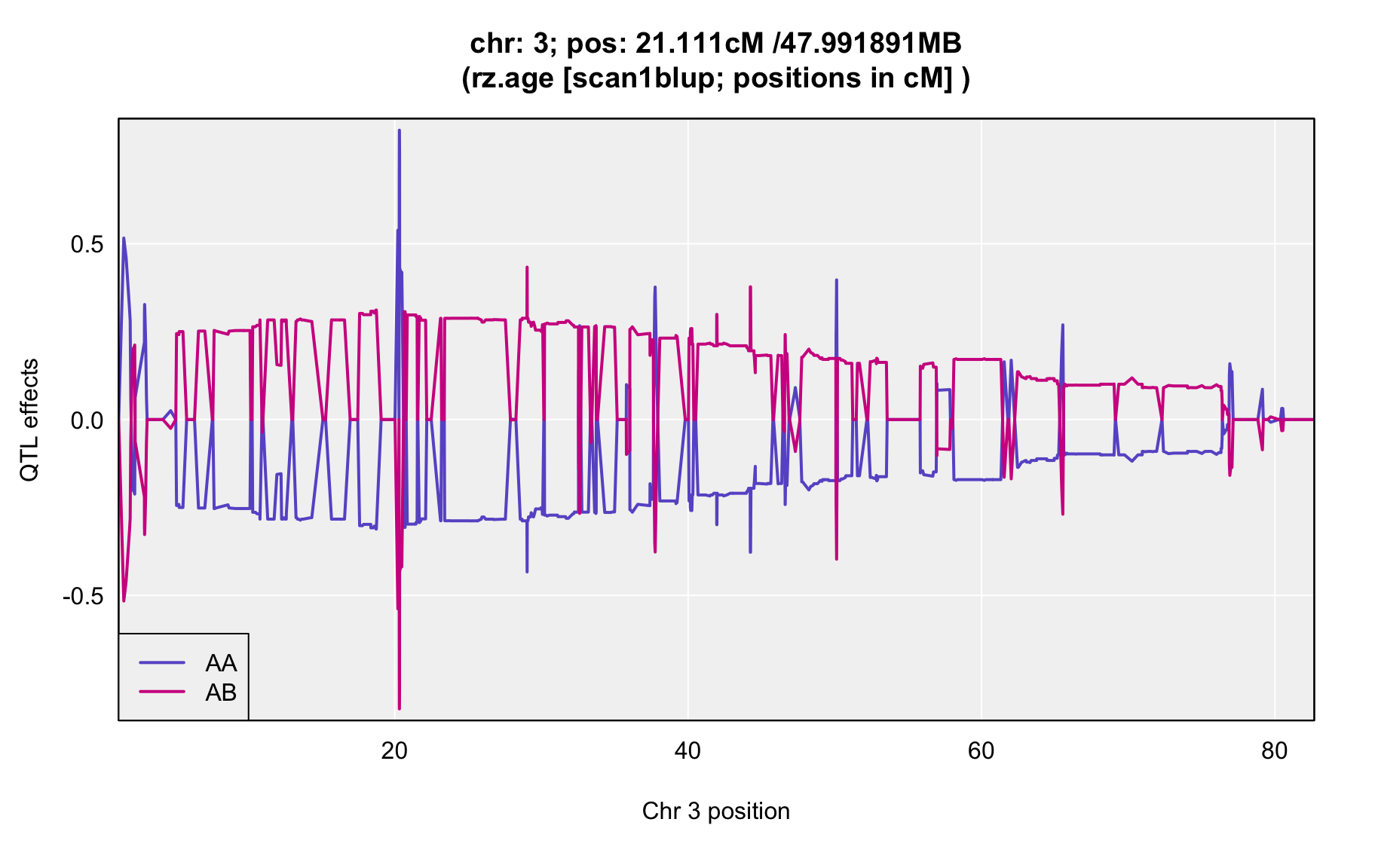
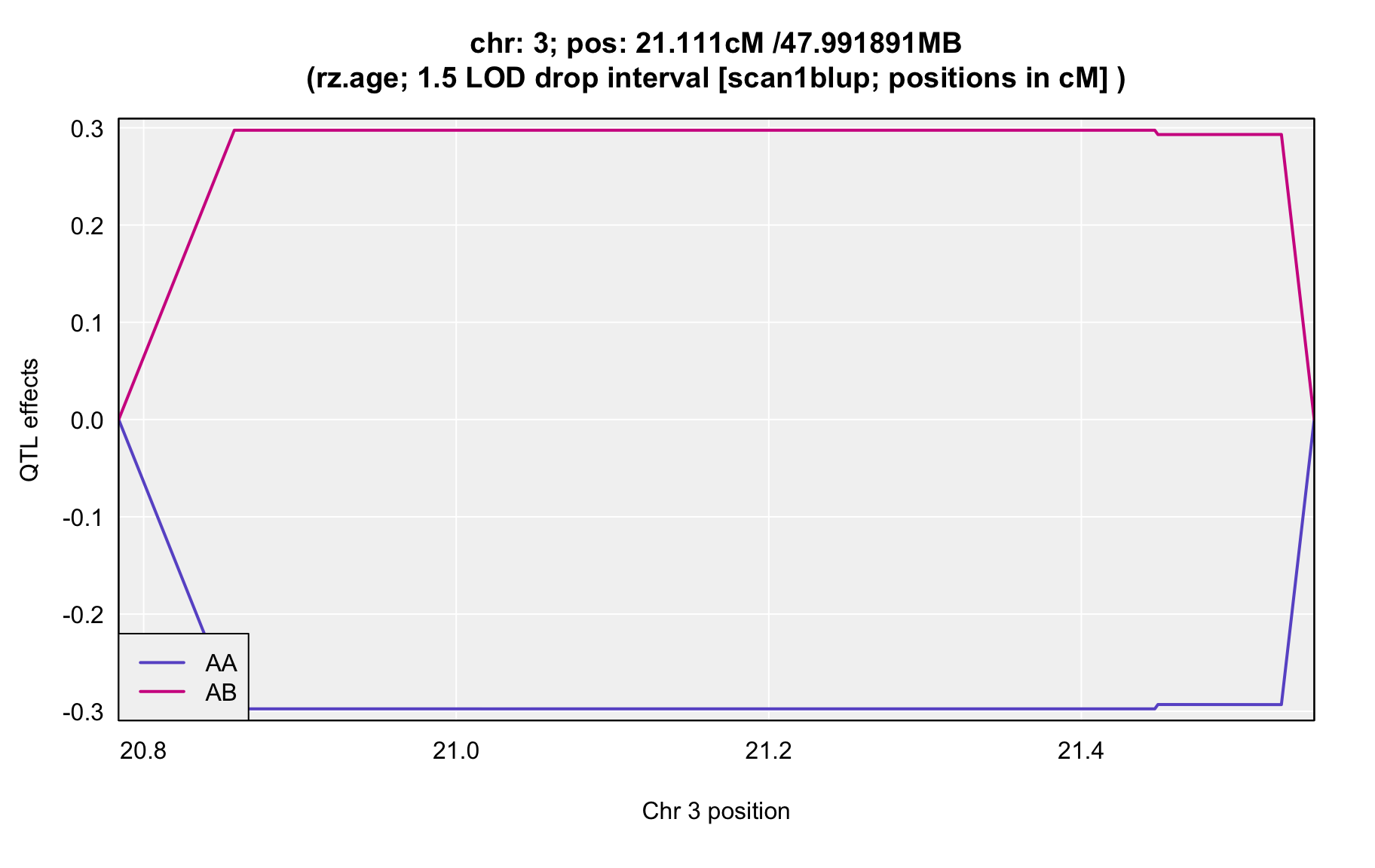
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | pseudogene | 47.45961 | 47.46034 | negative | MGI_C57BL6J_3780399 | Gm2229 | NCBI_Gene:100039426,ENSEMBL:ENSMUSG00000103798 | MGI:3780399 | pseudogene | predicted gene 2229 |
| 3 | pseudogene | 48.02601 | 48.02676 | negative | MGI_C57BL6J_3648898 | Gm7977 | NCBI_Gene:666202,ENSEMBL:ENSMUSG00000103257 | MGI:3648898 | pseudogene | predicted gene 7977 |
| 3 | gene | 48.07834 | 48.07845 | positive | MGI_C57BL6J_5610523 | Gm37295 | ENSEMBL:ENSMUSG00000104110 | MGI:5610523 | unclassified gene | predicted gene, 37295 |
| 3 | gene | 48.54642 | 48.57947 | positive | MGI_C57BL6J_5826472 | Gm46835 | NCBI_Gene:108168915 | MGI:5826472 | lncRNA gene | predicted gene, 46835 |
| 3 | gene | 48.60573 | 48.60910 | negative | MGI_C57BL6J_1913582 | 1700018B24Rik | NCBI_Gene:66332,ENSEMBL:ENSMUSG00000037910 | MGI:1913582 | lncRNA gene | RIKEN cDNA 1700018B24 gene |
| 3 | gene | 48.74851 | 48.76095 | negative | MGI_C57BL6J_5610418 | Gm37190 | ENSEMBL:ENSMUSG00000102181 | MGI:5610418 | lncRNA gene | predicted gene, 37190 |
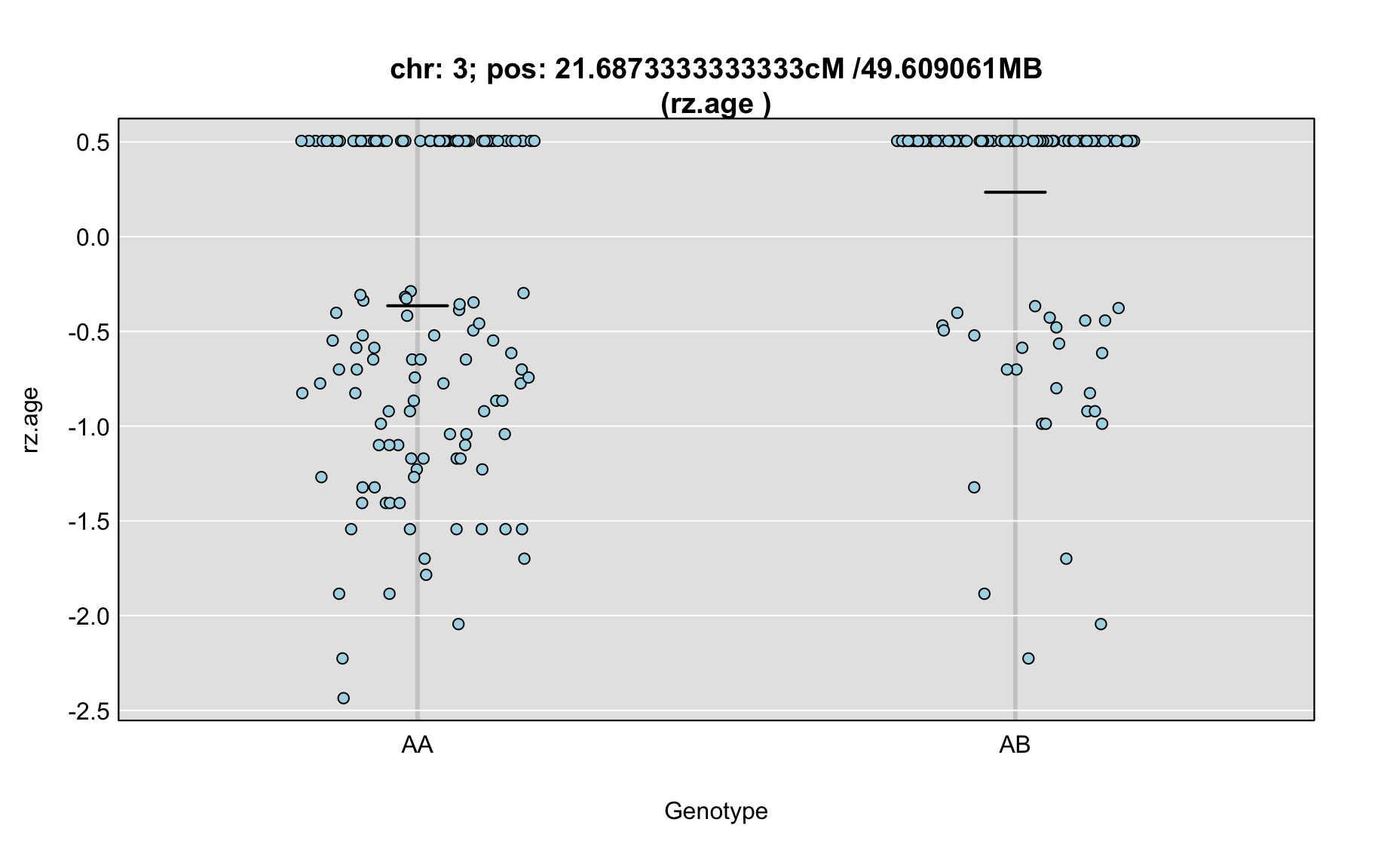
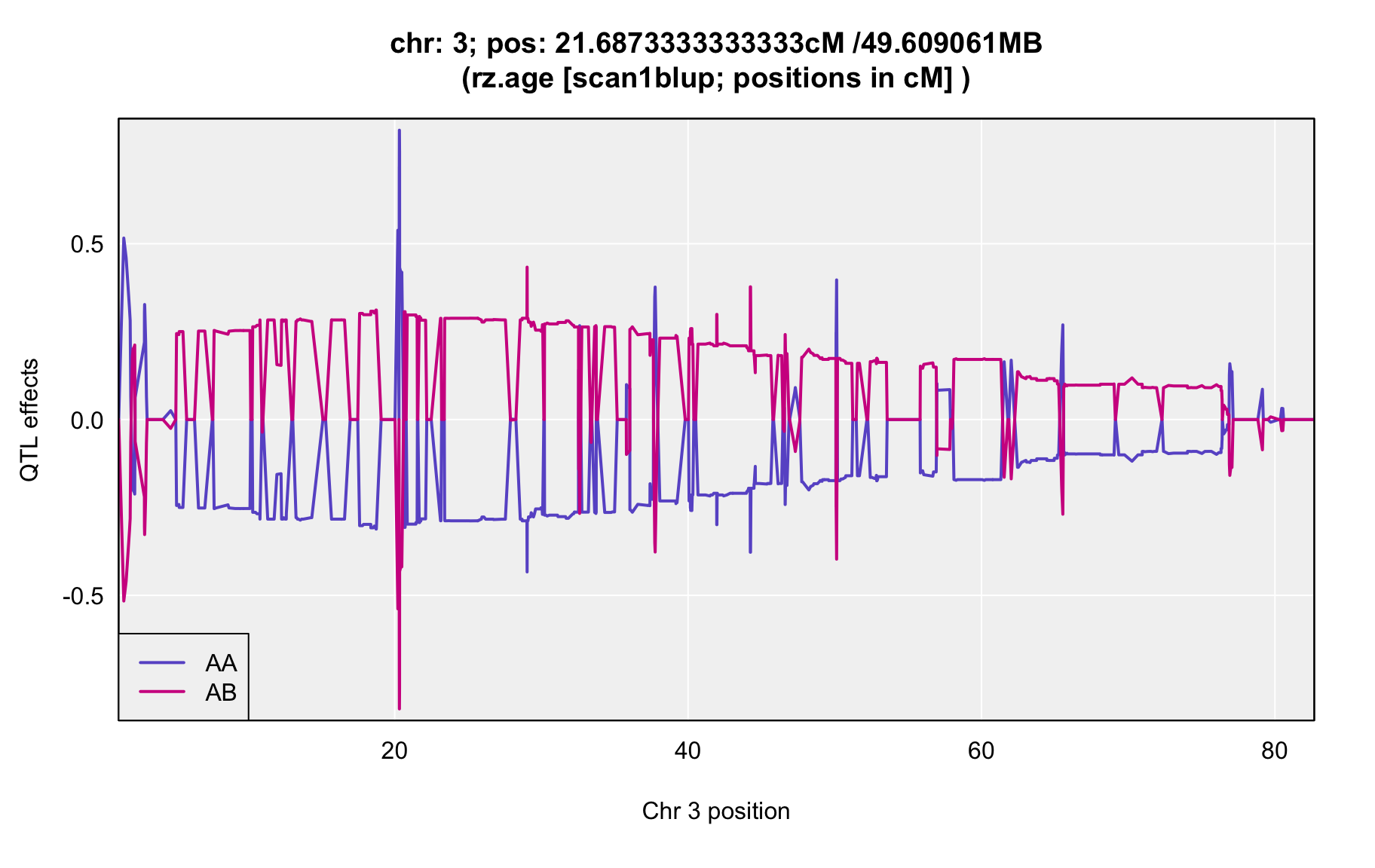
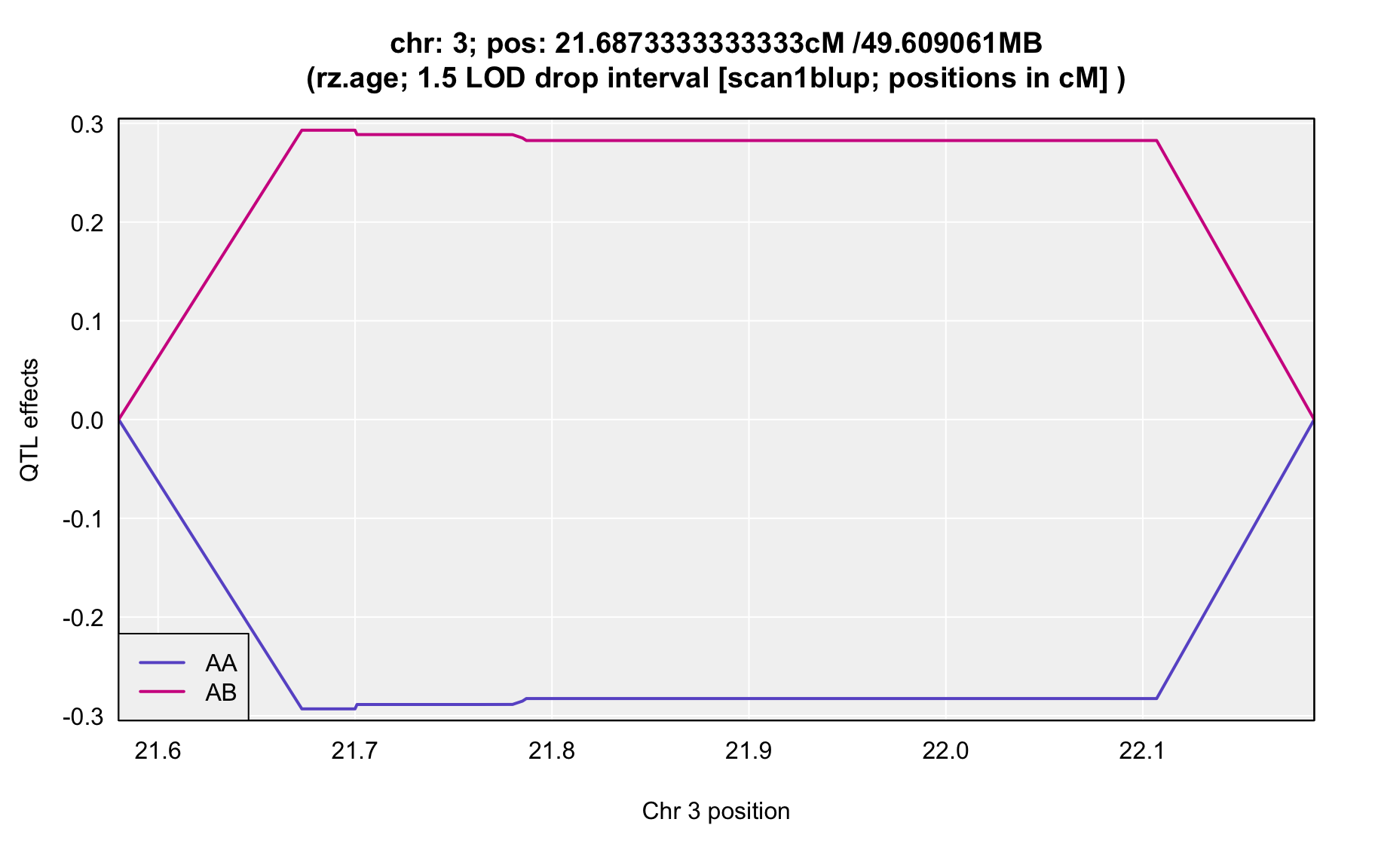
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 48.95472 | 48.95488 | positive | MGI_C57BL6J_5453288 | Gm23511 | ENSEMBL:ENSMUSG00000065018 | MGI:5453288 | snRNA gene | predicted gene, 23511 |
| 3 | gene | 49.06237 | 49.09636 | positive | MGI_C57BL6J_5590574 | Gm31415 | NCBI_Gene:102633632,ENSEMBL:ENSMUSG00000102263 | MGI:5590574 | lncRNA gene | predicted gene, 31415 |
| 3 | gene | 49.13746 | 49.13853 | positive | MGI_C57BL6J_5611047 | Gm37819 | ENSEMBL:ENSMUSG00000104186 | MGI:5611047 | unclassified gene | predicted gene, 37819 |
| 3 | pseudogene | 49.20771 | 49.20807 | negative | MGI_C57BL6J_5610912 | Gm37684 | ENSEMBL:ENSMUSG00000103027 | MGI:5610912 | pseudogene | predicted gene, 37684 |
| 3 | gene | 49.32568 | 49.36736 | positive | MGI_C57BL6J_5295664 | Gm20557 | NCBI_Gene:329633,ENSEMBL:ENSMUSG00000102679 | MGI:5295664 | lncRNA gene | predicted gene, 20557 |
| 3 | pseudogene | 49.38115 | 49.38285 | negative | MGI_C57BL6J_3643624 | Gm5846 | NCBI_Gene:545519,ENSEMBL:ENSMUSG00000104309 | MGI:3643624 | pseudogene | predicted gene 5846 |
| 3 | pseudogene | 49.40082 | 49.46744 | negative | MGI_C57BL6J_3780091 | Gm9683 | NCBI_Gene:676568 | MGI:3780091 | pseudogene | predicted gene 9683 |
| 3 | gene | 49.49135 | 49.49148 | positive | MGI_C57BL6J_5452257 | Gm22480 | ENSEMBL:ENSMUSG00000084517 | MGI:5452257 | snoRNA gene | predicted gene, 22480 |
| 3 | pseudogene | 49.53075 | 49.53131 | positive | MGI_C57BL6J_5610652 | Gm37424 | ENSEMBL:ENSMUSG00000104437 | MGI:5610652 | pseudogene | predicted gene, 37424 |
| 3 | pseudogene | 49.60356 | 49.60388 | positive | MGI_C57BL6J_5611489 | Gm38261 | ENSEMBL:ENSMUSG00000102731 | MGI:5611489 | pseudogene | predicted gene, 38261 |
| 3 | pseudogene | 49.60423 | 49.60447 | positive | MGI_C57BL6J_5611316 | Gm38088 | ENSEMBL:ENSMUSG00000103249 | MGI:5611316 | pseudogene | predicted gene, 38088 |
| 3 | pseudogene | 49.63244 | 49.63297 | negative | MGI_C57BL6J_3780498 | Gm2328 | NCBI_Gene:100039587,ENSEMBL:ENSMUSG00000104134 | MGI:3780498 | pseudogene | predicted gene 2328 |
| 3 | pseudogene | 49.64348 | 49.64413 | positive | MGI_C57BL6J_5588900 | Gm29741 | NCBI_Gene:101055903,ENSEMBL:ENSMUSG00000103568 | MGI:5588900 | pseudogene | predicted gene, 29741 |
| 3 | gene | 49.65876 | 49.65887 | positive | MGI_C57BL6J_5452873 | Gm23096 | ENSEMBL:ENSMUSG00000065381 | MGI:5452873 | snRNA gene | predicted gene, 23096 |
| 3 | gene | 49.74329 | 49.75738 | negative | MGI_C57BL6J_1920423 | Pcdh18 | NCBI_Gene:73173,ENSEMBL:ENSMUSG00000037892 | MGI:1920423 | protein coding gene | protocadherin 18 |
| 3 | pseudogene | 49.78043 | 49.78089 | negative | MGI_C57BL6J_5610778 | Gm37550 | ENSEMBL:ENSMUSG00000102943 | MGI:5610778 | pseudogene | predicted gene, 37550 |
| 3 | gene | 49.89253 | 50.49909 | negative | MGI_C57BL6J_1347355 | Slc7a11 | NCBI_Gene:26570,ENSEMBL:ENSMUSG00000027737 | MGI:1347355 | protein coding gene | solute carrier family 7 (cationic amino acid transporter, y+ system), member 11 |
| 3 | gene | 50.00197 | 50.01078 | positive | MGI_C57BL6J_5611082 | Gm37854 | ENSEMBL:ENSMUSG00000102892 | MGI:5611082 | lncRNA gene | predicted gene, 37854 |
| 3 | gene | 50.01177 | 50.02624 | negative | MGI_C57BL6J_5622927 | Gm40042 | NCBI_Gene:105244434 | MGI:5622927 | lncRNA gene | predicted gene, 40042 |
| 3 | pseudogene | 50.02967 | 50.02999 | positive | MGI_C57BL6J_5611054 | Gm37826 | ENSEMBL:ENSMUSG00000104457 | MGI:5611054 | pseudogene | predicted gene, 37826 |
| 3 | pseudogene | 50.16666 | 50.16689 | negative | MGI_C57BL6J_5610920 | Gm37692 | ENSEMBL:ENSMUSG00000104349 | MGI:5610920 | pseudogene | predicted gene, 37692 |
| 3 | pseudogene | 50.25485 | 50.25544 | positive | MGI_C57BL6J_3780515 | Gm2345 | NCBI_Gene:100039628,ENSEMBL:ENSMUSG00000102167 | MGI:3780515 | pseudogene | predicted gene 2345 |
| 3 | gene | 50.40028 | 50.40428 | negative | MGI_C57BL6J_5610726 | Gm37498 | ENSEMBL:ENSMUSG00000103907 | MGI:5610726 | unclassified gene | predicted gene, 37498 |
| 3 | gene | 50.51640 | 50.60731 | negative | MGI_C57BL6J_3643374 | Gm6209 | NCBI_Gene:621304,ENSEMBL:ENSMUSG00000102715 | MGI:3643374 | lncRNA gene | predicted gene 6209 |
| 3 | gene | 50.57519 | 50.57652 | negative | MGI_C57BL6J_1923259 | 5830415G21Rik | ENSEMBL:ENSMUSG00000102895 | MGI:1923259 | unclassified gene | RIKEN cDNA 5830415G21 gene |
| 3 | pseudogene | 50.59799 | 50.59850 | negative | MGI_C57BL6J_5611465 | Gm38237 | NCBI_Gene:108168851,ENSEMBL:ENSMUSG00000103745 | MGI:5611465 | pseudogene | predicted gene, 38237 |
| 3 | gene | 50.62435 | 50.62841 | negative | MGI_C57BL6J_5610427 | Gm37199 | ENSEMBL:ENSMUSG00000104068 | MGI:5610427 | unclassified gene | predicted gene, 37199 |
| 3 | gene | 50.67970 | 50.68340 | negative | MGI_C57BL6J_5610689 | Gm37461 | ENSEMBL:ENSMUSG00000104281 | MGI:5610689 | unclassified gene | predicted gene, 37461 |
| 3 | gene | 50.70690 | 50.71189 | negative | MGI_C57BL6J_5622928 | Gm40043 | NCBI_Gene:105244435 | MGI:5622928 | lncRNA gene | predicted gene, 40043 |
| 3 | gene | 50.83858 | 50.83872 | negative | MGI_C57BL6J_5454280 | Gm24503 | ENSEMBL:ENSMUSG00000089079 | MGI:5454280 | snoRNA gene | predicted gene, 24503 |
| 3 | gene | 50.88324 | 50.88889 | negative | MGI_C57BL6J_5610776 | Gm37548 | NCBI_Gene:105244436,ENSEMBL:ENSMUSG00000103012 | MGI:5610776 | lncRNA gene | predicted gene, 37548 |
| 3 | gene | 50.91889 | 50.92153 | negative | MGI_C57BL6J_5611071 | Gm37843 | ENSEMBL:ENSMUSG00000102614 | MGI:5611071 | unclassified gene | predicted gene, 37843 |
| 3 | gene | 50.96656 | 50.96677 | positive | MGI_C57BL6J_2139603 | AA672641 | NA | NA | unclassified gene | expressed sequence AA672641 |
| 3 | gene | 50.97754 | 50.97881 | negative | MGI_C57BL6J_5610437 | Gm37209 | ENSEMBL:ENSMUSG00000103652 | MGI:5610437 | unclassified gene | predicted gene, 37209 |
| 3 | gene | 51.03335 | 51.04284 | negative | MGI_C57BL6J_5594915 | Gm35756 | NCBI_Gene:102639442 | MGI:5594915 | lncRNA gene | predicted gene, 35756 |
| 3 | gene | 51.05891 | 51.08934 | negative | MGI_C57BL6J_5579936 | Gm29230 | NCBI_Gene:102639521,ENSEMBL:ENSMUSG00000102067 | MGI:5579936 | lncRNA gene | predicted gene 29230 |
| 3 | gene | 51.10366 | 51.10517 | positive | MGI_C57BL6J_5611474 | Gm38246 | ENSEMBL:ENSMUSG00000103464 | MGI:5611474 | lncRNA gene | predicted gene, 38246 |
| 3 | gene | 51.12669 | 51.12894 | negative | MGI_C57BL6J_5595050 | Gm35891 | NCBI_Gene:102639626 | MGI:5595050 | lncRNA gene | predicted gene, 35891 |
| 3 | gene | 51.21847 | 51.22033 | negative | MGI_C57BL6J_5622929 | Gm40044 | NCBI_Gene:105244437 | MGI:5622929 | lncRNA gene | predicted gene, 40044 |
| 3 | gene | 51.22445 | 51.25165 | positive | MGI_C57BL6J_109382 | Noct | NCBI_Gene:12457,ENSEMBL:ENSMUSG00000023087 | MGI:109382 | protein coding gene | nocturnin |
| 3 | gene | 51.23192 | 51.23438 | positive | MGI_C57BL6J_5611585 | Gm38357 | ENSEMBL:ENSMUSG00000103313 | MGI:5611585 | unclassified gene | predicted gene, 38357 |
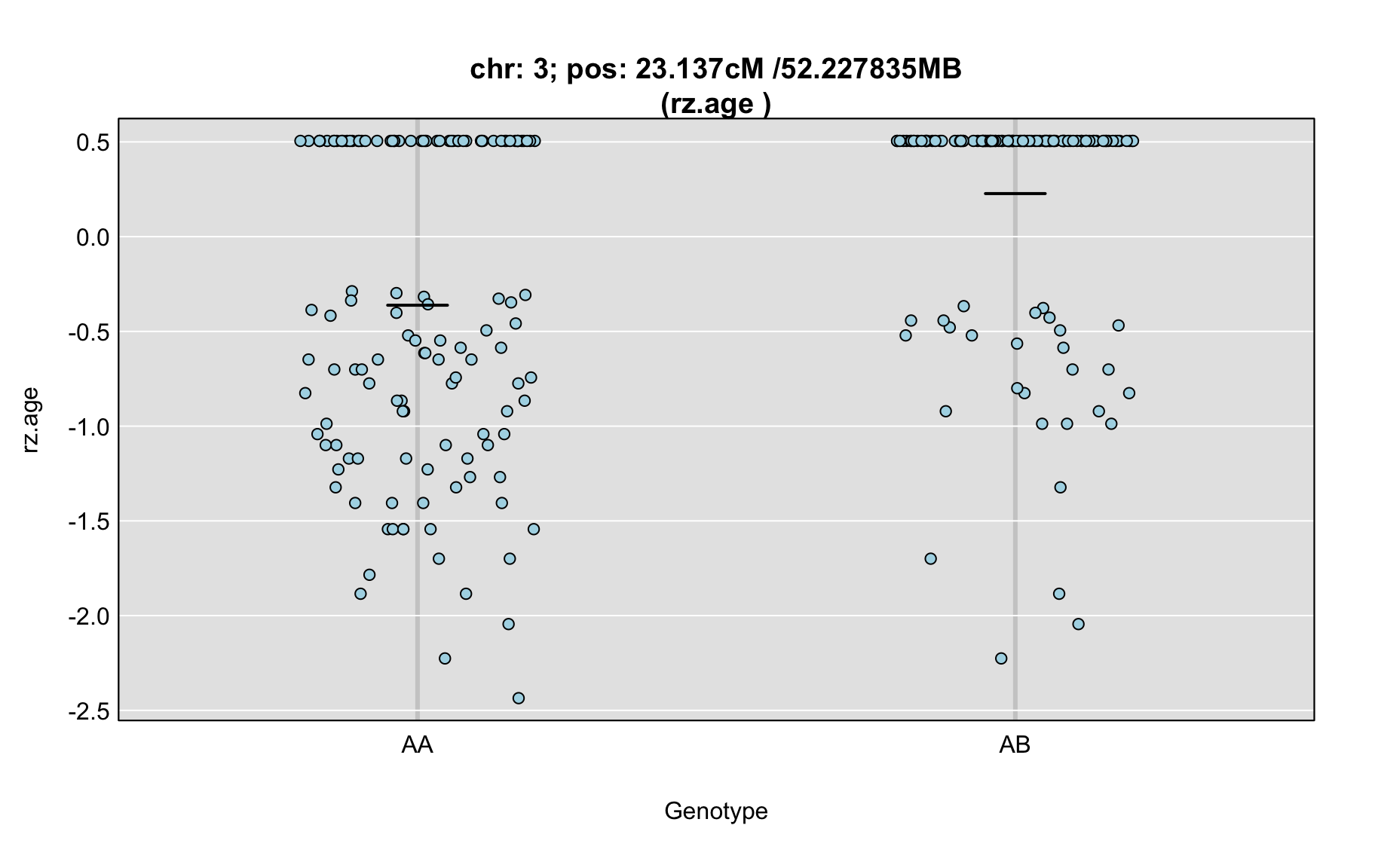
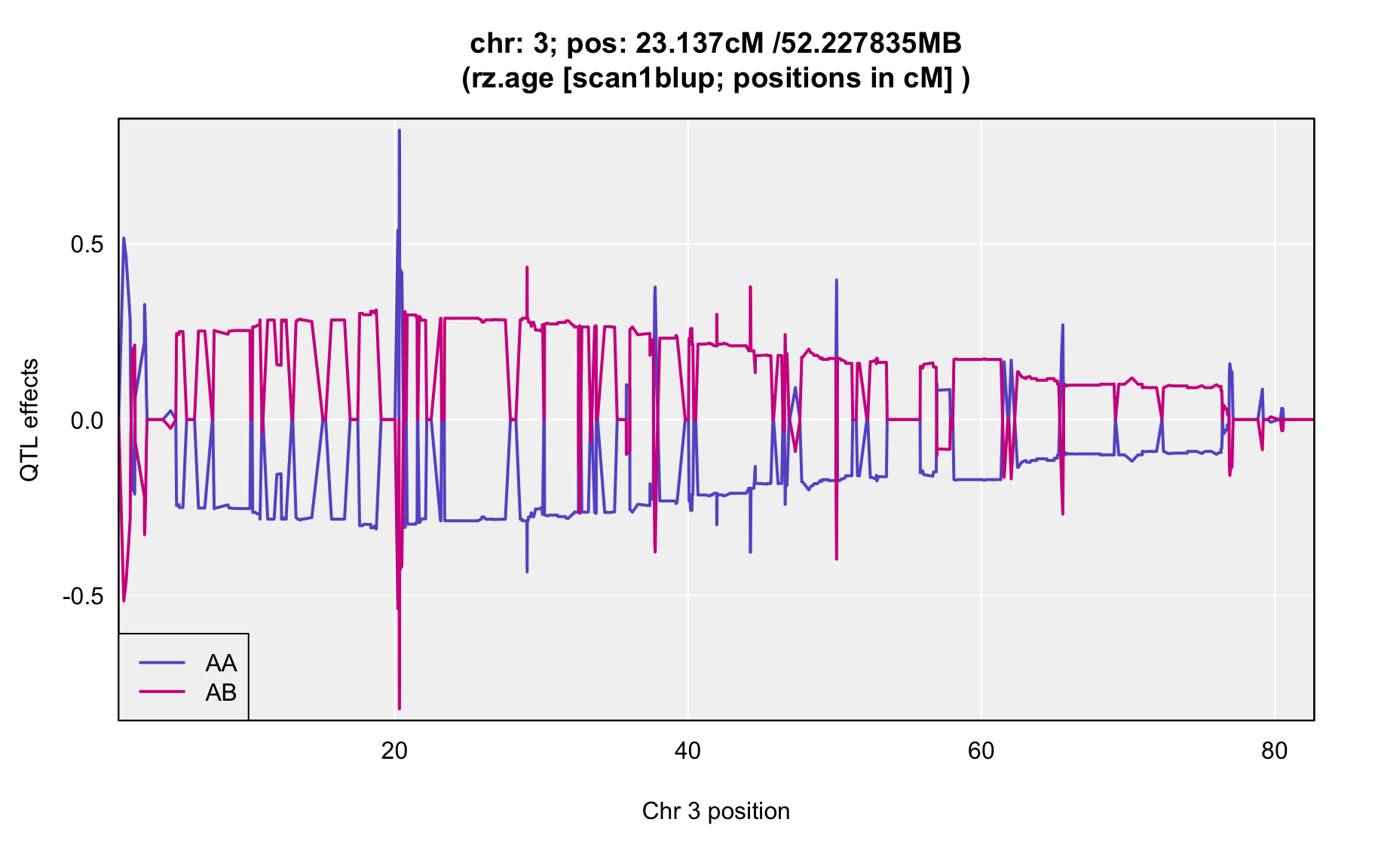
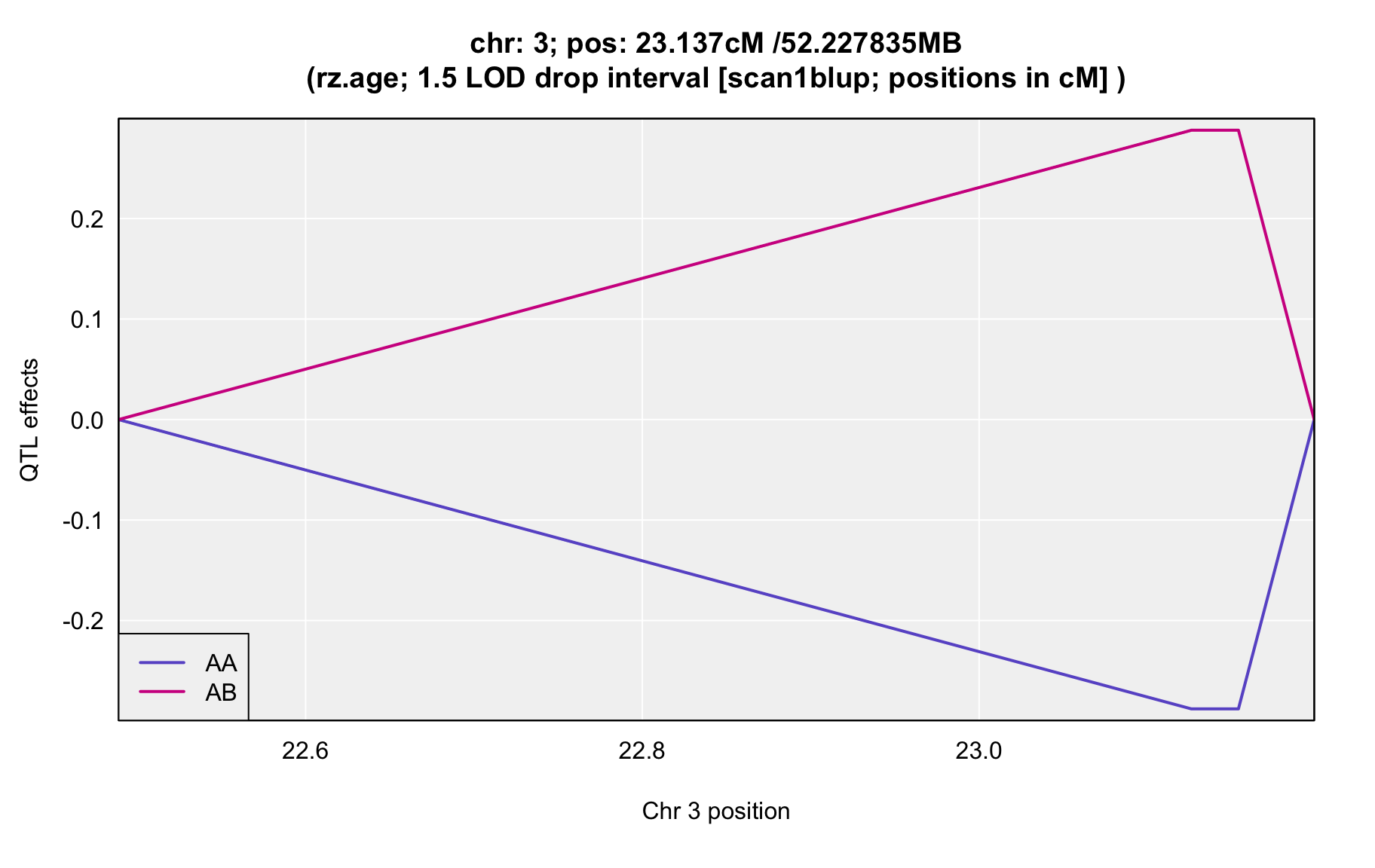
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 51.51099 | 51.51430 | positive | MGI_C57BL6J_5595167 | Gm36008 | NCBI_Gene:102639777 | MGI:5595167 | lncRNA gene | predicted gene, 36008 |
| 3 | gene | 51.51532 | 51.56088 | negative | MGI_C57BL6J_1920501 | Setd7 | NCBI_Gene:73251,ENSEMBL:ENSMUSG00000037111 | MGI:1920501 | protein coding gene | SET domain containing (lysine methyltransferase) 7 |
| 3 | gene | 51.55754 | 51.55965 | negative | MGI_C57BL6J_5611428 | Gm38200 | ENSEMBL:ENSMUSG00000102466 | MGI:5611428 | unclassified gene | predicted gene, 38200 |
| 3 | gene | 51.55976 | 51.56712 | positive | MGI_C57BL6J_1923230 | 5031434O11Rik | NCBI_Gene:100039684,ENSEMBL:ENSMUSG00000097885 | MGI:1923230 | lncRNA gene | RIKEN cDNA 5031434O11 gene |
| 3 | pseudogene | 51.57111 | 51.57142 | negative | MGI_C57BL6J_5611562 | Gm38334 | ENSEMBL:ENSMUSG00000104001 | MGI:5611562 | pseudogene | predicted gene, 38334 |
| 3 | gene | 51.58231 | 51.58285 | positive | MGI_C57BL6J_1918620 | 5430433H01Rik | ENSEMBL:ENSMUSG00000104075 | MGI:1918620 | unclassified gene | RIKEN cDNA 5430433H01 gene |
| 3 | pseudogene | 51.63649 | 51.63705 | negative | MGI_C57BL6J_5611475 | Gm38247 | NCBI_Gene:108168852,ENSEMBL:ENSMUSG00000104166 | MGI:5611475 | pseudogene | predicted gene, 38247 |
| 3 | gene | 51.66036 | 51.68268 | positive | MGI_C57BL6J_2448481 | Mgst2 | NCBI_Gene:211666,ENSEMBL:ENSMUSG00000074604 | MGI:2448481 | protein coding gene | microsomal glutathione S-transferase 2 |
| 3 | gene | 51.67280 | 51.67390 | positive | MGI_C57BL6J_5610489 | Gm37261 | ENSEMBL:ENSMUSG00000103527 | MGI:5610489 | unclassified gene | predicted gene, 37261 |
| 3 | gene | 51.68591 | 52.10509 | negative | MGI_C57BL6J_2389461 | Maml3 | NCBI_Gene:433586,ENSEMBL:ENSMUSG00000061143 | MGI:2389461 | protein coding gene | mastermind like transcriptional coactivator 3 |
| 3 | gene | 51.69372 | 51.69509 | positive | MGI_C57BL6J_3642905 | Gm10729 | ENSEMBL:ENSMUSG00000074603 | MGI:3642905 | protein coding gene | predicted gene 10729 |
| 3 | gene | 51.70772 | 51.71225 | positive | MGI_C57BL6J_5826440 | Gm46803 | NCBI_Gene:108168869 | MGI:5826440 | lncRNA gene | predicted gene, 46803 |
| 3 | gene | 51.75712 | 51.76209 | negative | MGI_C57BL6J_5610570 | Gm37342 | ENSEMBL:ENSMUSG00000102732 | MGI:5610570 | unclassified gene | predicted gene, 37342 |
| 3 | gene | 51.81295 | 51.81931 | positive | MGI_C57BL6J_5610457 | Gm37229 | ENSEMBL:ENSMUSG00000102539 | MGI:5610457 | lncRNA gene | predicted gene, 37229 |
| 3 | gene | 51.82891 | 51.82903 | positive | MGI_C57BL6J_4422061 | n-R5s196 | ENSEMBL:ENSMUSG00000065070 | MGI:4422061 | rRNA gene | nuclear encoded rRNA 5S 196 |
| 3 | gene | 51.82939 | 51.83040 | negative | MGI_C57BL6J_1920491 | 3110080O07Rik | ENSEMBL:ENSMUSG00000103284 | MGI:1920491 | unclassified gene | RIKEN cDNA 3110080O07 gene |
| 3 | gene | 51.86735 | 51.86856 | negative | MGI_C57BL6J_5611228 | Gm38000 | ENSEMBL:ENSMUSG00000103484 | MGI:5611228 | unclassified gene | predicted gene, 38000 |
| 3 | gene | 51.87299 | 51.87584 | negative | MGI_C57BL6J_5610903 | Gm37675 | ENSEMBL:ENSMUSG00000102620 | MGI:5610903 | unclassified gene | predicted gene, 37675 |
| 3 | gene | 51.88428 | 51.88689 | negative | MGI_C57BL6J_5611480 | Gm38252 | ENSEMBL:ENSMUSG00000102786 | MGI:5611480 | unclassified gene | predicted gene, 38252 |
| 3 | pseudogene | 51.90699 | 51.90882 | negative | MGI_C57BL6J_3708672 | Gm10728 | ENSEMBL:ENSMUSG00000103701 | MGI:3708672 | pseudogene | predicted gene 10728 |
| 3 | gene | 51.98094 | 51.99007 | positive | MGI_C57BL6J_5012274 | Gm20089 | ENSEMBL:ENSMUSG00000104161 | MGI:5012274 | lncRNA gene | predicted gene, 20089 |
| 3 | gene | 51.98763 | 51.98985 | negative | MGI_C57BL6J_5611283 | Gm38055 | ENSEMBL:ENSMUSG00000102147 | MGI:5611283 | unclassified gene | predicted gene, 38055 |
| 3 | gene | 51.99134 | 51.99196 | negative | MGI_C57BL6J_5610754 | Gm37526 | ENSEMBL:ENSMUSG00000103784 | MGI:5610754 | unclassified gene | predicted gene, 37526 |
| 3 | gene | 52.00074 | 52.00402 | negative | MGI_C57BL6J_5610693 | Gm37465 | ENSEMBL:ENSMUSG00000102423 | MGI:5610693 | unclassified gene | predicted gene, 37465 |
| 3 | gene | 52.05729 | 52.06424 | negative | MGI_C57BL6J_2444420 | C130089K02Rik | ENSEMBL:ENSMUSG00000104339 | MGI:2444420 | unclassified gene | RIKEN cDNA C130089K02 gene |
| 3 | gene | 52.07159 | 52.07339 | negative | MGI_C57BL6J_1921741 | 5430420F09Rik | ENSEMBL:ENSMUSG00000102808 | MGI:1921741 | unclassified gene | RIKEN cDNA 5430420F09 gene |
| 3 | gene | 52.13405 | 52.22200 | negative | MGI_C57BL6J_5610430 | Gm37202 | ENSEMBL:ENSMUSG00000103655 | MGI:5610430 | lncRNA gene | predicted gene, 37202 |
| 3 | gene | 52.17108 | 52.18013 | negative | MGI_C57BL6J_5589233 | Gm30074 | NCBI_Gene:102631843,ENSEMBL:ENSMUSG00000103726 | MGI:5589233 | lncRNA gene | predicted gene, 30074 |
| 3 | gene | 52.18657 | 52.18796 | positive | MGI_C57BL6J_5611360 | Gm38132 | ENSEMBL:ENSMUSG00000104055 | MGI:5611360 | lncRNA gene | predicted gene, 38132 |
| 3 | gene | 52.19825 | 52.20049 | positive | MGI_C57BL6J_5610398 | Gm37170 | ENSEMBL:ENSMUSG00000103170 | MGI:5610398 | lncRNA gene | predicted gene, 37170 |
| 3 | pseudogene | 52.23684 | 52.24694 | negative | MGI_C57BL6J_3031232 | Olfr1398-ps1 | NCBI_Gene:404507,ENSEMBL:ENSMUSG00000102510 | MGI:3031232 | pseudogene | olfactory receptor 1398, pseudogene 1 |
| 3 | gene | 52.23745 | 52.25650 | positive | MGI_C57BL6J_5589284 | Gm30125 | NCBI_Gene:102631910 | MGI:5589284 | lncRNA gene | predicted gene, 30125 |
| 3 | gene | 52.26834 | 52.35322 | positive | MGI_C57BL6J_1890077 | Foxo1 | NCBI_Gene:56458,ENSEMBL:ENSMUSG00000044167 | MGI:1890077 | protein coding gene | forkhead box O1 |
| 3 | gene | 52.26843 | 52.26944 | negative | MGI_C57BL6J_5141867 | Gm20402 | ENSEMBL:ENSMUSG00000092405 | MGI:5141867 | lncRNA gene | predicted gene 20402 |
| 3 | gene | 52.28224 | 52.28431 | positive | MGI_C57BL6J_5611262 | Gm38034 | ENSEMBL:ENSMUSG00000102844 | MGI:5611262 | unclassified gene | predicted gene, 38034 |
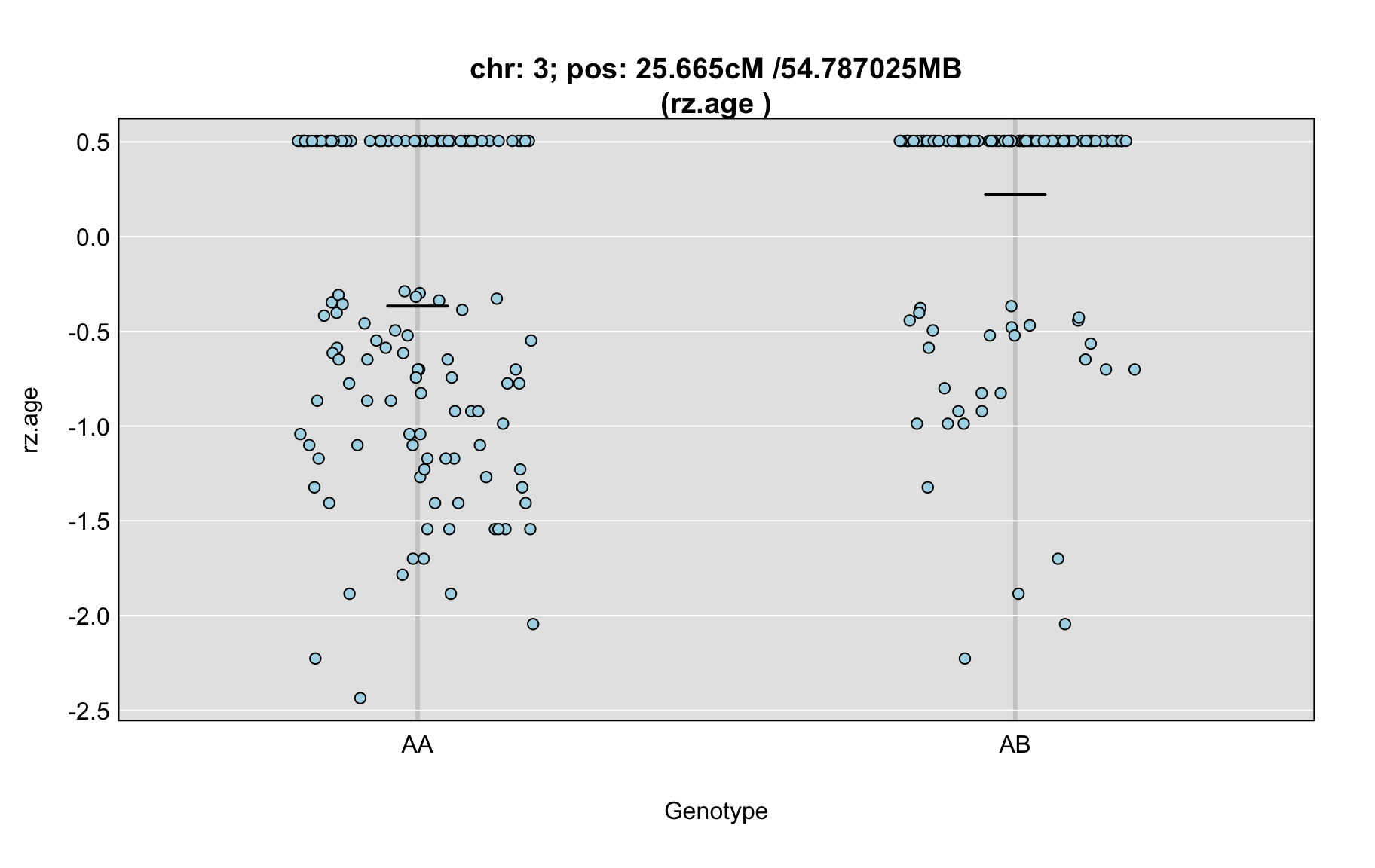
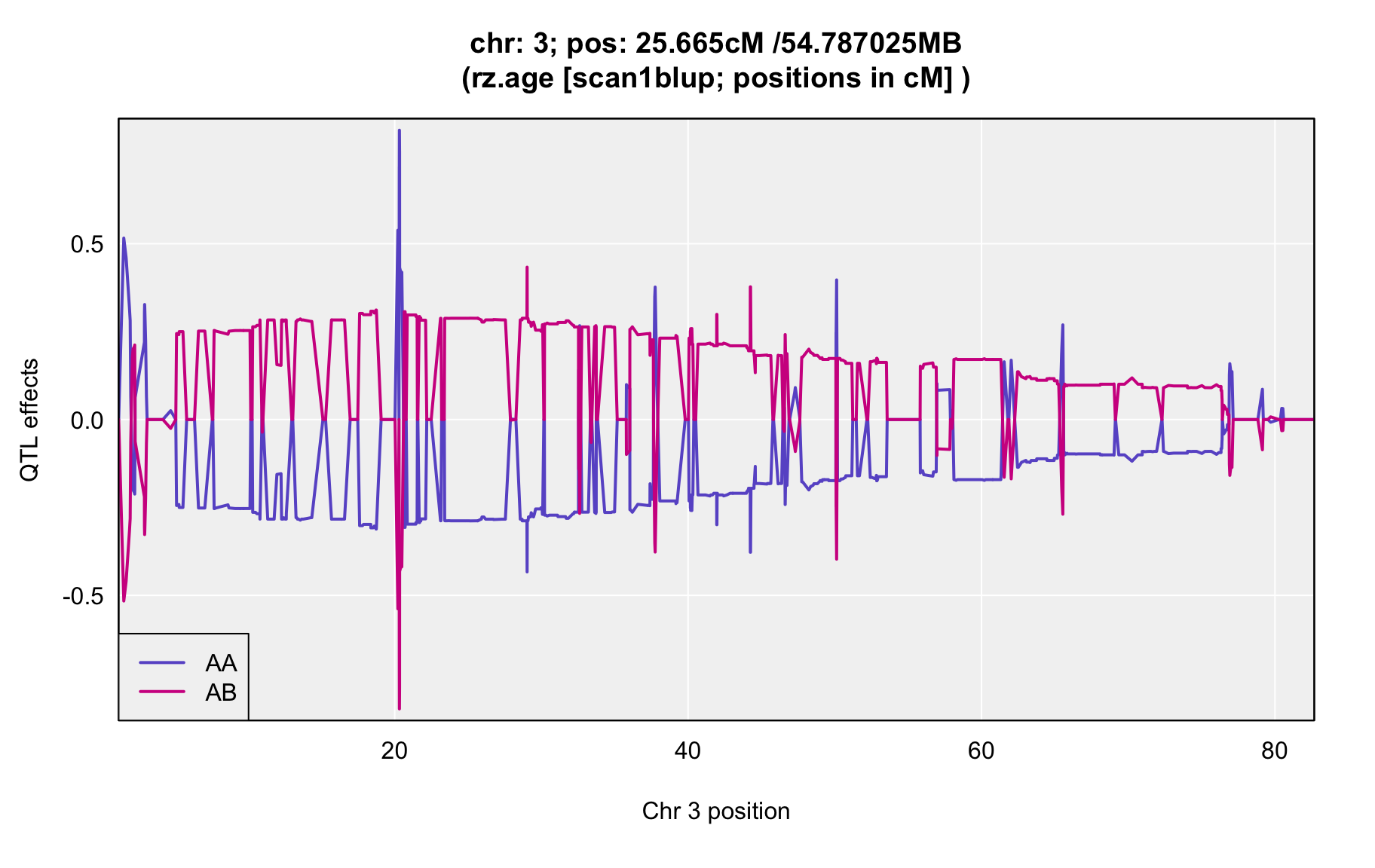
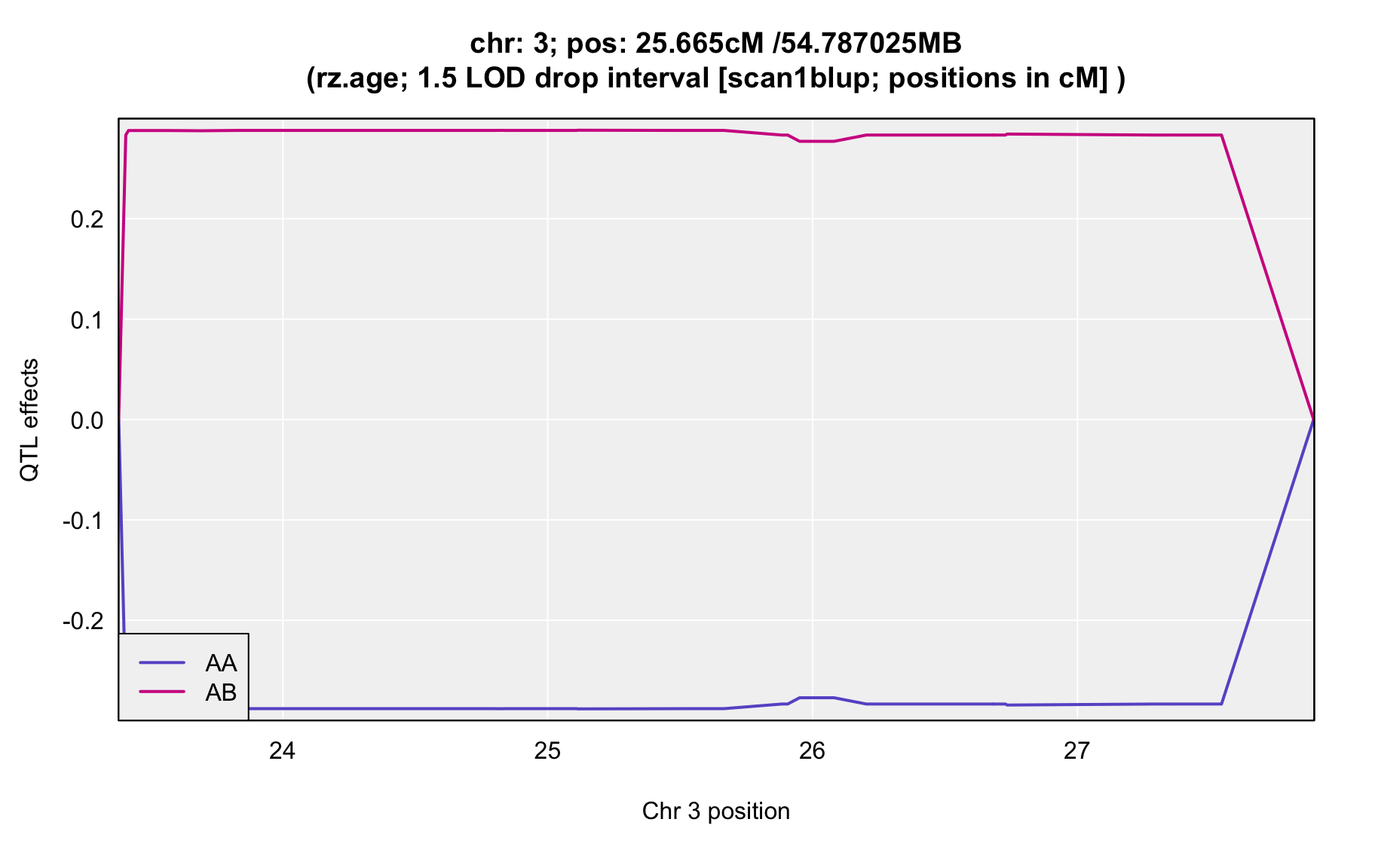
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 52.46717 | 52.49327 | positive | MGI_C57BL6J_5621422 | Gm38537 | NCBI_Gene:102641152 | MGI:5621422 | lncRNA gene | predicted gene, 38537 |
| 3 | gene | 52.50277 | 52.55022 | positive | MGI_C57BL6J_5589332 | Gm30173 | NCBI_Gene:102631981,ENSEMBL:ENSMUSG00000103965 | MGI:5589332 | lncRNA gene | predicted gene, 30173 |
| 3 | gene | 52.58013 | 52.58907 | positive | MGI_C57BL6J_5622932 | Gm40047 | NCBI_Gene:105244440 | MGI:5622932 | lncRNA gene | predicted gene, 40047 |
| 3 | pseudogene | 52.61282 | 52.61402 | positive | MGI_C57BL6J_3704216 | Gm10293 | NCBI_Gene:100039762,ENSEMBL:ENSMUSG00000070490 | MGI:3704216 | pseudogene | predicted pseudogene 10293 |
| 3 | gene | 52.64989 | 52.70971 | positive | MGI_C57BL6J_5589394 | Gm30235 | NCBI_Gene:102632062 | MGI:5589394 | lncRNA gene | predicted gene, 30235 |
| 3 | gene | 52.72265 | 52.72275 | negative | MGI_C57BL6J_5452575 | Gm22798 | ENSEMBL:ENSMUSG00000065926 | MGI:5452575 | snRNA gene | predicted gene, 22798 |
| 3 | gene | 52.73972 | 52.77666 | positive | MGI_C57BL6J_3780614 | Gm2447 | NCBI_Gene:329639,ENSEMBL:ENSMUSG00000102785 | MGI:3780614 | lncRNA gene | predicted gene 2447 |
| 3 | gene | 52.78900 | 52.79781 | negative | MGI_C57BL6J_5589451 | Gm30292 | NCBI_Gene:102632139,ENSEMBL:ENSMUSG00000104109 | MGI:5589451 | lncRNA gene | predicted gene, 30292 |
| 3 | gene | 52.90676 | 52.92767 | negative | MGI_C57BL6J_5434106 | Gm20750 | NCBI_Gene:622124,ENSEMBL:ENSMUSG00000104401 | MGI:5434106 | lncRNA gene | predicted gene, 20750 |
| 3 | gene | 52.93325 | 52.95628 | positive | MGI_C57BL6J_5589510 | Gm30351 | NCBI_Gene:102632211 | MGI:5589510 | lncRNA gene | predicted gene, 30351 |
| 3 | gene | 52.95618 | 52.95789 | negative | MGI_C57BL6J_5589561 | Gm30402 | NCBI_Gene:102632287 | MGI:5589561 | lncRNA gene | predicted gene, 30402 |
| 3 | pseudogene | 52.96005 | 52.96037 | negative | MGI_C57BL6J_5610235 | Gm37007 | ENSEMBL:ENSMUSG00000103886 | MGI:5610235 | pseudogene | predicted gene, 37007 |
| 3 | gene | 52.98188 | 53.01724 | negative | MGI_C57BL6J_1914792 | Cog6 | NCBI_Gene:67542,ENSEMBL:ENSMUSG00000027742 | MGI:1914792 | protein coding gene | component of oligomeric golgi complex 6 |
| 3 | gene | 53.03858 | 53.04176 | negative | MGI_C57BL6J_5663038 | Gm42901 | ENSEMBL:ENSMUSG00000104850 | MGI:5663038 | lncRNA gene | predicted gene 42901 |
| 3 | gene | 53.04153 | 53.26168 | positive | MGI_C57BL6J_1920048 | Lhfp | NCBI_Gene:108927,ENSEMBL:ENSMUSG00000048332 | MGI:1920048 | protein coding gene | lipoma HMGIC fusion partner |
| 3 | gene | 53.07216 | 53.07524 | negative | MGI_C57BL6J_5663035 | Gm42898 | ENSEMBL:ENSMUSG00000104828 | MGI:5663035 | unclassified gene | predicted gene 42898 |
| 3 | gene | 53.08322 | 53.08588 | positive | MGI_C57BL6J_5663036 | Gm42899 | ENSEMBL:ENSMUSG00000104687 | MGI:5663036 | unclassified gene | predicted gene 42899 |
| 3 | gene | 53.09900 | 53.10318 | positive | MGI_C57BL6J_2443940 | C130075A20Rik | ENSEMBL:ENSMUSG00000104735 | MGI:2443940 | unclassified gene | RIKEN cDNA C130075A20 gene |
| 3 | gene | 53.10967 | 53.11012 | positive | MGI_C57BL6J_5663941 | Gm43804 | ENSEMBL:ENSMUSG00000104615 | MGI:5663941 | unclassified gene | predicted gene 43804 |
| 3 | gene | 53.11040 | 53.11341 | negative | MGI_C57BL6J_5663940 | Gm43803 | ENSEMBL:ENSMUSG00000104968 | MGI:5663940 | lncRNA gene | predicted gene 43803 |
| 3 | gene | 53.11217 | 53.11299 | positive | MGI_C57BL6J_5663942 | Gm43805 | ENSEMBL:ENSMUSG00000105971 | MGI:5663942 | unclassified gene | predicted gene 43805 |
| 3 | pseudogene | 53.34649 | 53.34723 | negative | MGI_C57BL6J_5663609 | Gm43472 | ENSEMBL:ENSMUSG00000106660 | MGI:5663609 | pseudogene | predicted gene 43472 |
| 3 | gene | 53.39575 | 53.39850 | positive | MGI_C57BL6J_5622933 | Gm40048 | NCBI_Gene:105244441 | MGI:5622933 | lncRNA gene | predicted gene, 40048 |
| 3 | pseudogene | 53.40161 | 53.42457 | negative | MGI_C57BL6J_5663608 | Gm43471 | NCBI_Gene:108168853,ENSEMBL:ENSMUSG00000105397 | MGI:5663608 | pseudogene | predicted gene 43471 |
| 3 | gene | 53.44858 | 53.46333 | negative | MGI_C57BL6J_2444520 | Nhlrc3 | NCBI_Gene:212114,ENSEMBL:ENSMUSG00000042997 | MGI:2444520 | protein coding gene | NHL repeat containing 3 |
| 3 | gene | 53.45262 | 53.45805 | positive | MGI_C57BL6J_3801870 | Gm16206 | NCBI_Gene:102632669,ENSEMBL:ENSMUSG00000085174 | MGI:3801870 | lncRNA gene | predicted gene 16206 |
| 3 | gene | 53.46367 | 53.48175 | positive | MGI_C57BL6J_1919933 | Proser1 | NCBI_Gene:212127,ENSEMBL:ENSMUSG00000049504 | MGI:1919933 | protein coding gene | proline and serine rich 1 |
| 3 | gene | 53.48865 | 53.50850 | positive | MGI_C57BL6J_2388072 | Stoml3 | NCBI_Gene:229277,ENSEMBL:ENSMUSG00000027744 | MGI:2388072 | protein coding gene | stomatin (Epb7.2)-like 3 |
| 3 | gene | 53.50893 | 53.51010 | positive | MGI_C57BL6J_1924132 | 6430500D05Rik | ENSEMBL:ENSMUSG00000105481 | MGI:1924132 | unclassified gene | RIKEN cDNA 6430500D05 gene |
| 3 | gene | 53.51200 | 53.51213 | positive | MGI_C57BL6J_5453452 | Gm23675 | ENSEMBL:ENSMUSG00000087936 | MGI:5453452 | snoRNA gene | predicted gene, 23675 |
| 3 | gene | 53.51394 | 53.65736 | negative | MGI_C57BL6J_2444465 | Frem2 | NCBI_Gene:242022,ENSEMBL:ENSMUSG00000037016 | MGI:2444465 | protein coding gene | Fras1 related extracellular matrix protein 2 |
| 3 | pseudogene | 53.66666 | 53.66739 | negative | MGI_C57BL6J_3648425 | Gm6073 | NCBI_Gene:606547,ENSEMBL:ENSMUSG00000105840 | MGI:3648425 | pseudogene | predicted gene 6073 |
| 3 | pseudogene | 53.69237 | 53.69292 | negative | MGI_C57BL6J_3649038 | Gm6204 | NCBI_Gene:621155,ENSEMBL:ENSMUSG00000105879 | MGI:3649038 | pseudogene | predicted gene 6204 |
| 3 | gene | 53.70304 | 53.73576 | positive | MGI_C57BL6J_5589894 | Gm30735 | NCBI_Gene:102632743,ENSEMBL:ENSMUSG00000105168 | MGI:5589894 | lncRNA gene | predicted gene, 30735 |
| 3 | gene | 53.73622 | 53.73783 | positive | MGI_C57BL6J_3642187 | Gm10727 | ENSEMBL:ENSMUSG00000105444 | MGI:3642187 | unclassified gene | predicted gene 10727 |
| 3 | gene | 53.75673 | 53.77295 | positive | MGI_C57BL6J_2443105 | C820005J03Rik | ENSEMBL:ENSMUSG00000106270 | MGI:2443105 | lncRNA gene | RIKEN cDNA C820005J03 gene |
| 3 | gene | 53.84341 | 53.84351 | positive | MGI_C57BL6J_5454628 | Gm24851 | ENSEMBL:ENSMUSG00000088653 | MGI:5454628 | snRNA gene | predicted gene, 24851 |
| 3 | gene | 53.84509 | 53.84528 | positive | MGI_C57BL6J_3779200 | Gm10985 | ENSEMBL:ENSMUSG00000078742 | MGI:3779200 | protein coding gene | predicted gene 10985 |
| 3 | gene | 53.85338 | 53.86383 | negative | MGI_C57BL6J_1915140 | Ufm1 | NCBI_Gene:67890,ENSEMBL:ENSMUSG00000027746 | MGI:1915140 | protein coding gene | ubiquitin-fold modifier 1 |
| 3 | gene | 53.92484 | 53.94353 | positive | MGI_C57BL6J_5590185 | Gm31026 | NCBI_Gene:102633119,ENSEMBL:ENSMUSG00000106577 | MGI:5590185 | lncRNA gene | predicted gene, 31026 |
| 3 | pseudogene | 54.02115 | 54.02195 | positive | MGI_C57BL6J_3645533 | Gm8109 | NCBI_Gene:666444,ENSEMBL:ENSMUSG00000103155 | MGI:3645533 | pseudogene | predicted gene 8109 |
| 3 | pseudogene | 54.02988 | 54.03041 | positive | MGI_C57BL6J_5010203 | Gm18018 | NCBI_Gene:100416282,ENSEMBL:ENSMUSG00000105067 | MGI:5010203 | pseudogene | predicted gene, 18018 |
| 3 | gene | 54.15604 | 54.31847 | positive | MGI_C57BL6J_109525 | Trpc4 | NCBI_Gene:22066,ENSEMBL:ENSMUSG00000027748 | MGI:109525 | protein coding gene | transient receptor potential cation channel, subfamily C, member 4 |
| 3 | gene | 54.35927 | 54.39104 | positive | MGI_C57BL6J_1926321 | Postn | NCBI_Gene:50706,ENSEMBL:ENSMUSG00000027750 | MGI:1926321 | protein coding gene | periostin, osteoblast specific factor |
| 3 | gene | 54.35966 | 54.36101 | negative | MGI_C57BL6J_5590254 | Gm31095 | NCBI_Gene:102633208 | MGI:5590254 | lncRNA gene | predicted gene, 31095 |
| 3 | pseudogene | 54.48141 | 54.48293 | positive | MGI_C57BL6J_3645731 | Gm5641 | NCBI_Gene:434807,ENSEMBL:ENSMUSG00000069014 | MGI:3645731 | pseudogene | predicted gene 5641 |
| 3 | gene | 54.48856 | 54.53489 | negative | MGI_C57BL6J_5590313 | Gm31154 | NCBI_Gene:102633293 | MGI:5590313 | lncRNA gene | predicted gene, 31154 |
| 3 | gene | 54.59055 | 54.60139 | negative | MGI_C57BL6J_5622935 | Gm40050 | NCBI_Gene:105244443 | MGI:5622935 | lncRNA gene | predicted gene, 40050 |
| 3 | gene | 54.63144 | 54.63463 | positive | MGI_C57BL6J_5477172 | Gm26678 | ENSEMBL:ENSMUSG00000097599 | MGI:5477172 | lncRNA gene | predicted gene, 26678 |
| 3 | pseudogene | 54.63288 | 54.63457 | negative | MGI_C57BL6J_3588285 | B020017C02Rik | NCBI_Gene:100503708,ENSEMBL:ENSMUSG00000104831 | MGI:3588285 | pseudogene | RIKEN cDNA B020017C02 gene |
| 3 | gene | 54.69276 | 54.72877 | positive | MGI_C57BL6J_1929651 | Supt20 | NCBI_Gene:56790,ENSEMBL:ENSMUSG00000027751 | MGI:1929651 | protein coding gene | SPT20 SAGA complex component |
| 3 | gene | 54.71318 | 54.71435 | negative | MGI_C57BL6J_5439427 | Gm21958 | ENSEMBL:ENSMUSG00000094487 | MGI:5439427 | protein coding gene | predicted gene, 21958 |
| 3 | gene | 54.72868 | 54.73539 | negative | MGI_C57BL6J_1916889 | Exosc8 | NCBI_Gene:69639,ENSEMBL:ENSMUSG00000027752 | MGI:1916889 | protein coding gene | exosome component 8 |
| 3 | gene | 54.73554 | 54.75132 | positive | MGI_C57BL6J_1913498 | Alg5 | NCBI_Gene:66248,ENSEMBL:ENSMUSG00000036632 | MGI:1913498 | protein coding gene | asparagine-linked glycosylation 5 (dolichyl-phosphate beta-glucosyltransferase) |
| 3 | gene | 54.75545 | 54.80166 | positive | MGI_C57BL6J_1859993 | Smad9 | NCBI_Gene:55994,ENSEMBL:ENSMUSG00000027796 | MGI:1859993 | protein coding gene | SMAD family member 9 |
| 3 | gene | 54.80311 | 54.80779 | negative | MGI_C57BL6J_2180854 | Rfxap | NCBI_Gene:170767,ENSEMBL:ENSMUSG00000036615 | MGI:2180854 | protein coding gene | regulatory factor X-associated protein |
| 3 | pseudogene | 54.80997 | 54.81140 | negative | MGI_C57BL6J_3708673 | Gm10254 | NCBI_Gene:545524,ENSEMBL:ENSMUSG00000069011 | MGI:3708673 | pseudogene | predicted gene 10254 |
| 3 | gene | 54.89707 | 54.91589 | negative | MGI_C57BL6J_3607715 | Sertm1 | NCBI_Gene:329641,ENSEMBL:ENSMUSG00000056306 | MGI:3607715 | protein coding gene | serine rich and transmembrane domain containing 1 |
| 3 | gene | 54.91602 | 54.95326 | positive | MGI_C57BL6J_5590517 | Gm31358 | NCBI_Gene:102633562 | MGI:5590517 | lncRNA gene | predicted gene, 31358 |
| 3 | gene | 55.04547 | 55.05550 | negative | MGI_C57BL6J_108042 | Ccna1 | NCBI_Gene:12427,ENSEMBL:ENSMUSG00000027793 | MGI:108042 | protein coding gene | cyclin A1 |
| 3 | gene | 55.05524 | 55.08400 | positive | MGI_C57BL6J_1918220 | 4931419H13Rik | NCBI_Gene:70970,ENSEMBL:ENSMUSG00000106589 | MGI:1918220 | lncRNA gene | RIKEN cDNA 4931419H13 gene |
| 3 | pseudogene | 55.05735 | 55.05766 | negative | MGI_C57BL6J_5663692 | Gm43555 | ENSEMBL:ENSMUSG00000105977 | MGI:5663692 | pseudogene | predicted gene 43555 |
| 3 | gene | 55.11207 | 55.13733 | positive | MGI_C57BL6J_2139806 | Spg20 | NCBI_Gene:229285,ENSEMBL:ENSMUSG00000036580 | MGI:2139806 | protein coding gene | spastic paraplegia 20, spartin (Troyer syndrome) homolog (human) |
| 3 | gene | 55.13734 | 55.17525 | positive | MGI_C57BL6J_2444356 | Ccdc169 | NCBI_Gene:320604,ENSEMBL:ENSMUSG00000048655 | MGI:2444356 | protein coding gene | coiled-coil domain containing 169 |
| 3 | gene | 55.16231 | 55.16243 | negative | MGI_C57BL6J_5451806 | Gm22029 | ENSEMBL:ENSMUSG00000080590 | MGI:5451806 | miRNA gene | predicted gene, 22029 |
| 3 | gene | 55.18203 | 55.20996 | positive | MGI_C57BL6J_1921684 | Sohlh2 | NCBI_Gene:74434,ENSEMBL:ENSMUSG00000027794 | MGI:1921684 | protein coding gene | spermatogenesis and oogenesis specific basic helix-loop-helix 2 |
| 3 | gene | 55.24236 | 55.53907 | positive | MGI_C57BL6J_1330861 | Dclk1 | NCBI_Gene:13175,ENSEMBL:ENSMUSG00000027797 | MGI:1330861 | protein coding gene | doublecortin-like kinase 1 |
| 3 | gene | 55.29094 | 55.29194 | positive | MGI_C57BL6J_1924494 | 9430012M22Rik | NA | NA | unclassified gene | RIKEN cDNA 9430012M22 gene |
| 3 | gene | 55.30532 | 55.30542 | positive | MGI_C57BL6J_5454909 | Gm25132 | ENSEMBL:ENSMUSG00000064586 | MGI:5454909 | snRNA gene | predicted gene, 25132 |
| 3 | gene | 55.33084 | 55.33297 | positive | MGI_C57BL6J_5012002 | Gm19817 | ENSEMBL:ENSMUSG00000105302 | MGI:5012002 | unclassified gene | predicted gene, 19817 |
| 3 | gene | 55.35240 | 55.38079 | negative | MGI_C57BL6J_5622936 | Gm40051 | NCBI_Gene:105244444 | MGI:5622936 | lncRNA gene | predicted gene, 40051 |
| 3 | gene | 55.37746 | 55.40312 | negative | MGI_C57BL6J_5662746 | Gm42609 | ENSEMBL:ENSMUSG00000105975 | MGI:5662746 | lncRNA gene | predicted gene 42609 |
| 3 | gene | 55.39181 | 55.40420 | negative | MGI_C57BL6J_3642047 | Gm9831 | NCBI_Gene:101056032 | MGI:3642047 | protein coding gene | predicted gene 9831 |
| 3 | gene | 55.46198 | 55.47221 | negative | MGI_C57BL6J_5439423 | Gm21954 | ENSEMBL:ENSMUSG00000094962 | MGI:5439423 | protein coding gene | predicted gene, 21954 |
| 3 | gene | 55.47609 | 55.48162 | negative | MGI_C57BL6J_5590602 | Gm31443 | NCBI_Gene:102633676 | MGI:5590602 | lncRNA gene | predicted gene, 31443 |
| 3 | gene | 55.48379 | 55.48718 | positive | MGI_C57BL6J_5662744 | Gm42607 | ENSEMBL:ENSMUSG00000106354 | MGI:5662744 | unclassified gene | predicted gene 42607 |
| 3 | gene | 55.51757 | 55.51959 | positive | MGI_C57BL6J_5662745 | Gm42608 | ENSEMBL:ENSMUSG00000106651 | MGI:5662745 | unclassified gene | predicted gene 42608 |
| 3 | gene | 55.54045 | 55.54338 | negative | MGI_C57BL6J_5590955 | Gm31796 | NCBI_Gene:102634140 | MGI:5590955 | lncRNA gene | predicted gene, 31796 |
| 3 | gene | 55.54354 | 55.54855 | positive | MGI_C57BL6J_5455181 | Gm25404 | NCBI_Gene:102634218,ENSEMBL:ENSMUSG00000064904 | MGI:5455181 | snRNA gene | predicted gene, 25404 |
| 3 | gene | 55.57492 | 55.57727 | positive | MGI_C57BL6J_5663685 | Gm43548 | ENSEMBL:ENSMUSG00000105627 | MGI:5663685 | unclassified gene | predicted gene 43548 |
| 3 | gene | 55.57811 | 55.58273 | negative | MGI_C57BL6J_5663686 | Gm43549 | NCBI_Gene:108168870,ENSEMBL:ENSMUSG00000106334 | MGI:5663686 | lncRNA gene | predicted gene 43549 |
| 3 | gene | 55.62519 | 56.18427 | negative | MGI_C57BL6J_1347075 | Nbea | NCBI_Gene:26422,ENSEMBL:ENSMUSG00000027799 | MGI:1347075 | protein coding gene | neurobeachin |
| 3 | gene | 55.70589 | 55.71565 | positive | MGI_C57BL6J_5826441 | Gm46804 | NCBI_Gene:108168871 | MGI:5826441 | lncRNA gene | predicted gene, 46804 |
| 3 | gene | 55.72593 | 55.72844 | negative | MGI_C57BL6J_3609650 | D430004P15Rik | NA | NA | unclassified gene | Riken cDNA D430004P15 gene |
| 3 | gene | 55.77534 | 55.78795 | negative | MGI_C57BL6J_3698047 | 9630024C22Rik | NA | NA | unclassified non-coding RNA gene | RIKEN cDNA 9630024C22 gene |
| 3 | gene | 55.78251 | 55.78529 | positive | MGI_C57BL6J_1333773 | Mab21l1 | NCBI_Gene:17116,ENSEMBL:ENSMUSG00000056947 | MGI:1333773 | protein coding gene | mab-21-like 1 |
| 3 | gene | 55.83367 | 55.83446 | negative | MGI_C57BL6J_1924979 | 6720451E02Rik | NA | NA | unclassified gene | RIKEN cDNA 6720451E02 gene |
| 3 | pseudogene | 55.84710 | 55.84840 | positive | MGI_C57BL6J_1918432 | 4933417G07Rik | NCBI_Gene:71182,ENSEMBL:ENSMUSG00000106572 | MGI:1918432 | pseudogene | RIKEN cDNA 4933417G07 gene |
| 3 | gene | 55.85525 | 55.85615 | negative | MGI_C57BL6J_1926125 | B230206L23Rik | NA | NA | unclassified gene | RIKEN cDNA B230206L23 gene |
| 3 | gene | 55.88327 | 56.00032 | positive | MGI_C57BL6J_5591073 | Gm31914 | NCBI_Gene:102634293 | MGI:5591073 | lncRNA gene | predicted gene, 31914 |
| 3 | gene | 55.89031 | 55.93670 | positive | MGI_C57BL6J_5663513 | Gm43376 | ENSEMBL:ENSMUSG00000104677 | MGI:5663513 | lncRNA gene | predicted gene 43376 |
| 3 | gene | 55.95678 | 55.95691 | negative | MGI_C57BL6J_5452561 | Gm22784 | ENSEMBL:ENSMUSG00000077276 | MGI:5452561 | snoRNA gene | predicted gene, 22784 |
| 3 | gene | 56.01492 | 56.01717 | negative | MGI_C57BL6J_5662977 | Gm42840 | ENSEMBL:ENSMUSG00000104938 | MGI:5662977 | unclassified gene | predicted gene 42840 |
| 3 | gene | 56.12316 | 56.12616 | negative | MGI_C57BL6J_5662979 | Gm42842 | ENSEMBL:ENSMUSG00000106211 | MGI:5662979 | unclassified gene | predicted gene 42842 |
| 3 | gene | 56.12860 | 56.13053 | negative | MGI_C57BL6J_5662978 | Gm42841 | ENSEMBL:ENSMUSG00000106528 | MGI:5662978 | unclassified gene | predicted gene 42841 |
| 3 | gene | 56.16395 | 56.16757 | negative | MGI_C57BL6J_5662980 | Gm42843 | ENSEMBL:ENSMUSG00000105293 | MGI:5662980 | unclassified gene | predicted gene 42843 |
| 3 | gene | 56.21791 | 56.22124 | negative | MGI_C57BL6J_5622937 | Gm40052 | NCBI_Gene:105244445 | MGI:5622937 | lncRNA gene | predicted gene, 40052 |
| 3 | gene | 56.25590 | 56.26994 | negative | MGI_C57BL6J_5622938 | Gm40053 | NCBI_Gene:105244446 | MGI:5622938 | lncRNA gene | predicted gene, 40053 |
| 3 | gene | 56.34703 | 56.35755 | negative | MGI_C57BL6J_5622939 | Gm40054 | NCBI_Gene:105244447 | MGI:5622939 | lncRNA gene | predicted gene, 40054 |
| 3 | gene | 56.35224 | 56.35230 | positive | MGI_C57BL6J_102792 | Mcptl | NA | NA | protein coding gene | mast cell protease-like |
| 3 | gene | 56.43600 | 56.43875 | negative | MGI_C57BL6J_5591336 | Gm32177 | NCBI_Gene:102634641 | MGI:5591336 | lncRNA gene | predicted gene, 32177 |
| 3 | gene | 56.44528 | 56.44541 | negative | MGI_C57BL6J_5455504 | Gm25727 | ENSEMBL:ENSMUSG00000077710 | MGI:5455504 | snoRNA gene | predicted gene, 25727 |
| 3 | gene | 56.69842 | 56.70506 | positive | MGI_C57BL6J_5591396 | Gm32237 | NCBI_Gene:102634719 | MGI:5591396 | lncRNA gene | predicted gene, 32237 |
| 3 | pseudogene | 56.82832 | 56.82884 | negative | MGI_C57BL6J_3780790 | Gm2622 | NCBI_Gene:100040139,ENSEMBL:ENSMUSG00000105113 | MGI:3780790 | pseudogene | predicted gene 2622 |
| 3 | gene | 56.88806 | 56.89133 | negative | MGI_C57BL6J_3588185 | F830104G03Rik | NA | NA | unclassified gene | RIKEN cDNA F830104G03 gene |
| 3 | gene | 57.05415 | 57.05428 | positive | MGI_C57BL6J_5452046 | Gm22269 | ENSEMBL:ENSMUSG00000077190 | MGI:5452046 | snoRNA gene | predicted gene, 22269 |
| 3 | pseudogene | 57.13549 | 57.13683 | negative | MGI_C57BL6J_3805962 | Gm8177 | NCBI_Gene:666585,ENSEMBL:ENSMUSG00000097366 | MGI:3805962 | pseudogene | predicted gene 8177 |
| 3 | gene | 57.22772 | 57.22918 | negative | MGI_C57BL6J_1918389 | 4933408K01Rik | NA | NA | unclassified gene | RIKEN cDNA 4933408K01 gene |
| 3 | pseudogene | 57.24925 | 57.24950 | positive | MGI_C57BL6J_5663561 | Gm43424 | ENSEMBL:ENSMUSG00000106286 | MGI:5663561 | pseudogene | predicted gene 43424 |
| 3 | pseudogene | 57.27174 | 57.27237 | negative | MGI_C57BL6J_5010845 | Gm18660 | NCBI_Gene:100417512,ENSEMBL:ENSMUSG00000104534 | MGI:5010845 | pseudogene | predicted gene, 18660 |
| 3 | gene | 57.28561 | 57.30199 | negative | MGI_C57BL6J_104678 | Tm4sf1 | NCBI_Gene:17112,ENSEMBL:ENSMUSG00000027800 | MGI:104678 | protein coding gene | transmembrane 4 superfamily member 1 |
| 3 | pseudogene | 57.36225 | 57.36361 | positive | MGI_C57BL6J_3646382 | Gm5276 | NCBI_Gene:383862,ENSEMBL:ENSMUSG00000105452 | MGI:3646382 | pseudogene | predicted gene 5276 |
| 3 | gene | 57.37929 | 57.39455 | negative | MGI_C57BL6J_5591526 | Gm32367 | NCBI_Gene:102634892 | MGI:5591526 | lncRNA gene | predicted gene, 32367 |
| 3 | gene | 57.42531 | 57.44168 | positive | MGI_C57BL6J_2385173 | Tm4sf4 | NCBI_Gene:229302,ENSEMBL:ENSMUSG00000027801 | MGI:2385173 | protein coding gene | transmembrane 4 superfamily member 4 |
| 3 | gene | 57.45564 | 57.57591 | negative | MGI_C57BL6J_1917649 | Wwtr1 | NCBI_Gene:97064,ENSEMBL:ENSMUSG00000027803 | MGI:1917649 | protein coding gene | WW domain containing transcription regulator 1 |
| 3 | pseudogene | 57.52655 | 57.53048 | positive | MGI_C57BL6J_3802068 | Gm16016 | NCBI_Gene:105244448,ENSEMBL:ENSMUSG00000083538 | MGI:3802068 | pseudogene | predicted gene 16016 |
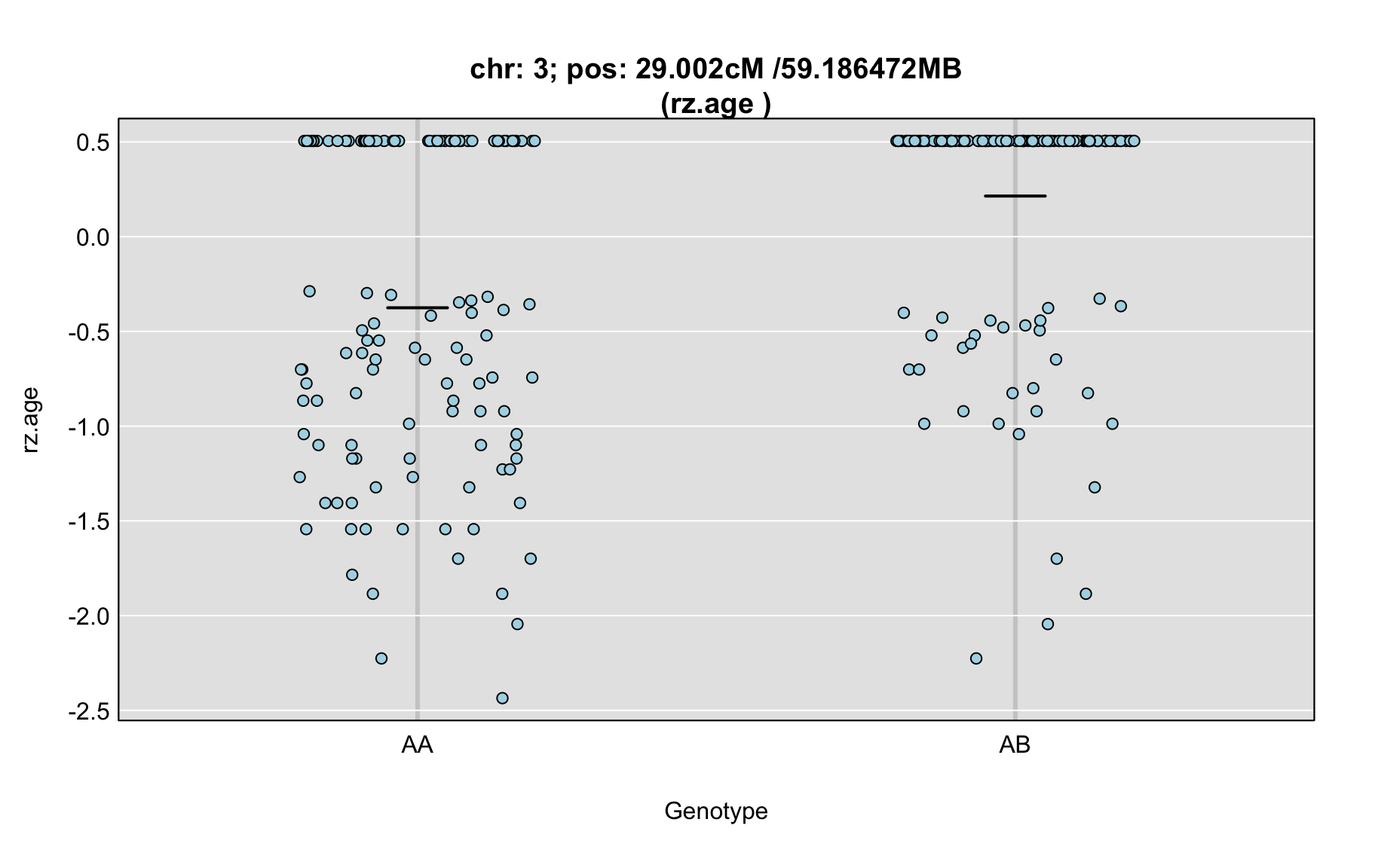
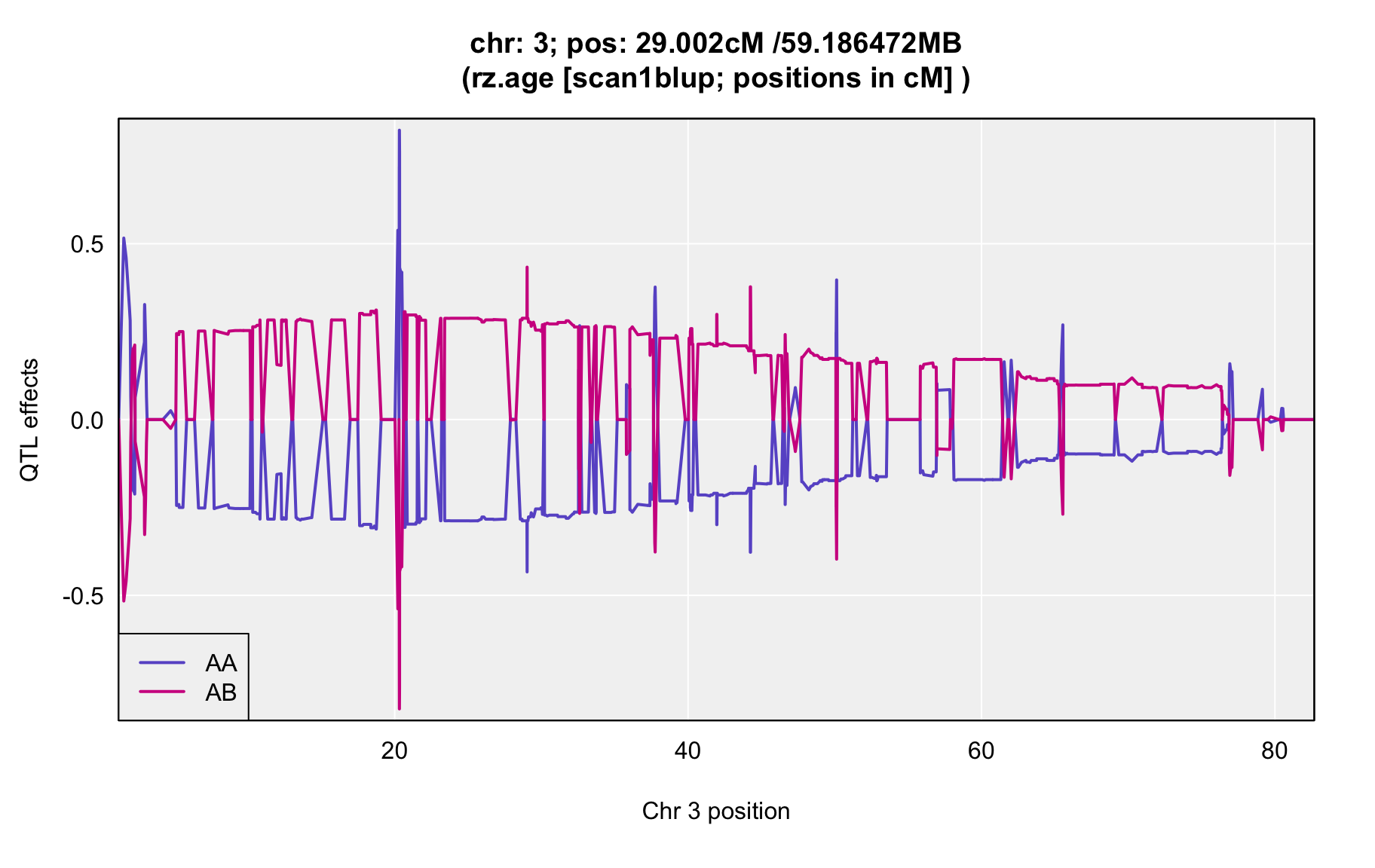
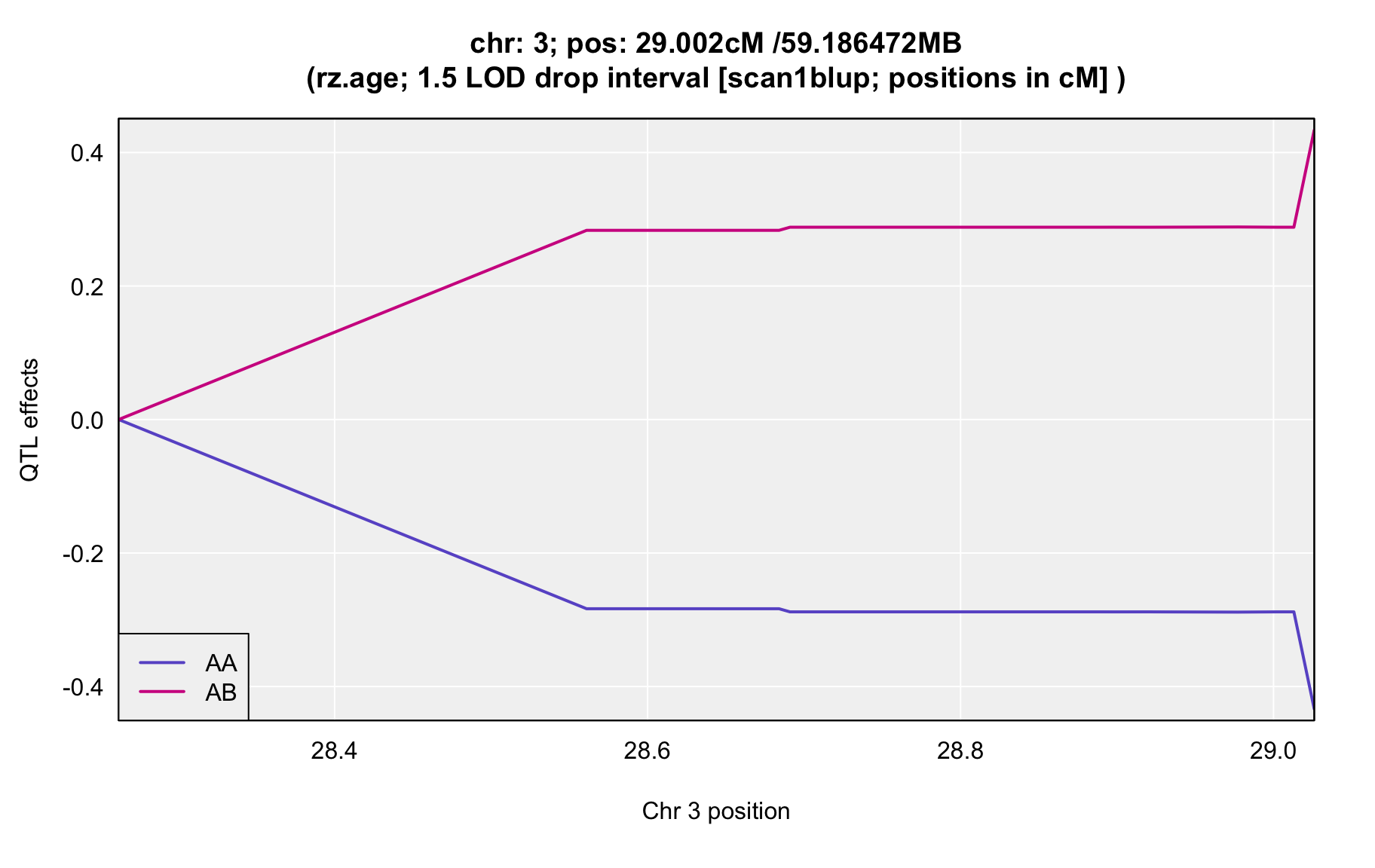
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 57.95198 | 58.03171 | negative | MGI_C57BL6J_5622940 | Gm40055 | NCBI_Gene:105244449,ENSEMBL:ENSMUSG00000105003 | MGI:5622940 | lncRNA gene | predicted gene, 40055 |
| 3 | gene | 58.07972 | 58.08183 | positive | MGI_C57BL6J_5662650 | Gm42513 | ENSEMBL:ENSMUSG00000105065 | MGI:5662650 | unclassified gene | predicted gene 42513 |
| 3 | gene | 58.08960 | 58.16235 | positive | MGI_C57BL6J_5622941 | Gm40056 | NCBI_Gene:105244450 | MGI:5622941 | lncRNA gene | predicted gene, 40056 |
| 3 | pseudogene | 58.10256 | 58.10430 | negative | MGI_C57BL6J_3643179 | Gm5537 | NCBI_Gene:433594,ENSEMBL:ENSMUSG00000069008 | MGI:3643179 | pseudogene | predicted gene 5537 |
| 3 | gene | 58.11705 | 58.16762 | positive | MGI_C57BL6J_1925506 | 4921539H07Rik | ENSEMBL:ENSMUSG00000104586 | MGI:1925506 | lncRNA gene | RIKEN cDNA 4921539H07 gene |
| 3 | gene | 58.14171 | 58.16381 | negative | MGI_C57BL6J_1915100 | 1700007F19Rik | NCBI_Gene:67850,ENSEMBL:ENSMUSG00000100666 | MGI:1915100 | lncRNA gene | RIKEN cDNA 1700007F19 gene |
| 3 | gene | 58.21192 | 58.21337 | positive | MGI_C57BL6J_5622942 | Gm40057 | NCBI_Gene:105244451 | MGI:5622942 | lncRNA gene | predicted gene, 40057 |
| 3 | gene | 58.29025 | 58.29036 | negative | MGI_C57BL6J_5455943 | Gm26166 | ENSEMBL:ENSMUSG00000065722 | MGI:5455943 | snRNA gene | predicted gene, 26166 |
| 3 | gene | 58.30186 | 58.31762 | positive | MGI_C57BL6J_5592006 | Gm32847 | NCBI_Gene:102635539 | MGI:5592006 | lncRNA gene | predicted gene, 32847 |
| 3 | pseudogene | 58.31494 | 58.31531 | negative | MGI_C57BL6J_5663867 | Gm43730 | ENSEMBL:ENSMUSG00000106176 | MGI:5663867 | pseudogene | predicted gene 43730 |
| 3 | pseudogene | 58.40465 | 58.40512 | positive | MGI_C57BL6J_5012313 | Gm20128 | NCBI_Gene:100504238,ENSEMBL:ENSMUSG00000105154 | MGI:5012313 | pseudogene | predicted gene, 20128 |
| 3 | gene | 58.41472 | 58.46679 | positive | MGI_C57BL6J_1919283 | Tsc22d2 | NCBI_Gene:72033,ENSEMBL:ENSMUSG00000027806 | MGI:1919283 | protein coding gene | TSC22 domain family, member 2 |
| 3 | gene | 58.43268 | 58.43301 | positive | MGI_C57BL6J_1338064 | Gdap9 | NA | NA | unclassified gene | ganglioside-induced differentiation-associated-protein 9 |
| 3 | gene | 58.51982 | 58.52589 | negative | MGI_C57BL6J_92638 | Serp1 | NCBI_Gene:28146,ENSEMBL:ENSMUSG00000027808 | MGI:92638 | protein coding gene | stress-associated endoplasmic reticulum protein 1 |
| 3 | gene | 58.52582 | 58.55750 | positive | MGI_C57BL6J_1098684 | Eif2a | NCBI_Gene:229317,ENSEMBL:ENSMUSG00000027810 | MGI:1098684 | protein coding gene | eukaryotic translation initiation factor 2A |
| 3 | gene | 58.52693 | 58.52812 | positive | MGI_C57BL6J_1925148 | 6720482G16Rik | ENSEMBL:ENSMUSG00000106553 | MGI:1925148 | unclassified gene | RIKEN cDNA 6720482G16 gene |
| 3 | gene | 58.57664 | 58.59354 | positive | MGI_C57BL6J_1916477 | Selenot | NCBI_Gene:69227,ENSEMBL:ENSMUSG00000075700 | MGI:1916477 | protein coding gene | selenoprotein T |
| 3 | gene | 58.61630 | 58.63721 | negative | MGI_C57BL6J_3588212 | Erich6 | NCBI_Gene:545527,ENSEMBL:ENSMUSG00000070471 | MGI:3588212 | protein coding gene | glutamate rich 6 |
| 3 | gene | 58.63671 | 58.63749 | positive | MGI_C57BL6J_5826443 | Gm46806 | NCBI_Gene:108168873 | MGI:5826443 | lncRNA gene | predicted gene, 46806 |
| 3 | pseudogene | 58.65204 | 58.65404 | positive | MGI_C57BL6J_3779790 | Gm8234 | NCBI_Gene:666680,ENSEMBL:ENSMUSG00000105021 | MGI:3779790 | pseudogene | predicted gene 8234 |
| 3 | pseudogene | 58.66547 | 58.66664 | negative | MGI_C57BL6J_2139888 | AU022133 | NCBI_Gene:100417266,ENSEMBL:ENSMUSG00000105336 | MGI:2139888 | pseudogene | expressed sequence AU022133 |
| 3 | gene | 58.67494 | 58.69244 | negative | MGI_C57BL6J_108062 | Siah2 | NCBI_Gene:20439,ENSEMBL:ENSMUSG00000036432 | MGI:108062 | protein coding gene | siah E3 ubiquitin protein ligase 2 |
| 3 | gene | 58.69259 | 58.79733 | positive | MGI_C57BL6J_1923132 | 4930593A02Rik | NCBI_Gene:71301,ENSEMBL:ENSMUSG00000097725 | MGI:1923132 | lncRNA gene | RIKEN cDNA 4930593A02 gene |
| 3 | gene | 58.69756 | 58.69867 | positive | MGI_C57BL6J_1925070 | A930028O11Rik | ENSEMBL:ENSMUSG00000105226 | MGI:1925070 | unclassified gene | RIKEN cDNA A930028O11 gene |
| 3 | pseudogene | 58.76149 | 58.76275 | negative | MGI_C57BL6J_5592251 | Gm33092 | NCBI_Gene:102635856 | MGI:5592251 | pseudogene | predicted gene, 33092 |
| 3 | pseudogene | 58.78841 | 58.82231 | negative | MGI_C57BL6J_2140018 | Mindy4b-ps | NCBI_Gene:100040293,ENSEMBL:ENSMUSG00000101860 | MGI:2140018 | pseudogene | MINDY lysine 48 deubiquitinase 4B, pseudogene |
| 3 | gene | 58.82499 | 58.83590 | positive | MGI_C57BL6J_5622944 | Gm40059 | NCBI_Gene:105244453 | MGI:5622944 | lncRNA gene | predicted gene, 40059 |
| 3 | gene | 58.84210 | 58.86675 | positive | MGI_C57BL6J_5622945 | Gm40060 | NCBI_Gene:105244454 | MGI:5622945 | lncRNA gene | predicted gene, 40060 |
| 3 | gene | 58.84403 | 58.88534 | negative | MGI_C57BL6J_2388124 | Clrn1 | NCBI_Gene:229320,ENSEMBL:ENSMUSG00000043850 | MGI:2388124 | protein coding gene | clarin 1 |
| 3 | gene | 58.97563 | 58.97573 | positive | MGI_C57BL6J_5452268 | Gm22491 | ENSEMBL:ENSMUSG00000087815 | MGI:5452268 | snRNA gene | predicted gene, 22491 |
| 3 | gene | 58.97886 | 58.97903 | negative | MGI_C57BL6J_5455349 | Gm25572 | ENSEMBL:ENSMUSG00000088152 | MGI:5455349 | snRNA gene | predicted gene, 25572 |
| 3 | pseudogene | 58.98713 | 58.98786 | positive | MGI_C57BL6J_3642685 | Rpl13-ps6 | NCBI_Gene:100040416,ENSEMBL:ENSMUSG00000059776 | MGI:3642685 | pseudogene | ribosomal protein L13, pseudogene 6 |
| 3 | gene | 59.00162 | 59.00169 | positive | MGI_C57BL6J_5531175 | Gm27793 | ENSEMBL:ENSMUSG00000099052 | MGI:5531175 | unclassified non-coding RNA gene | predicted gene, 27793 |
| 3 | gene | 59.00247 | 59.00616 | negative | MGI_C57BL6J_5663707 | Gm43570 | ENSEMBL:ENSMUSG00000105945 | MGI:5663707 | lncRNA gene | predicted gene 43570 |
| 3 | gene | 59.00559 | 59.31868 | positive | MGI_C57BL6J_2139916 | Med12l | NCBI_Gene:329650,ENSEMBL:ENSMUSG00000056476 | MGI:2139916 | protein coding gene | mediator complex subunit 12-like |
| 3 | gene | 59.09645 | 59.10182 | negative | MGI_C57BL6J_2442043 | Gpr171 | NCBI_Gene:229323,ENSEMBL:ENSMUSG00000050075 | MGI:2442043 | protein coding gene | G protein-coupled receptor 171 |
| 3 | gene | 59.11386 | 59.15362 | negative | MGI_C57BL6J_2155705 | P2ry14 | NCBI_Gene:140795,ENSEMBL:ENSMUSG00000036381 | MGI:2155705 | protein coding gene | purinergic receptor P2Y, G-protein coupled, 14 |
| 3 | gene | 59.14630 | 59.15363 | negative | MGI_C57BL6J_2444074 | F630111L10Rik | NCBI_Gene:320463 | MGI:2444074 | lncRNA gene | RIKEN cDNA F630111L10 gene |
| 3 | gene | 59.17890 | 59.19547 | negative | MGI_C57BL6J_1934133 | Gpr87 | NCBI_Gene:84111,ENSEMBL:ENSMUSG00000051431 | MGI:1934133 | protein coding gene | G protein-coupled receptor 87 |
| 3 | gene | 59.20789 | 59.21088 | negative | MGI_C57BL6J_1921441 | P2ry13 | NCBI_Gene:74191,ENSEMBL:ENSMUSG00000036362 | MGI:1921441 | protein coding gene | purinergic receptor P2Y, G-protein coupled 13 |
| 3 | gene | 59.21627 | 59.26301 | negative | MGI_C57BL6J_1918089 | P2ry12 | NCBI_Gene:70839,ENSEMBL:ENSMUSG00000036353 | MGI:1918089 | protein coding gene | purinergic receptor P2Y, G-protein coupled 12 |
| 3 | gene | 59.26979 | 59.27131 | positive | MGI_C57BL6J_5663726 | Gm43589 | ENSEMBL:ENSMUSG00000104550 | MGI:5663726 | unclassified gene | predicted gene 43589 |
| 3 | gene | 59.31674 | 59.34444 | negative | MGI_C57BL6J_1923481 | Igsf10 | NCBI_Gene:242050,ENSEMBL:ENSMUSG00000036334 | MGI:1923481 | protein coding gene | immunoglobulin superfamily, member 10 |
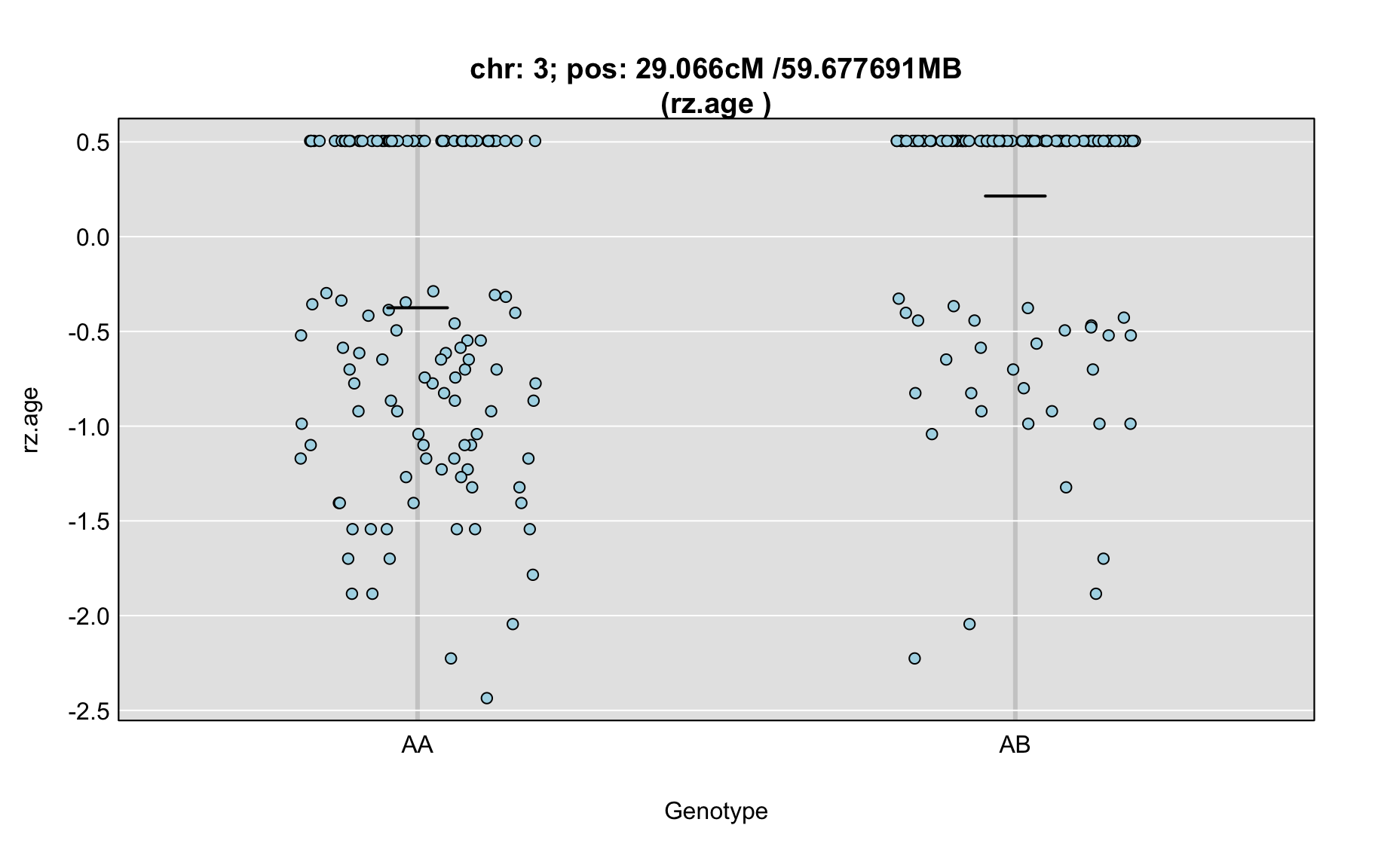
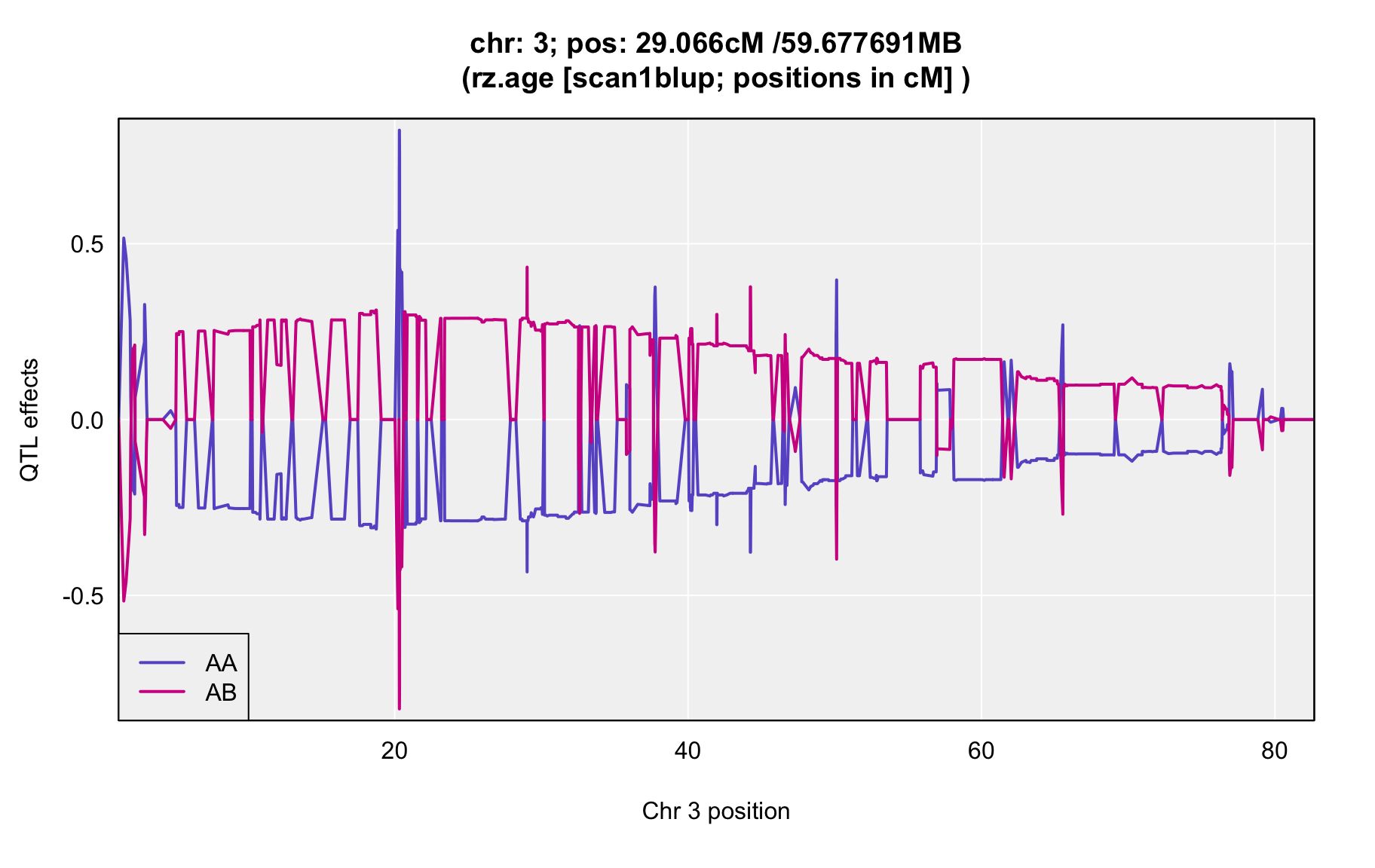
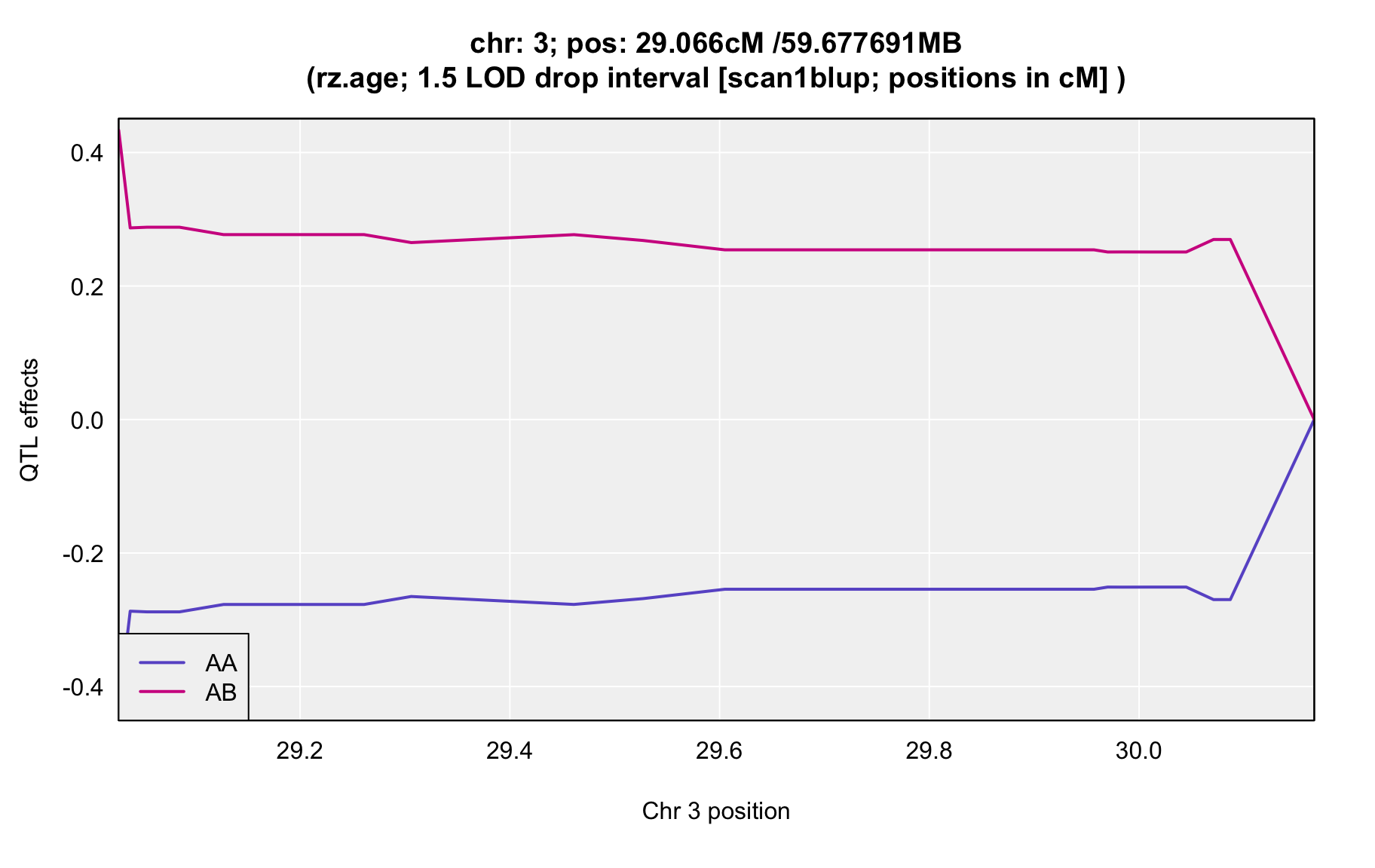
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 59.31674 | 59.34444 | negative | MGI_C57BL6J_1923481 | Igsf10 | NCBI_Gene:242050,ENSEMBL:ENSMUSG00000036334 | MGI:1923481 | protein coding gene | immunoglobulin superfamily, member 10 |
| 3 | gene | 59.34427 | 59.35117 | positive | MGI_C57BL6J_5611414 | Gm38186 | ENSEMBL:ENSMUSG00000103668 | MGI:5611414 | lncRNA gene | predicted gene, 38186 |
| 3 | pseudogene | 59.42453 | 59.42476 | positive | MGI_C57BL6J_5610943 | Gm37715 | ENSEMBL:ENSMUSG00000102247 | MGI:5610943 | pseudogene | predicted gene, 37715 |
| 3 | pseudogene | 59.43154 | 59.43335 | positive | MGI_C57BL6J_3648019 | Gm8276 | NCBI_Gene:666756,ENSEMBL:ENSMUSG00000102550 | MGI:3648019 | pseudogene | predicted gene 8276 |
| 3 | pseudogene | 59.46967 | 59.47409 | negative | MGI_C57BL6J_5611516 | Gm38288 | ENSEMBL:ENSMUSG00000103211 | MGI:5611516 | pseudogene | predicted gene, 38288 |
| 3 | gene | 59.47493 | 59.47500 | negative | MGI_C57BL6J_4414092 | n-TVcac1 | NCBI_Gene:102467620 | MGI:4414092 | tRNA gene | nuclear encoded tRNA valine 1 (anticodon CAC) |
| 3 | pseudogene | 59.48302 | 59.48329 | positive | MGI_C57BL6J_5610999 | Gm37771 | ENSEMBL:ENSMUSG00000104038 | MGI:5610999 | pseudogene | predicted gene, 37771 |
| 3 | pseudogene | 59.50558 | 59.53523 | negative | MGI_C57BL6J_3644725 | Gm4856 | NCBI_Gene:229330,ENSEMBL:ENSMUSG00000103598 | MGI:3644725 | pseudogene | predicted gene 4856 |
| 3 | pseudogene | 59.60245 | 59.63576 | negative | MGI_C57BL6J_3644280 | Gm5709 | NCBI_Gene:435732,ENSEMBL:ENSMUSG00000095128 | MGI:3644280 | pseudogene | predicted gene 5709 |
| 3 | gene | 59.66286 | 59.66528 | positive | MGI_C57BL6J_5611154 | Gm37926 | ENSEMBL:ENSMUSG00000102480 | MGI:5611154 | unclassified gene | predicted gene, 37926 |
| 3 | gene | 59.72979 | 59.75240 | positive | MGI_C57BL6J_3779495 | Aadacl2fm2 | NCBI_Gene:433597,ENSEMBL:ENSMUSG00000090527 | MGI:3779495 | protein coding gene | AADACL2 family member 2 |
| 3 | pseudogene | 59.73675 | 59.73686 | negative | MGI_C57BL6J_5611436 | Gm38208 | ENSEMBL:ENSMUSG00000104471 | MGI:5611436 | pseudogene | predicted gene, 38208 |
| 3 | pseudogene | 59.73718 | 59.73732 | negative | MGI_C57BL6J_5610807 | Gm37579 | ENSEMBL:ENSMUSG00000102390 | MGI:5610807 | pseudogene | predicted gene, 37579 |
| 3 | pseudogene | 59.76721 | 59.76891 | positive | MGI_C57BL6J_3644451 | Gm5539 | NCBI_Gene:433598,ENSEMBL:ENSMUSG00000103986 | MGI:3644451 | pseudogene | predicted gene 5539 |
| 3 | gene | 59.77781 | 59.84682 | positive | MGI_C57BL6J_1921922 | 4930449A18Rik | ENSEMBL:ENSMUSG00000074589 | MGI:1921922 | protein coding gene | RIKEN cDNA 4930449A18 gene |
| 3 | pseudogene | 59.78322 | 59.78863 | negative | MGI_C57BL6J_3780925 | Gm2756 | NCBI_Gene:100040407,ENSEMBL:ENSMUSG00000102757 | MGI:3780925 | pseudogene | predicted gene 2756 |
| 3 | gene | 59.83837 | 59.84682 | positive | MGI_C57BL6J_4361133 | Gm16527 | NCBI_Gene:669780 | MGI:4361133 | protein coding gene | predicted gene, 16527 |
| 3 | gene | 59.86105 | 59.87731 | positive | MGI_C57BL6J_3643798 | Aadacl2fm3 | NCBI_Gene:666803,ENSEMBL:ENSMUSG00000095522 | MGI:3643798 | protein coding gene | AADACL2 family member 3 |
| 3 | gene | 59.92521 | 59.93795 | positive | MGI_C57BL6J_3028051 | Aadacl2fm1 | NCBI_Gene:229333,ENSEMBL:ENSMUSG00000036951 | MGI:3028051 | protein coding gene | AADACL2 family member 1 |
| 3 | pseudogene | 59.95231 | 59.97359 | positive | MGI_C57BL6J_3819177 | Gm9696 | NCBI_Gene:676914,ENSEMBL:ENSMUSG00000070465 | MGI:3819177 | pseudogene | predicted gene 9696 |
| 3 | gene | 60.00674 | 60.02542 | positive | MGI_C57BL6J_3646333 | Aadacl2 | NCBI_Gene:639634,ENSEMBL:ENSMUSG00000091376 | MGI:3646333 | protein coding gene | arylacetamide deacetylase like 2 |
| 3 | pseudogene | 60.02036 | 60.02087 | negative | MGI_C57BL6J_5010612 | Gm18427 | NCBI_Gene:100417154,ENSEMBL:ENSMUSG00000103177 | MGI:5010612 | pseudogene | predicted gene, 18427 |
| 3 | gene | 60.02572 | 60.04016 | positive | MGI_C57BL6J_1915008 | Aadac | NCBI_Gene:67758,ENSEMBL:ENSMUSG00000027761 | MGI:1915008 | protein coding gene | arylacetamide deacetylase |
| 3 | gene | 60.08187 | 60.08757 | positive | MGI_C57BL6J_1934135 | Sucnr1 | NCBI_Gene:84112,ENSEMBL:ENSMUSG00000027762 | MGI:1934135 | protein coding gene | succinate receptor 1 |
| 3 | gene | 60.12764 | 60.12775 | negative | MGI_C57BL6J_5454159 | Gm24382 | ENSEMBL:ENSMUSG00000084647 | MGI:5454159 | snRNA gene | predicted gene, 24382 |
| 3 | gene | 60.29342 | 60.30941 | negative | MGI_C57BL6J_5592809 | Gm33650 | NCBI_Gene:102636642 | MGI:5592809 | lncRNA gene | predicted gene, 33650 |
| 3 | pseudogene | 60.34219 | 60.34309 | negative | MGI_C57BL6J_3779880 | Tgif1-ps | NCBI_Gene:669823,ENSEMBL:ENSMUSG00000107270 | MGI:3779880 | pseudogene | TGFB-induced factor homeobox 1, pseudogene |
| 3 | gene | 60.47279 | 60.62975 | positive | MGI_C57BL6J_1928482 | Mbnl1 | NCBI_Gene:56758,ENSEMBL:ENSMUSG00000027763 | MGI:1928482 | protein coding gene | muscleblind like splicing factor 1 |
| 3 | gene | 60.53575 | 60.53974 | positive | MGI_C57BL6J_5610817 | Gm37589 | ENSEMBL:ENSMUSG00000104235 | MGI:5610817 | unclassified gene | predicted gene, 37589 |
| 3 | gene | 60.57770 | 60.58136 | positive | MGI_C57BL6J_5012001 | Gm19816 | NA | NA | unclassified gene | predicted gene%2c 19816 |
| 3 | gene | 60.60438 | 60.60776 | positive | MGI_C57BL6J_5610716 | Gm37488 | ENSEMBL:ENSMUSG00000104125 | MGI:5610716 | unclassified gene | predicted gene, 37488 |
| 3 | pseudogene | 60.73763 | 60.73779 | negative | MGI_C57BL6J_5611508 | Gm38280 | ENSEMBL:ENSMUSG00000104337 | MGI:5611508 | pseudogene | predicted gene, 38280 |
| 3 | pseudogene | 60.76958 | 60.77164 | negative | MGI_C57BL6J_5592866 | Gm33707 | NCBI_Gene:102636711,ENSEMBL:ENSMUSG00000102298 | MGI:5592866 | pseudogene | predicted gene, 33707 |
| 3 | gene | 60.78148 | 61.00379 | negative | MGI_C57BL6J_5610263 | Gm37035 | NCBI_Gene:105244455,ENSEMBL:ENSMUSG00000102564 | MGI:5610263 | lncRNA gene | predicted gene, 37035 |
| 3 | gene | 60.78836 | 60.79473 | positive | MGI_C57BL6J_5611554 | Gm38326 | NCBI_Gene:102642162,ENSEMBL:ENSMUSG00000103587 | MGI:5611554 | lncRNA gene | predicted gene, 38326 |
| 3 | pseudogene | 60.86071 | 60.86109 | positive | MGI_C57BL6J_5610213 | Gm36985 | ENSEMBL:ENSMUSG00000102490 | MGI:5610213 | pseudogene | predicted gene, 36985 |
| 3 | pseudogene | 60.87700 | 60.87884 | positive | MGI_C57BL6J_3644852 | Gm8325 | NCBI_Gene:666853,ENSEMBL:ENSMUSG00000027694 | MGI:3644852 | pseudogene | predicted pseudogene 8325 |
| 3 | gene | 61.00279 | 61.00898 | positive | MGI_C57BL6J_105049 | P2ry1 | NCBI_Gene:18441,ENSEMBL:ENSMUSG00000027765 | MGI:105049 | protein coding gene | purinergic receptor P2Y, G-protein coupled 1 |
| 3 | pseudogene | 61.29116 | 61.29145 | negative | MGI_C57BL6J_5610947 | Gm37719 | ENSEMBL:ENSMUSG00000102242 | MGI:5610947 | pseudogene | predicted gene, 37719 |
| 3 | pseudogene | 61.32849 | 61.32923 | negative | MGI_C57BL6J_5012086 | Gm19901 | NCBI_Gene:100503798,ENSEMBL:ENSMUSG00000102667 | MGI:5012086 | pseudogene | predicted gene, 19901 |
| 3 | gene | 61.35855 | 61.36190 | negative | MGI_C57BL6J_5610924 | Gm37696 | ENSEMBL:ENSMUSG00000103473 | MGI:5610924 | lncRNA gene | predicted gene, 37696 |
| 3 | gene | 61.36164 | 61.36870 | positive | MGI_C57BL6J_1921262 | Rap2b | NCBI_Gene:74012,ENSEMBL:ENSMUSG00000036894 | MGI:1921262 | protein coding gene | RAP2B, member of RAS oncogene family |
| 3 | gene | 61.36225 | 61.36595 | negative | MGI_C57BL6J_3697707 | B430305J03Rik | ENSEMBL:ENSMUSG00000053706 | MGI:3697707 | protein coding gene | RIKEN cDNA B430305J03 gene |
| 3 | gene | 61.39215 | 61.41303 | positive | MGI_C57BL6J_5622946 | Gm40061 | NCBI_Gene:105244456 | MGI:5622946 | lncRNA gene | predicted gene, 40061 |
| 3 | pseudogene | 61.46166 | 61.46288 | negative | MGI_C57BL6J_3643135 | Gm8349 | NCBI_Gene:666891,ENSEMBL:ENSMUSG00000098208 | MGI:3643135 | pseudogene | predicted gene 8349 |
| 3 | pseudogene | 62.06754 | 62.06979 | negative | MGI_C57BL6J_3780107 | Gm9700 | NCBI_Gene:676965,ENSEMBL:ENSMUSG00000104489 | MGI:3780107 | pseudogene | predicted gene 9700 |
| 3 | gene | 62.12321 | 62.12356 | positive | MGI_C57BL6J_5610632 | Gm37404 | ENSEMBL:ENSMUSG00000103322 | MGI:5610632 | unclassified gene | predicted gene, 37404 |
| 3 | gene | 62.33752 | 62.34053 | negative | MGI_C57BL6J_3704218 | 9330121J05Rik | ENSEMBL:ENSMUSG00000103502 | MGI:3704218 | lncRNA gene | RIKEN cDNA 9330121J05 gene |
| 3 | gene | 62.33834 | 62.46222 | positive | MGI_C57BL6J_1918053 | Arhgef26 | NCBI_Gene:622434,ENSEMBL:ENSMUSG00000036885 | MGI:1918053 | protein coding gene | Rho guanine nucleotide exchange factor (GEF) 26 |
| 3 | gene | 62.34139 | 62.35052 | negative | MGI_C57BL6J_5593082 | Gm33923 | NCBI_Gene:102637008 | MGI:5593082 | lncRNA gene | predicted gene, 33923 |
| 3 | gene | 62.46801 | 62.50703 | negative | MGI_C57BL6J_1919412 | Dhx36 | NCBI_Gene:72162,ENSEMBL:ENSMUSG00000027770 | MGI:1919412 | protein coding gene | DEAH (Asp-Glu-Ala-His) box polypeptide 36 |
| 3 | gene | 62.52908 | 62.60544 | negative | MGI_C57BL6J_2443628 | Gpr149 | NCBI_Gene:229357,ENSEMBL:ENSMUSG00000043441 | MGI:2443628 | protein coding gene | G protein-coupled receptor 149 |
| 3 | gene | 62.60493 | 62.76393 | positive | MGI_C57BL6J_5593207 | Gm34048 | NCBI_Gene:102637165 | MGI:5593207 | lncRNA gene | predicted gene, 34048 |
| 3 | pseudogene | 62.76144 | 62.76202 | positive | MGI_C57BL6J_5010567 | Gm18382 | NCBI_Gene:100417044,ENSEMBL:ENSMUSG00000103764 | MGI:5010567 | pseudogene | predicted gene, 18382 |
| 3 | gene | 62.77198 | 62.86064 | positive | MGI_C57BL6J_5622947 | Gm40062 | NCBI_Gene:105244457 | MGI:5622947 | lncRNA gene | predicted gene, 40062 |
| 3 | gene | 62.86768 | 62.88900 | positive | MGI_C57BL6J_5622948 | Gm40063 | NCBI_Gene:105244458 | MGI:5622948 | lncRNA gene | predicted gene, 40063 |
| 3 | pseudogene | 62.92376 | 62.92426 | negative | MGI_C57BL6J_3780108 | Gm9701 | NCBI_Gene:676984,ENSEMBL:ENSMUSG00000063001 | MGI:3780108 | pseudogene | predicted gene 9701 |
| 3 | gene | 63.15222 | 63.15236 | positive | MGI_C57BL6J_5452210 | Gm22433 | ENSEMBL:ENSMUSG00000089175 | MGI:5452210 | snoRNA gene | predicted gene, 22433 |
| 3 | pseudogene | 63.17311 | 63.17386 | negative | MGI_C57BL6J_3645001 | Gm8388 | NCBI_Gene:666967,ENSEMBL:ENSMUSG00000102270 | MGI:3645001 | pseudogene | predicted gene 8388 |
| 3 | gene | 63.23282 | 63.24545 | negative | MGI_C57BL6J_5622949 | Gm40064 | NCBI_Gene:105244459 | MGI:5622949 | lncRNA gene | predicted gene, 40064 |
| 3 | gene | 63.24154 | 63.38603 | positive | MGI_C57BL6J_97004 | Mme | NCBI_Gene:17380,ENSEMBL:ENSMUSG00000027820 | MGI:97004 | protein coding gene | membrane metallo endopeptidase |
| 3 | gene | 63.43925 | 63.45790 | positive | MGI_C57BL6J_5593399 | Gm34240 | NCBI_Gene:102637430,ENSEMBL:ENSMUSG00000104336 | MGI:5593399 | lncRNA gene | predicted gene, 34240 |
| 3 | gene | 63.48110 | 63.48391 | negative | MGI_C57BL6J_5593461 | Strit1 | NCBI_Gene:102637511,ENSEMBL:ENSMUSG00000103476 | MGI:5593461 | protein coding gene | small transmembrane regulator of ion transport 1 |
| 3 | gene | 63.48388 | 63.48437 | positive | MGI_C57BL6J_5610507 | Gm37279 | ENSEMBL:ENSMUSG00000102883 | MGI:5610507 | unclassified gene | predicted gene, 37279 |
| 3 | gene | 63.48899 | 63.48910 | positive | MGI_C57BL6J_5454723 | Gm24946 | ENSEMBL:ENSMUSG00000088957 | MGI:5454723 | snRNA gene | predicted gene, 24946 |
| 3 | gene | 63.69620 | 63.89972 | negative | MGI_C57BL6J_2683547 | Plch1 | NCBI_Gene:269437,ENSEMBL:ENSMUSG00000036834 | MGI:2683547 | protein coding gene | phospholipase C, eta 1 |
| 3 | gene | 63.77021 | 63.77031 | positive | MGI_C57BL6J_5453277 | Gm23500 | ENSEMBL:ENSMUSG00000088360 | MGI:5453277 | snRNA gene | predicted gene, 23500 |
| 3 | gene | 63.79610 | 63.83685 | positive | MGI_C57BL6J_5826444 | Gm46807 | NCBI_Gene:108168874 | MGI:5826444 | lncRNA gene | predicted gene, 46807 |
| 3 | pseudogene | 63.82579 | 63.82644 | positive | MGI_C57BL6J_3802047 | Gm16162 | ENSEMBL:ENSMUSG00000090131 | MGI:3802047 | pseudogene | predicted gene 16162 |
| 3 | pseudogene | 63.84312 | 63.84407 | positive | MGI_C57BL6J_3801947 | Gm16129 | ENSEMBL:ENSMUSG00000085290 | MGI:3801947 | pseudogene | predicted gene 16129 |
| 3 | pseudogene | 63.90543 | 63.90573 | positive | MGI_C57BL6J_5593538 | Gm34379 | NCBI_Gene:102637617,ENSEMBL:ENSMUSG00000103905 | MGI:5593538 | pseudogene | predicted gene, 34379 |
| 3 | gene | 63.91468 | 63.92947 | negative | MGI_C57BL6J_3607716 | E130311K13Rik | NCBI_Gene:329659,ENSEMBL:ENSMUSG00000048581 | MGI:3607716 | protein coding gene | RIKEN cDNA E130311K13 gene |
| 3 | gene | 63.93351 | 63.96477 | negative | MGI_C57BL6J_1332247 | Slc33a1 | NCBI_Gene:11416,ENSEMBL:ENSMUSG00000027822 | MGI:1332247 | protein coding gene | solute carrier family 33 (acetyl-CoA transporter), member 1 |
| 3 | gene | 63.96363 | 63.96866 | positive | MGI_C57BL6J_5477344 | Gm26850 | ENSEMBL:ENSMUSG00000097724 | MGI:5477344 | lncRNA gene | predicted gene, 26850 |
| 3 | gene | 63.96482 | 64.02258 | positive | MGI_C57BL6J_2448526 | Gmps | NCBI_Gene:229363,ENSEMBL:ENSMUSG00000027823 | MGI:2448526 | protein coding gene | guanine monophosphate synthetase |
| 3 | pseudogene | 64.05303 | 64.05561 | negative | MGI_C57BL6J_3757653 | Vmn2r-ps3 | NCBI_Gene:100124546,ENSEMBL:ENSMUSG00000090303 | MGI:3757653 | pseudogene | vomeronasal 2, receptor, pseudogene 3 |
| 3 | gene | 64.07977 | 64.22085 | positive | MGI_C57BL6J_3645892 | Vmn2r1 | NCBI_Gene:56544,ENSEMBL:ENSMUSG00000027824 | MGI:3645892 | protein coding gene | vomeronasal 2, receptor 1 |
| 3 | gene | 64.10963 | 64.10974 | positive | MGI_C57BL6J_5451942 | Gm22165 | ENSEMBL:ENSMUSG00000089039 | MGI:5451942 | snRNA gene | predicted gene, 22165 |
| 3 | gene | 64.11528 | 64.17092 | negative | MGI_C57BL6J_3757666 | Vmn2r2 | NCBI_Gene:100125586,ENSEMBL:ENSMUSG00000043897 | MGI:3757666 | protein coding gene | vomeronasal 2, receptor 2 |
| 3 | pseudogene | 64.16339 | 64.16385 | positive | MGI_C57BL6J_3779465 | Gm5139 | NCBI_Gene:380739,ENSEMBL:ENSMUSG00000093447 | MGI:3779465 | pseudogene | predicted gene 5139 |
| 3 | pseudogene | 64.19557 | 64.21368 | negative | MGI_C57BL6J_5010479 | Gm18294 | NCBI_Gene:100416861,ENSEMBL:ENSMUSG00000093416 | MGI:5010479 | pseudogene | predicted gene, 18294 |
| 3 | gene | 64.23373 | 64.23546 | positive | MGI_C57BL6J_5593621 | Gm34462 | NCBI_Gene:102637724 | MGI:5593621 | lncRNA gene | predicted gene, 34462 |
| 3 | gene | 64.25722 | 64.28971 | negative | MGI_C57BL6J_3643995 | Vmn2r3 | NCBI_Gene:637004,ENSEMBL:ENSMUSG00000091572 | MGI:3643995 | protein coding gene | vomeronasal 2, receptor 3 |
| 3 | gene | 64.27582 | 64.27619 | negative | MGI_C57BL6J_1345615 | Ifld3 | NA | NA | unclassified gene | induced in fatty liver dystrophy 3 |
| 3 | gene | 64.31841 | 64.31856 | negative | MGI_C57BL6J_5453674 | Gm23897 | ENSEMBL:ENSMUSG00000088735 | MGI:5453674 | snoRNA gene | predicted gene, 23897 |
| 3 | pseudogene | 64.32866 | 64.33285 | positive | MGI_C57BL6J_3757669 | Vmn2r-ps4 | NCBI_Gene:100124547,ENSEMBL:ENSMUSG00000093503 | MGI:3757669 | pseudogene | vomeronasal 2, receptor, pseudogene 4 |
| 3 | pseudogene | 64.33956 | 64.35772 | negative | MGI_C57BL6J_3645404 | Vmn2r-ps5 | NCBI_Gene:667031,ENSEMBL:ENSMUSG00000091967 | MGI:3645404 | polymorphic pseudogene | vomeronasal 2, receptor, pseudogene 5 |
| 3 | gene | 64.38304 | 64.41537 | negative | MGI_C57BL6J_3648229 | Vmn2r4 | NCBI_Gene:637053,ENSEMBL:ENSMUSG00000092049 | MGI:3648229 | protein coding gene | vomeronasal 2, receptor 4 |
| 3 | pseudogene | 64.43688 | 64.43829 | negative | MGI_C57BL6J_3757678 | Vmn2r-ps6 | NCBI_Gene:100124548,ENSEMBL:ENSMUSG00000093395 | MGI:3757678 | pseudogene | vomeronasal 2, receptor, pseudogene 6 |
| 3 | pseudogene | 64.44260 | 64.44397 | negative | MGI_C57BL6J_3757679 | Vmn2r-ps7 | NCBI_Gene:672951,ENSEMBL:ENSMUSG00000093546 | MGI:3757679 | pseudogene | vomeronasal 2, receptor, pseudogene 7 |
| 3 | pseudogene | 64.44797 | 64.46009 | positive | MGI_C57BL6J_5313103 | Gm20656 | ENSEMBL:ENSMUSG00000093671 | MGI:5313103 | pseudogene | predicted gene 20656 |
| 3 | pseudogene | 64.47702 | 64.47932 | positive | MGI_C57BL6J_3757680 | Vmn2r-ps8 | NCBI_Gene:100124549,ENSEMBL:ENSMUSG00000094482 | MGI:3757680 | pseudogene | vomeronasal 2, receptor, pseudogene 8 |
| 3 | gene | 64.48861 | 64.50984 | negative | MGI_C57BL6J_3649074 | Vmn2r5 | NCBI_Gene:667060,ENSEMBL:ENSMUSG00000068999 | MGI:3649074 | protein coding gene | vomeronasal 2, receptor 5 |
| 3 | gene | 64.53131 | 64.56583 | negative | MGI_C57BL6J_3649068 | Vmn2r6 | NCBI_Gene:667069,ENSEMBL:ENSMUSG00000090581 | MGI:3649068 | protein coding gene | vomeronasal 2, receptor 6 |
| 3 | pseudogene | 64.59152 | 64.59290 | negative | MGI_C57BL6J_3757686 | Vmn2r-ps9 | NCBI_Gene:100124550,ENSEMBL:ENSMUSG00000093699 | MGI:3757686 | pseudogene | vomeronasal 2, receptor, pseudogene 9 |
| 3 | pseudogene | 64.59691 | 64.60577 | positive | MGI_C57BL6J_5313104 | Gm20657 | ENSEMBL:ENSMUSG00000093484 | MGI:5313104 | pseudogene | predicted gene 20657 |
| 3 | pseudogene | 64.60043 | 64.60114 | positive | MGI_C57BL6J_3781218 | Gm3040 | NCBI_Gene:100416170 | MGI:3781218 | pseudogene | predicted gene 3040 |
| 3 | pseudogene | 64.62169 | 64.62399 | positive | MGI_C57BL6J_3757943 | Vmn2r-ps10 | NCBI_Gene:100124554,ENSEMBL:ENSMUSG00000095107 | MGI:3757943 | pseudogene | vomeronasal 2, receptor, pseudogene 10 |
| 3 | pseudogene | 64.63257 | 64.84474 | negative | MGI_C57BL6J_2441682 | Vmn2r-ps11 | NCBI_Gene:319210,ENSEMBL:ENSMUSG00000093434 | MGI:2441682 | pseudogene | vomeronasal 2, receptor, pseudogene 11 |
| 3 | gene | 64.63286 | 64.84474 | negative | MGI_C57BL6J_5504054 | Gm26939 | ENSEMBL:ENSMUSG00000097538 | MGI:5504054 | lncRNA gene | predicted gene, 26939 |
| 3 | gene | 64.68504 | 64.84476 | negative | MGI_C57BL6J_2441693 | Vmn2r7 | NCBI_Gene:319217,ENSEMBL:ENSMUSG00000062200 | MGI:2441693 | protein coding gene | vomeronasal 2, receptor 7 |
| 3 | pseudogene | 64.74475 | 64.74564 | negative | MGI_C57BL6J_3757944 | Vmn2r-ps12 | NCBI_Gene:100125273,ENSEMBL:ENSMUSG00000093474 | MGI:3757944 | pseudogene | vomeronasal 2, receptor, pseudogene 12 |
| 3 | gene | 64.89447 | 64.89460 | negative | MGI_C57BL6J_5454999 | Gm25222 | ENSEMBL:ENSMUSG00000088824 | MGI:5454999 | snoRNA gene | predicted gene, 25222 |
| 3 | pseudogene | 64.89885 | 64.89976 | positive | MGI_C57BL6J_3757946 | Vmn2r-ps13 | NCBI_Gene:670110,ENSEMBL:ENSMUSG00000093767 | MGI:3757946 | pseudogene | vomeronasal 2, receptor, pseudogene 13 |
| 3 | gene | 64.94905 | 65.37823 | positive | MGI_C57BL6J_109155 | Kcnab1 | NCBI_Gene:16497,ENSEMBL:ENSMUSG00000027827 | MGI:109155 | protein coding gene | potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| 3 | gene | 64.94922 | 65.22370 | positive | MGI_C57BL6J_5611276 | Gm38048 | ENSEMBL:ENSMUSG00000102437 | MGI:5611276 | lncRNA gene | predicted gene, 38048 |
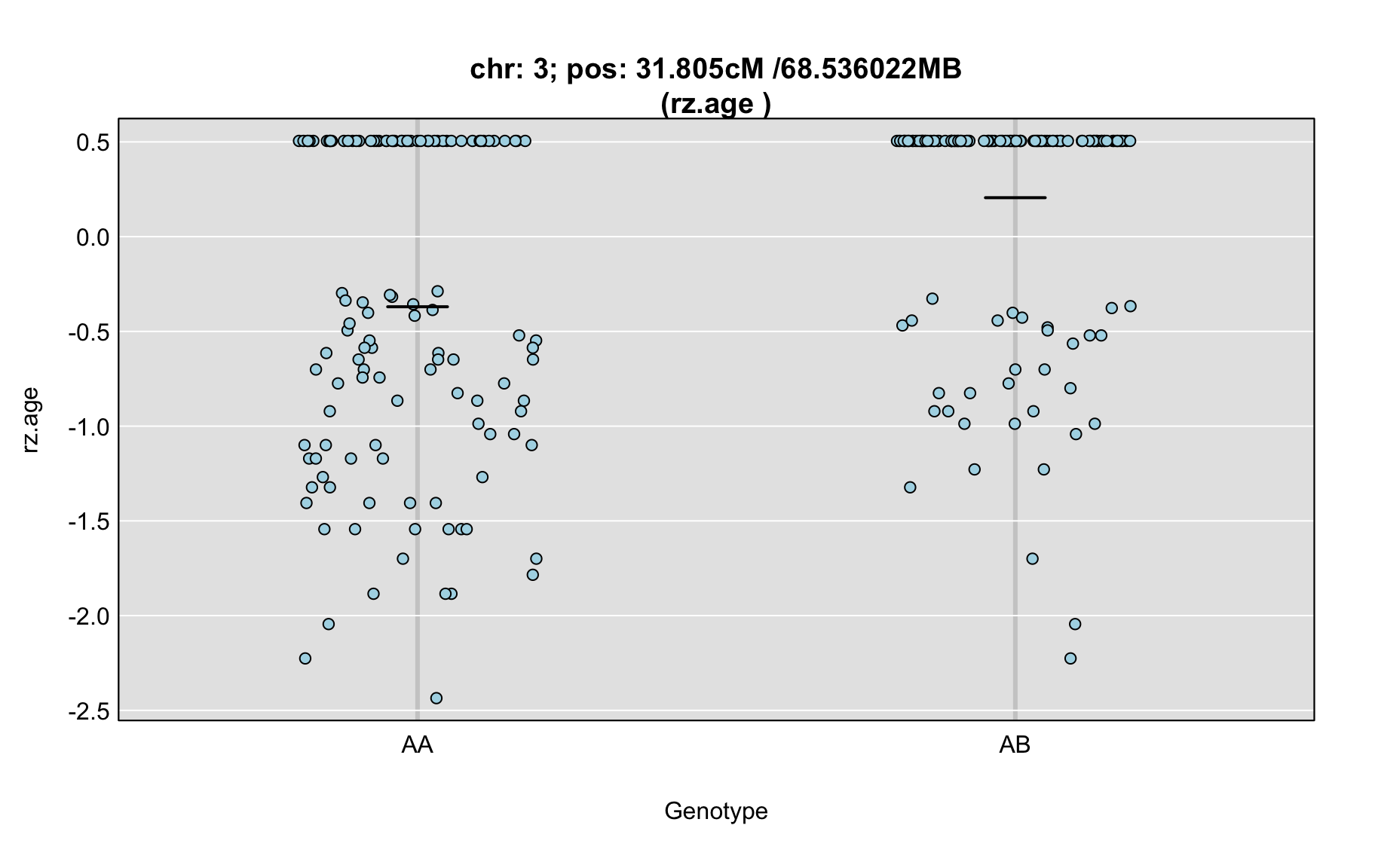
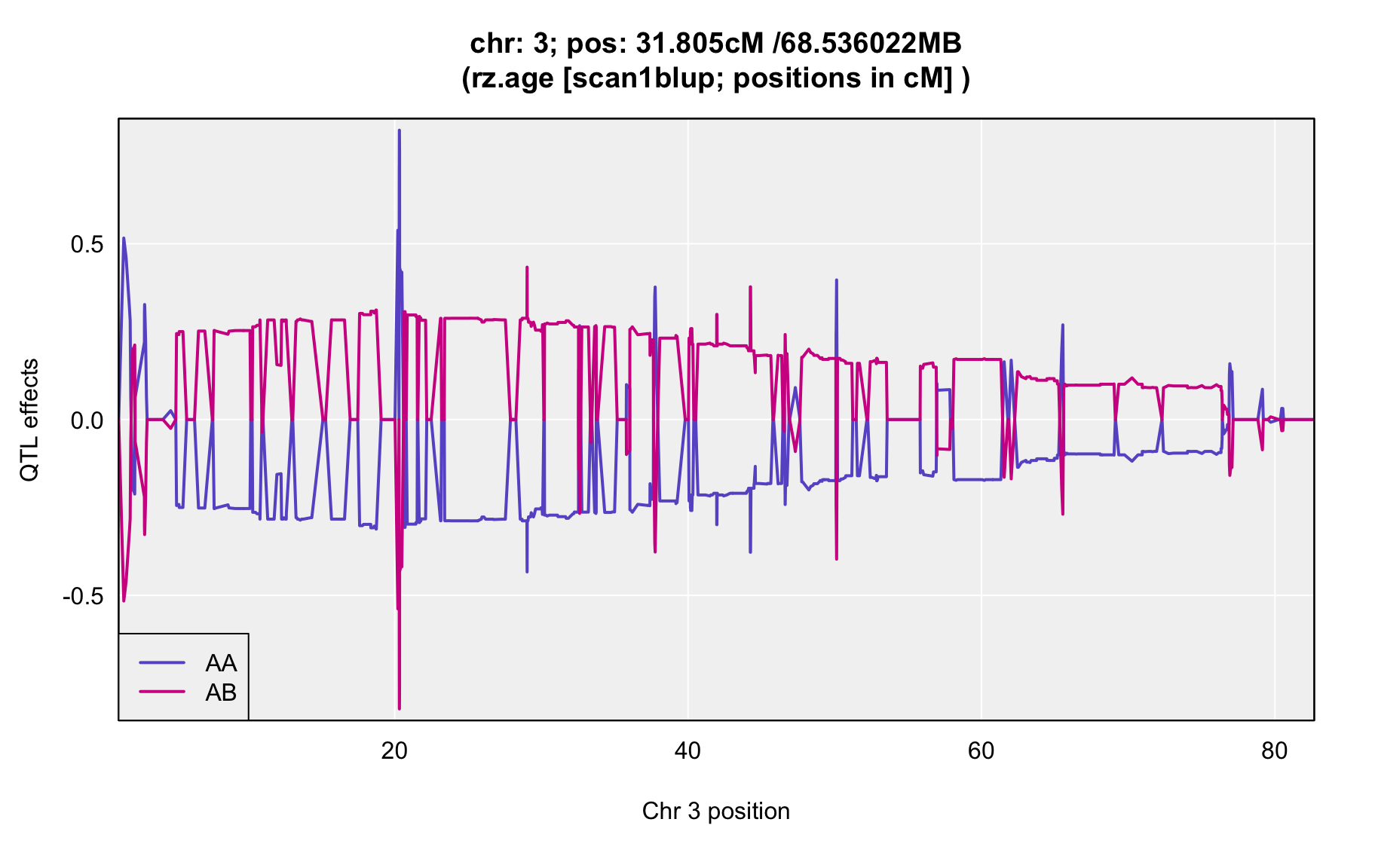
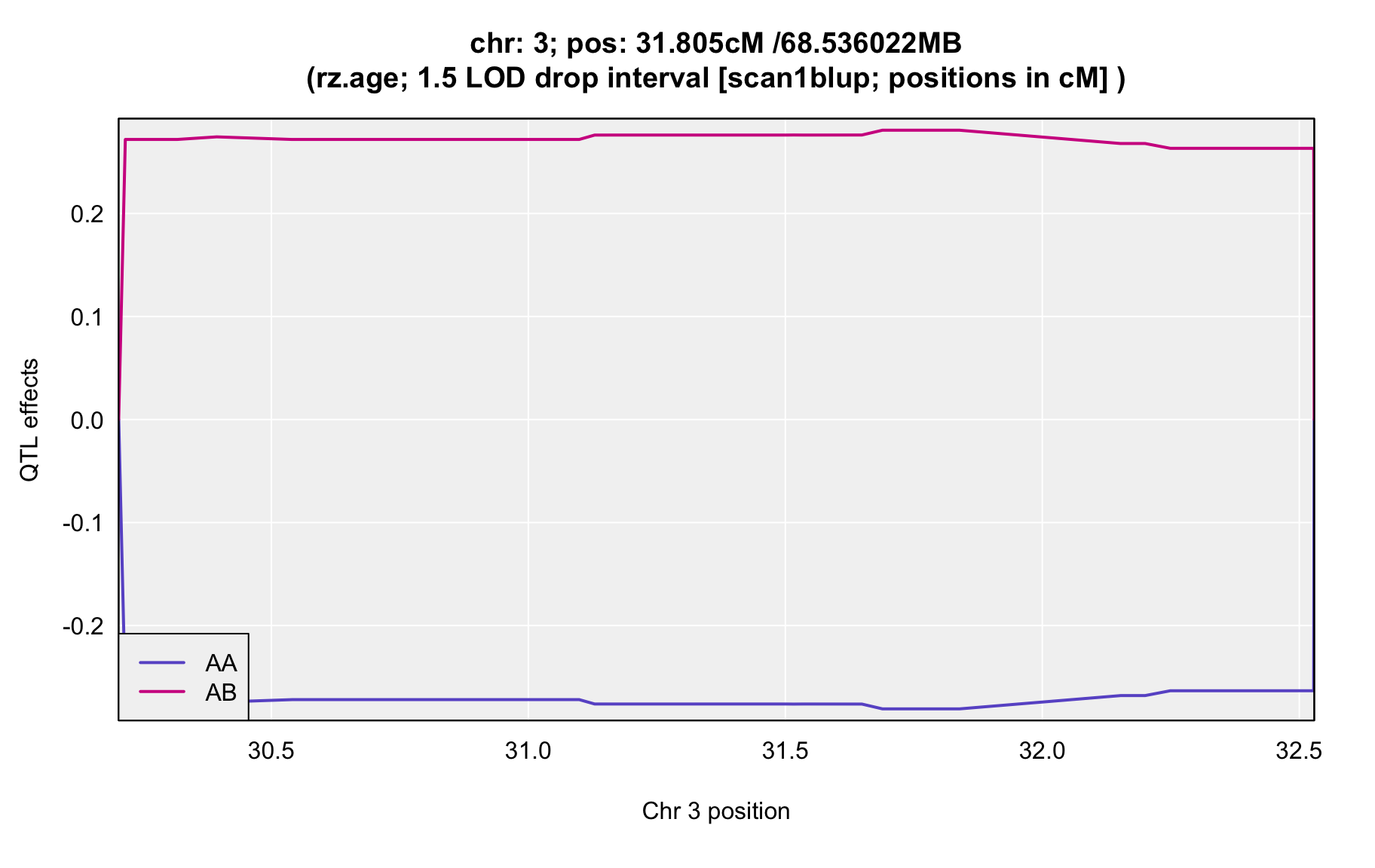
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 65.49409 | 65.49563 | negative | MGI_C57BL6J_5593939 | Gm34780 | NCBI_Gene:102638146,ENSEMBL:ENSMUSG00000103513 | MGI:5593939 | lncRNA gene | predicted gene, 34780 |
| 3 | gene | 65.49517 | 65.51729 | positive | MGI_C57BL6J_5622951 | Gm40066 | NCBI_Gene:105244461 | MGI:5622951 | lncRNA gene | predicted gene, 40066 |
| 3 | pseudogene | 65.50628 | 65.50664 | negative | MGI_C57BL6J_5610237 | Gm37009 | ENSEMBL:ENSMUSG00000103880 | MGI:5610237 | pseudogene | predicted gene, 37009 |
| 3 | gene | 65.50646 | 65.52004 | negative | MGI_C57BL6J_5621388 | Gm38503 | NCBI_Gene:102637953 | MGI:5621388 | lncRNA gene | predicted gene, 38503 |
| 3 | gene | 65.50726 | 65.52941 | negative | MGI_C57BL6J_1918254 | 4931440P22Rik | NCBI_Gene:71004,ENSEMBL:ENSMUSG00000074580 | MGI:1918254 | lncRNA gene | RIKEN cDNA 4931440P22 gene |
| 3 | gene | 65.52570 | 65.52579 | positive | MGI_C57BL6J_5452056 | Gm22279 | ENSEMBL:ENSMUSG00000087825 | MGI:5452056 | snRNA gene | predicted gene, 22279 |
| 3 | gene | 65.52841 | 65.55552 | positive | MGI_C57BL6J_2159210 | Tiparp | NCBI_Gene:99929,ENSEMBL:ENSMUSG00000034640 | MGI:2159210 | protein coding gene | TCDD-inducible poly(ADP-ribose) polymerase |
| 3 | gene | 65.57522 | 65.58085 | negative | MGI_C57BL6J_5594083 | Gm34924 | NCBI_Gene:102638334 | MGI:5594083 | lncRNA gene | predicted gene, 34924 |
| 3 | pseudogene | 65.61350 | 65.61532 | positive | MGI_C57BL6J_3648046 | Gm5847 | NCBI_Gene:545530,ENSEMBL:ENSMUSG00000082280 | MGI:3648046 | pseudogene | predicted gene 5847 |
| 3 | pseudogene | 65.61537 | 65.61617 | negative | MGI_C57BL6J_5611461 | Gm38233 | ENSEMBL:ENSMUSG00000103294 | MGI:5611461 | pseudogene | predicted gene, 38233 |
| 3 | gene | 65.65929 | 65.65943 | positive | MGI_C57BL6J_5531019 | Mir8120 | miRBase:MI0026052,NCBI_Gene:102465905,ENSEMBL:ENSMUSG00000098944 | MGI:5531019 | miRNA gene | microRNA 8120 |
| 3 | gene | 65.65954 | 65.83248 | positive | MGI_C57BL6J_3645902 | Lekr1 | NCBI_Gene:624866,ENSEMBL:ENSMUSG00000074579 | MGI:3645902 | protein coding gene | leucine, glutamate and lysine rich 1 |
| 3 | gene | 65.67839 | 65.68088 | positive | MGI_C57BL6J_5611004 | Gm37776 | ENSEMBL:ENSMUSG00000104037 | MGI:5611004 | unclassified gene | predicted gene, 37776 |
| 3 | gene | 65.77151 | 65.77162 | negative | MGI_C57BL6J_5455532 | Gm25755 | ENSEMBL:ENSMUSG00000065330 | MGI:5455532 | snRNA gene | predicted gene, 25755 |
| 3 | gene | 65.82912 | 65.87228 | negative | MGI_C57BL6J_5610587 | Gm37359 | ENSEMBL:ENSMUSG00000103620 | MGI:5610587 | lncRNA gene | predicted gene, 37359 |
| 3 | gene | 65.86327 | 65.86483 | negative | MGI_C57BL6J_5610359 | Gm37131 | ENSEMBL:ENSMUSG00000103440 | MGI:5610359 | unclassified gene | predicted gene, 37131 |
| 3 | gene | 65.92610 | 65.92828 | negative | MGI_C57BL6J_5594198 | Gm35039 | NCBI_Gene:102638487 | MGI:5594198 | lncRNA gene | predicted gene, 35039 |
| 3 | pseudogene | 65.93349 | 65.93391 | negative | MGI_C57BL6J_5610265 | Gm37037 | ENSEMBL:ENSMUSG00000102566 | MGI:5610265 | pseudogene | predicted gene, 37037 |
| 3 | gene | 65.94591 | 65.95839 | negative | MGI_C57BL6J_1922664 | Ccnl1 | NCBI_Gene:56706,ENSEMBL:ENSMUSG00000027829 | MGI:1922664 | protein coding gene | cyclin L1 |
| 3 | gene | 65.95776 | 65.96204 | positive | MGI_C57BL6J_5610533 | Gm37305 | ENSEMBL:ENSMUSG00000103041 | MGI:5610533 | lncRNA gene | predicted gene, 37305 |
| 3 | pseudogene | 65.96873 | 65.97028 | negative | MGI_C57BL6J_5594265 | Gm35106 | NCBI_Gene:105244463,ENSEMBL:ENSMUSG00000102914 | MGI:5594265 | pseudogene | predicted gene, 35106 |
| 3 | gene | 65.96991 | 65.97192 | positive | MGI_C57BL6J_5610306 | Gm37078 | ENSEMBL:ENSMUSG00000102652 | MGI:5610306 | unclassified gene | predicted gene, 37078 |
| 3 | pseudogene | 65.98716 | 65.98975 | negative | MGI_C57BL6J_5011209 | Gm19024 | NCBI_Gene:100418133,ENSEMBL:ENSMUSG00000103641 | MGI:5011209 | pseudogene | predicted gene, 19024 |
| 3 | pseudogene | 66.02950 | 66.03152 | negative | MGI_C57BL6J_3643872 | Gm6546 | NCBI_Gene:625048,ENSEMBL:ENSMUSG00000103369 | MGI:3643872 | pseudogene | predicted gene 6546 |
| 3 | gene | 66.05356 | 66.29707 | negative | MGI_C57BL6J_1920039 | Veph1 | NCBI_Gene:72789,ENSEMBL:ENSMUSG00000027831 | MGI:1920039 | protein coding gene | ventricular zone expressed PH domain-containing 1 |
| 3 | gene | 66.08668 | 66.08779 | positive | MGI_C57BL6J_5611050 | Gm37822 | ENSEMBL:ENSMUSG00000104189 | MGI:5611050 | unclassified gene | predicted gene, 37822 |
| 3 | gene | 66.08811 | 66.08826 | negative | MGI_C57BL6J_5456219 | Gm26442 | ENSEMBL:ENSMUSG00000064522 | MGI:5456219 | snRNA gene | predicted gene, 26442 |
| 3 | pseudogene | 66.09981 | 66.10006 | positive | MGI_C57BL6J_5610201 | Gm36973 | ENSEMBL:ENSMUSG00000104307 | MGI:5610201 | pseudogene | predicted gene, 36973 |
| 3 | pseudogene | 66.12840 | 66.12975 | negative | MGI_C57BL6J_98800 | Tpi-rs2 | NCBI_Gene:111205,ENSEMBL:ENSMUSG00000097968 | MGI:98800 | pseudogene | triosephosphate isomerase related sequence 2 |
| 3 | pseudogene | 66.19109 | 66.19124 | negative | MGI_C57BL6J_5610558 | Gm37330 | ENSEMBL:ENSMUSG00000103208 | MGI:5610558 | pseudogene | predicted gene, 37330 |
| 3 | gene | 66.21989 | 66.22581 | positive | MGI_C57BL6J_104641 | Ptx3 | NCBI_Gene:19288,ENSEMBL:ENSMUSG00000027832 | MGI:104641 | protein coding gene | pentraxin related gene |
| 3 | pseudogene | 66.36454 | 66.36463 | positive | MGI_C57BL6J_6275305 | Gm50014 | ENSEMBL:ENSMUSG00000117623 | MGI:6275305 | pseudogene | predicted gene, 50014 |
| 3 | gene | 66.37430 | 66.38810 | positive | MGI_C57BL6J_5594343 | Gm35184 | NCBI_Gene:102638679 | MGI:5594343 | lncRNA gene | predicted gene, 35184 |
| 3 | gene | 66.40041 | 66.40607 | positive | MGI_C57BL6J_5594395 | Gm35236 | NCBI_Gene:102638746 | MGI:5594395 | lncRNA gene | predicted gene, 35236 |
| 3 | pseudogene | 66.56278 | 66.56355 | negative | MGI_C57BL6J_5010137 | Gm17952 | NCBI_Gene:100416166,ENSEMBL:ENSMUSG00000103600 | MGI:5010137 | pseudogene | predicted gene, 17952 |
| 3 | pseudogene | 66.88235 | 66.88314 | positive | MGI_C57BL6J_3646021 | Gm6555 | NCBI_Gene:625115,ENSEMBL:ENSMUSG00000103700 | MGI:3646021 | pseudogene | predicted gene 6555 |
| 3 | pseudogene | 66.90736 | 66.90777 | negative | MGI_C57BL6J_5611584 | Gm38356 | ENSEMBL:ENSMUSG00000103318 | MGI:5611584 | pseudogene | predicted gene, 38356 |
| 3 | pseudogene | 66.95354 | 66.95386 | positive | MGI_C57BL6J_5663653 | Gm43516 | ENSEMBL:ENSMUSG00000105794 | MGI:5663653 | pseudogene | predicted gene 43516 |
| 3 | gene | 66.97172 | 66.98177 | negative | MGI_C57BL6J_1201673 | Shox2 | NCBI_Gene:20429,ENSEMBL:ENSMUSG00000027833 | MGI:1201673 | protein coding gene | short stature homeobox 2 |
| 3 | gene | 66.98139 | 67.35840 | positive | MGI_C57BL6J_1914130 | Rsrc1 | NCBI_Gene:66880,ENSEMBL:ENSMUSG00000034544 | MGI:1914130 | protein coding gene | arginine/serine-rich coiled-coil 1 |
| 3 | gene | 67.05596 | 67.05606 | positive | MGI_C57BL6J_5452072 | Gm22295 | ENSEMBL:ENSMUSG00000065245 | MGI:5452072 | snRNA gene | predicted gene, 22295 |
| 3 | gene | 67.36546 | 67.37516 | negative | MGI_C57BL6J_4937036 | Gm17402 | ENSEMBL:ENSMUSG00000090408 | MGI:4937036 | protein coding gene | predicted gene, 17402 |
| 3 | gene | 67.37410 | 67.40000 | positive | MGI_C57BL6J_1341819 | Mlf1 | NCBI_Gene:17349,ENSEMBL:ENSMUSG00000048416 | MGI:1341819 | protein coding gene | myeloid leukemia factor 1 |
| 3 | gene | 67.43010 | 67.47653 | positive | MGI_C57BL6J_107339 | Gfm1 | NCBI_Gene:28030,ENSEMBL:ENSMUSG00000027774 | MGI:107339 | protein coding gene | G elongation factor, mitochondrial 1 |
| 3 | gene | 67.45800 | 67.46393 | negative | MGI_C57BL6J_107633 | Lxn | NCBI_Gene:17035,ENSEMBL:ENSMUSG00000047557 | MGI:107633 | protein coding gene | latexin |
| 3 | gene | 67.47698 | 67.47882 | positive | MGI_C57BL6J_5662705 | Gm42568 | ENSEMBL:ENSMUSG00000103703 | MGI:5662705 | unclassified gene | predicted gene 42568 |
| 3 | gene | 67.47889 | 67.51552 | negative | MGI_C57BL6J_1924461 | Rarres1 | NCBI_Gene:109222,ENSEMBL:ENSMUSG00000049404 | MGI:1924461 | protein coding gene | retinoic acid receptor responder (tazarotene induced) 1 |
| 3 | gene | 67.51567 | 67.51966 | positive | MGI_C57BL6J_5622952 | Gm40067 | NCBI_Gene:105244464 | MGI:5622952 | lncRNA gene | predicted gene, 40067 |
| 3 | gene | 67.52266 | 67.55343 | negative | MGI_C57BL6J_5594458 | Gm35299 | NCBI_Gene:102638825,ENSEMBL:ENSMUSG00000103718 | MGI:5594458 | lncRNA gene | predicted gene, 35299 |
| 3 | gene | 67.57057 | 67.57282 | negative | MGI_C57BL6J_1921179 | 4930402C01Rik | ENSEMBL:ENSMUSG00000104382 | MGI:1921179 | lncRNA gene | RIKEN cDNA 4930402C01 gene |
| 3 | gene | 67.57316 | 67.57329 | negative | MGI_C57BL6J_5455696 | Gm25919 | ENSEMBL:ENSMUSG00000077459 | MGI:5455696 | snoRNA gene | predicted gene, 25919 |
| 3 | gene | 67.58274 | 67.60424 | positive | MGI_C57BL6J_1914118 | Mfsd1 | NCBI_Gene:66868,ENSEMBL:ENSMUSG00000027775 | MGI:1914118 | protein coding gene | major facilitator superfamily domain containing 1 |
| 3 | pseudogene | 67.60575 | 67.60785 | negative | MGI_C57BL6J_3648243 | Gm8515 | NCBI_Gene:667207,ENSEMBL:ENSMUSG00000103206 | MGI:3648243 | pseudogene | predicted gene 8515 |
| 3 | pseudogene | 67.61978 | 67.62073 | negative | MGI_C57BL6J_2685258 | Gm412 | NCBI_Gene:208424,ENSEMBL:ENSMUSG00000103606 | MGI:2685258 | pseudogene | predicted gene 412 |
| 3 | pseudogene | 67.68436 | 67.68475 | negative | MGI_C57BL6J_5610829 | Gm37601 | ENSEMBL:ENSMUSG00000102746 | MGI:5610829 | pseudogene | predicted gene, 37601 |
| 3 | pseudogene | 67.74197 | 67.74330 | negative | MGI_C57BL6J_5010902 | Gm18717 | NCBI_Gene:100417618,ENSEMBL:ENSMUSG00000104477 | MGI:5010902 | pseudogene | predicted gene, 18717 |
| 3 | gene | 67.75360 | 67.75565 | positive | MGI_C57BL6J_5594566 | Gm35407 | NCBI_Gene:102638975 | MGI:5594566 | lncRNA gene | predicted gene, 35407 |
| 3 | gene | 67.75566 | 67.76628 | positive | MGI_C57BL6J_5622953 | Gm40068 | NCBI_Gene:105244465 | MGI:5622953 | lncRNA gene | predicted gene, 40068 |
| 3 | gene | 67.89222 | 68.05659 | positive | MGI_C57BL6J_3644166 | Iqcj | NCBI_Gene:208426,ENSEMBL:ENSMUSG00000051777 | MGI:3644166 | protein coding gene | IQ motif containing J |
| 3 | gene | 67.89222 | 68.62648 | positive | MGI_C57BL6J_5439400 | Iqschfp | NCBI_Gene:100505386,ENSEMBL:ENSMUSG00000102422 | MGI:5439400 | protein coding gene | Iqcj and Schip1 fusion protein |
| 3 | gene | 68.06480 | 68.62648 | positive | MGI_C57BL6J_1353557 | Schip1 | NCBI_Gene:30953,ENSEMBL:ENSMUSG00000027777 | MGI:1353557 | protein coding gene | schwannomin interacting protein 1 |
| 3 | pseudogene | 68.30377 | 68.31036 | negative | MGI_C57BL6J_3648047 | Gm5848 | NCBI_Gene:545531 | MGI:3648047 | pseudogene | predicted pseudogene 5848 |
| 3 | pseudogene | 68.33319 | 68.33378 | positive | MGI_C57BL6J_3642851 | Gm10292 | NCBI_Gene:631379,ENSEMBL:ENSMUSG00000102846 | MGI:3642851 | pseudogene | predicted gene 10292 |
| 3 | gene | 68.40778 | 68.41470 | positive | MGI_C57BL6J_3641783 | Gm10723 | ENSEMBL:ENSMUSG00000103345 | MGI:3641783 | unclassified gene | predicted gene 10723 |
| 3 | gene | 68.54752 | 68.54771 | negative | MGI_C57BL6J_5451850 | Gm22073 | ENSEMBL:ENSMUSG00000084617 | MGI:5451850 | snRNA gene | predicted gene, 22073 |
| 3 | gene | 68.69064 | 68.69855 | positive | MGI_C57BL6J_96539 | Il12a | NCBI_Gene:16159,ENSEMBL:ENSMUSG00000027776 | MGI:96539 | protein coding gene | interleukin 12a |
| 3 | pseudogene | 68.73314 | 68.73808 | positive | MGI_C57BL6J_3708676 | Gm10040 | ENSEMBL:ENSMUSG00000058360 | MGI:3708676 | pseudogene | predicted gene 10040 |
| 3 | gene | 68.75991 | 68.76143 | negative | MGI_C57BL6J_1922468 | 4930535E02Rik | ENSEMBL:ENSMUSG00000102677 | MGI:1922468 | lncRNA gene | RIKEN cDNA 4930535E02 gene |
| 3 | gene | 68.76136 | 68.76267 | positive | MGI_C57BL6J_5594743 | Gm35584 | NCBI_Gene:102639225,ENSEMBL:ENSMUSG00000102936 | MGI:5594743 | lncRNA gene | predicted gene, 35584 |
| 3 | pseudogene | 68.84608 | 68.84741 | positive | MGI_C57BL6J_3644317 | Gm7270 | NCBI_Gene:639530,ENSEMBL:ENSMUSG00000103218,ENSEMBL:ENSMUSG00000102853 | MGI:3644317 | pseudogene | predicted gene 7270 |
| 3 | gene | 68.86792 | 68.87027 | negative | MGI_C57BL6J_4937275 | Gm17641 | ENSEMBL:ENSMUSG00000091272 | MGI:4937275 | protein coding gene | predicted gene, 17641 |
| 3 | gene | 68.86959 | 68.87216 | positive | MGI_C57BL6J_1915975 | 1110032F04Rik | NCBI_Gene:68725,ENSEMBL:ENSMUSG00000046999 | MGI:1915975 | protein coding gene | RIKEN cDNA 1110032F04 gene |
| 3 | gene | 68.89250 | 69.00464 | negative | MGI_C57BL6J_1915509 | Ift80 | NCBI_Gene:68259,ENSEMBL:ENSMUSG00000027778 | MGI:1915509 | protein coding gene | intraflagellar transport 80 |
| 3 | pseudogene | 68.90017 | 68.90045 | negative | MGI_C57BL6J_5313136 | Gm20689 | ENSEMBL:ENSMUSG00000093384 | MGI:5313136 | pseudogene | predicted gene 20689 |
| 3 | pseudogene | 68.90181 | 68.90216 | positive | MGI_C57BL6J_5313137 | Gm20690 | ENSEMBL:ENSMUSG00000093730,ENSEMBL:ENSMUSG00000103007 | MGI:5313137 | pseudogene | predicted gene 20690 |
| 3 | pseudogene | 68.90216 | 68.90278 | negative | MGI_C57BL6J_5010773 | Gm18588 | NCBI_Gene:100417394,ENSEMBL:ENSMUSG00000093402 | MGI:5010773 | pseudogene | predicted gene, 18588 |
| 3 | pseudogene | 68.90382 | 68.90415 | negative | MGI_C57BL6J_5313138 | Gm20691 | ENSEMBL:ENSMUSG00000093505 | MGI:5313138 | pseudogene | predicted gene 20691 |
| 3 | gene | 68.91650 | 68.91732 | negative | MGI_C57BL6J_6097561 | Gm48183 | ENSEMBL:ENSMUSG00000111756 | MGI:6097561 | unclassified gene | predicted gene, 48183 |
| 3 | gene | 69.00474 | 69.03462 | positive | MGI_C57BL6J_1917349 | Smc4 | NCBI_Gene:70099,ENSEMBL:ENSMUSG00000034349 | MGI:1917349 | protein coding gene | structural maintenance of chromosomes 4 |
| 3 | gene | 69.00977 | 69.00983 | positive | MGI_C57BL6J_2676842 | Mir15b | miRBase:MI0000140,NCBI_Gene:387175,ENSEMBL:ENSMUSG00000065580 | MGI:2676842 | miRNA gene | microRNA 15b |
| 3 | gene | 69.00990 | 69.01000 | positive | MGI_C57BL6J_3618690 | Mir16-2 | miRBase:MI0000566,NCBI_Gene:723949,ENSEMBL:ENSMUSG00000065606 | MGI:3618690 | miRNA gene | microRNA 16-2 |
| 3 | gene | 69.03529 | 69.04640 | negative | MGI_C57BL6J_1914199 | Trim59 | NCBI_Gene:66949,ENSEMBL:ENSMUSG00000034317 | MGI:1914199 | protein coding gene | tripartite motif-containing 59 |
| 3 | gene | 69.06065 | 69.06525 | positive | MGI_C57BL6J_2442898 | 9630025H16Rik | NCBI_Gene:105244467 | MGI:2442898 | lncRNA gene | RIKEN cDNA 9630025H16 gene |
| 3 | gene | 69.06715 | 69.12711 | negative | MGI_C57BL6J_1100848 | Kpna4 | NCBI_Gene:16649,ENSEMBL:ENSMUSG00000027782 | MGI:1100848 | protein coding gene | karyopherin (importin) alpha 4 |
| 3 | gene | 69.08510 | 69.08545 | negative | MGI_C57BL6J_5451786 | Gm22009 | ENSEMBL:ENSMUSG00000089417 | MGI:5451786 | snoRNA gene | predicted gene, 22009 |
| 3 | gene | 69.11974 | 69.12272 | negative | MGI_C57BL6J_5610786 | Gm37558 | ENSEMBL:ENSMUSG00000103094 | MGI:5610786 | unclassified gene | predicted gene, 37558 |
| 3 | gene | 69.13123 | 69.15718 | positive | MGI_C57BL6J_2686493 | Gm1647 | NCBI_Gene:381452,ENSEMBL:ENSMUSG00000094763 | MGI:2686493 | lncRNA gene | predicted gene 1647 |
| 3 | gene | 69.22242 | 69.22362 | positive | MGI_C57BL6J_1918869 | Arl14 | NCBI_Gene:71619,ENSEMBL:ENSMUSG00000098207 | MGI:1918869 | protein coding gene | ADP-ribosylation factor-like 14 |
| 3 | gene | 69.25461 | 69.26383 | positive | MGI_C57BL6J_5622955 | Gm40070 | NCBI_Gene:105244468 | MGI:5622955 | lncRNA gene | predicted gene, 40070 |
| 3 | gene | 69.31686 | 69.56080 | positive | MGI_C57BL6J_2139740 | Ppm1l | NCBI_Gene:242083,ENSEMBL:ENSMUSG00000027784 | MGI:2139740 | protein coding gene | protein phosphatase 1 (formerly 2C)-like |
| 3 | gene | 69.31816 | 69.31929 | negative | MGI_C57BL6J_1920514 | 1700037N05Rik | NA | NA | unclassified gene | RIKEN cDNA 1700037N05 gene |
| 3 | pseudogene | 69.32001 | 69.32062 | negative | MGI_C57BL6J_4938041 | Gm17214 | NCBI_Gene:100416279,ENSEMBL:ENSMUSG00000091315 | MGI:4938041 | pseudogene | predicted gene 17214 |
| 3 | pseudogene | 69.33519 | 69.33579 | negative | MGI_C57BL6J_4938039 | Gm17212 | NCBI_Gene:100416280,ENSEMBL:ENSMUSG00000090941 | MGI:4938039 | pseudogene | predicted gene 17212 |
| 3 | gene | 69.37640 | 69.38370 | positive | MGI_C57BL6J_2443158 | A330078L11Rik | NA | NA | unclassified gene | RIKEN cDNA A330078L11 gene |
| 3 | pseudogene | 69.43101 | 69.43238 | negative | MGI_C57BL6J_4938040 | Gm17213 | NCBI_Gene:100417908,ENSEMBL:ENSMUSG00000090743 | MGI:4938040 | pseudogene | predicted gene 17213 |
| 3 | pseudogene | 69.56624 | 69.56747 | negative | MGI_C57BL6J_3644126 | Gm8565 | NCBI_Gene:667309,ENSEMBL:ENSMUSG00000098006 | MGI:3644126 | pseudogene | predicted gene 8565 |
| 3 | gene | 69.57392 | 69.59896 | negative | MGI_C57BL6J_1349405 | B3galnt1 | NCBI_Gene:26879,ENSEMBL:ENSMUSG00000043300 | MGI:1349405 | protein coding gene | UDP-GalNAc:betaGlcNAc beta 1,3-galactosaminyltransferase, polypeptide 1 |
| 3 | pseudogene | 69.66177 | 69.66216 | positive | MGI_C57BL6J_5610608 | Gm37380 | ENSEMBL:ENSMUSG00000102342 | MGI:5610608 | pseudogene | predicted gene, 37380 |
| 3 | gene | 69.68547 | 69.68558 | positive | MGI_C57BL6J_5455398 | Gm25621 | ENSEMBL:ENSMUSG00000087848 | MGI:5455398 | rRNA gene | predicted gene, 25621 |
| 3 | pseudogene | 69.71694 | 69.71744 | negative | MGI_C57BL6J_98039 | Rpl32-ps | NCBI_Gene:19952,ENSEMBL:ENSMUSG00000068969 | MGI:98039 | pseudogene | ribosomal protein L32, pseudogene |
| 3 | gene | 69.72199 | 69.75637 | positive | MGI_C57BL6J_2140103 | Nmd3 | NCBI_Gene:97112,ENSEMBL:ENSMUSG00000027787 | MGI:2140103 | protein coding gene | NMD3 ribosome export adaptor |
| 3 | gene | 69.75311 | 69.75637 | positive | MGI_C57BL6J_2442565 | 9530051E23Rik | NA | NA | unclassified gene | RIKEN cDNA 9530051E23 gene |
| 3 | pseudogene | 69.80846 | 69.80902 | negative | MGI_C57BL6J_5011240 | Gm19055 | NCBI_Gene:100418175 | MGI:5011240 | pseudogene | predicted gene, 19055 |
| 3 | gene | 69.81954 | 69.85994 | negative | MGI_C57BL6J_1913433 | Sptssb | NCBI_Gene:66183,ENSEMBL:ENSMUSG00000043461 | MGI:1913433 | protein coding gene | serine palmitoyltransferase, small subunit B |
| 3 | gene | 69.88247 | 69.88411 | positive | MGI_C57BL6J_5594976 | Gm35817 | NCBI_Gene:102639522 | MGI:5594976 | lncRNA gene | predicted gene, 35817 |
| 3 | pseudogene | 69.95481 | 69.95568 | negative | MGI_C57BL6J_5010080 | Gm17895 | NCBI_Gene:100416058,ENSEMBL:ENSMUSG00000103549 | MGI:5010080 | pseudogene | predicted gene, 17895 |
| 3 | gene | 69.96232 | 69.96245 | negative | MGI_C57BL6J_5453261 | Gm23484 | ENSEMBL:ENSMUSG00000077366 | MGI:5453261 | snoRNA gene | predicted gene, 23484 |
| 3 | pseudogene | 69.96328 | 69.96347 | positive | MGI_C57BL6J_5611012 | Gm37784 | ENSEMBL:ENSMUSG00000102525 | MGI:5611012 | pseudogene | predicted gene, 37784 |
| 3 | gene | 70.00761 | 70.02871 | positive | MGI_C57BL6J_2685260 | Otol1 | NCBI_Gene:229389,ENSEMBL:ENSMUSG00000027788 | MGI:2685260 | protein coding gene | otolin 1 |
| 3 | pseudogene | 70.05975 | 70.06047 | positive | MGI_C57BL6J_5610909 | Gm37681 | ENSEMBL:ENSMUSG00000103020 | MGI:5610909 | pseudogene | predicted gene, 37681 |
| 3 | pseudogene | 70.11625 | 70.11653 | negative | MGI_C57BL6J_5610813 | Gm37585 | ENSEMBL:ENSMUSG00000103113 | MGI:5610813 | pseudogene | predicted gene, 37585 |
| 3 | gene | 70.22875 | 70.22887 | negative | MGI_C57BL6J_5453254 | Gm23477 | ENSEMBL:ENSMUSG00000089507 | MGI:5453254 | snoRNA gene | predicted gene, 23477 |
| 3 | pseudogene | 70.41328 | 70.41388 | positive | MGI_C57BL6J_3644892 | Gm6631 | NCBI_Gene:625868,ENSEMBL:ENSMUSG00000102663 | MGI:3644892 | pseudogene | predicted gene 6631 |
| 3 | pseudogene | 70.56626 | 70.56730 | negative | MGI_C57BL6J_5610441 | Gm37213 | ENSEMBL:ENSMUSG00000104023 | MGI:5610441 | pseudogene | predicted gene, 37213 |
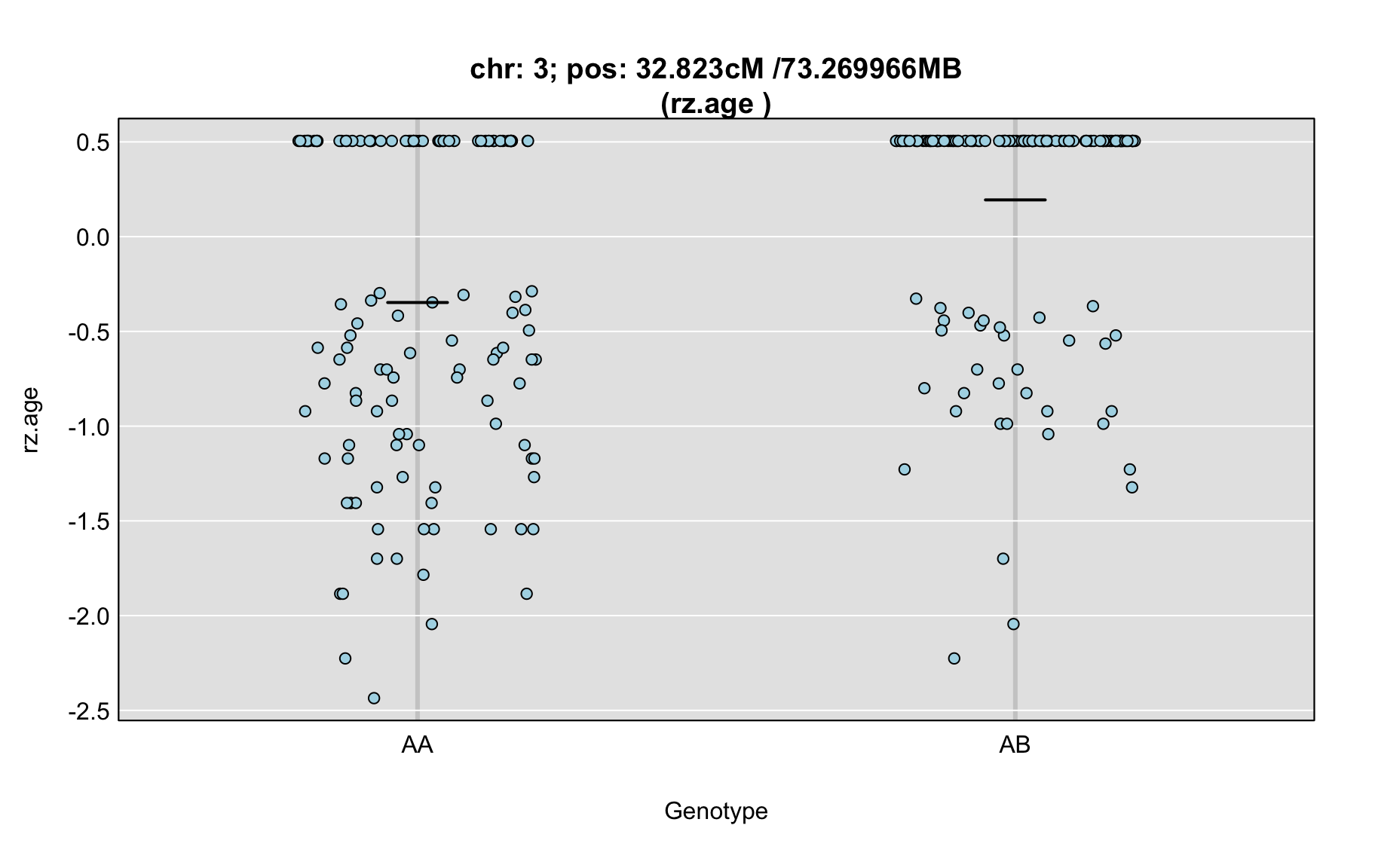
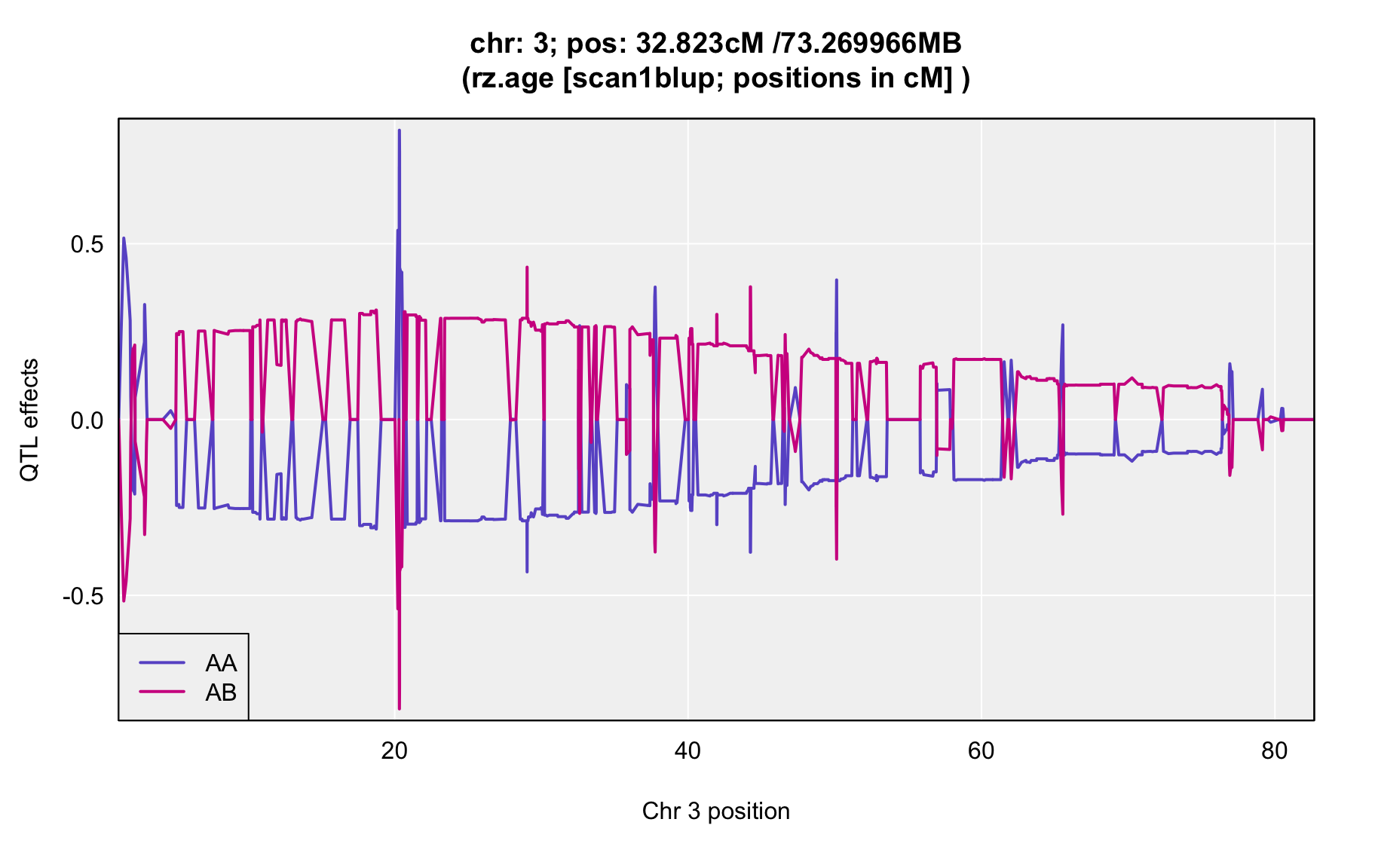
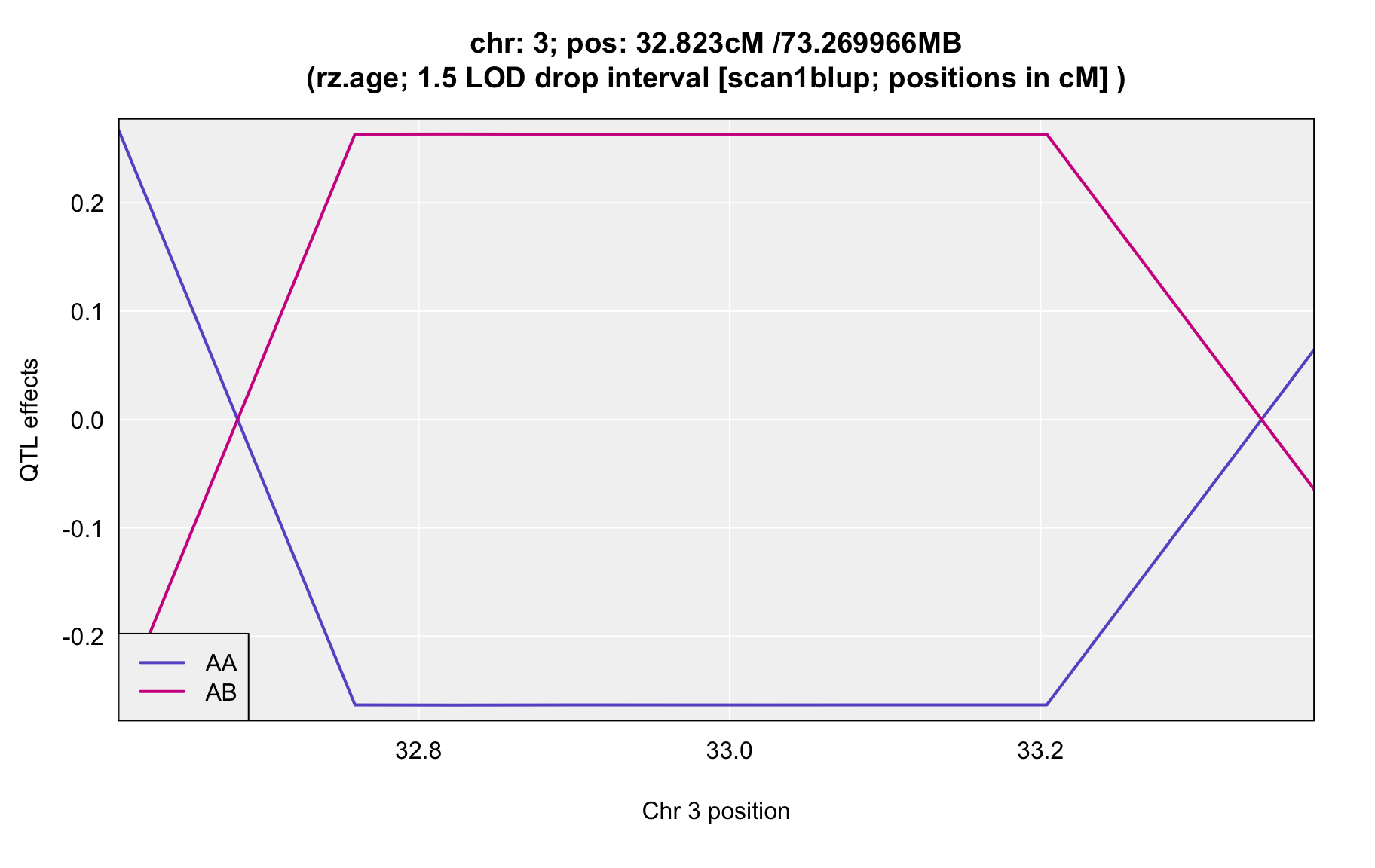
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | pseudogene | 72.60384 | 72.60435 | negative | MGI_C57BL6J_5611129 | Gm37901 | ENSEMBL:ENSMUSG00000103078 | MGI:5611129 | pseudogene | predicted gene, 37901 |
| 3 | gene | 72.73783 | 72.73798 | negative | MGI_C57BL6J_5452228 | Gm22451 | ENSEMBL:ENSMUSG00000084752 | MGI:5452228 | snRNA gene | predicted gene, 22451 |
| 3 | gene | 72.88855 | 72.96786 | negative | MGI_C57BL6J_1917233 | Sis | NCBI_Gene:69983,ENSEMBL:ENSMUSG00000027790 | MGI:1917233 | protein coding gene | sucrase isomaltase (alpha-glucosidase) |
| 3 | gene | 73.04197 | 73.04384 | negative | MGI_C57BL6J_5610284 | Gm37056 | ENSEMBL:ENSMUSG00000102753 | MGI:5610284 | unclassified gene | predicted gene, 37056 |
| 3 | gene | 73.04725 | 73.05788 | negative | MGI_C57BL6J_2679447 | Slitrk3 | NCBI_Gene:386750,ENSEMBL:ENSMUSG00000048304 | MGI:2679447 | protein coding gene | SLIT and NTRK-like family, member 3 |
| 3 | gene | 73.06632 | 73.59483 | positive | MGI_C57BL6J_5434110 | Gm20754 | NCBI_Gene:626082,ENSEMBL:ENSMUSG00000103428 | MGI:5434110 | lncRNA gene | predicted gene, 20754 |
| 3 | gene | 73.14685 | 73.14696 | positive | MGI_C57BL6J_5454321 | Gm24544 | ENSEMBL:ENSMUSG00000077050 | MGI:5454321 | miRNA gene | predicted gene, 24544 |
| 3 | gene | 73.16409 | 73.16621 | positive | MGI_C57BL6J_5610953 | Gm37725 | ENSEMBL:ENSMUSG00000102916 | MGI:5610953 | unclassified gene | predicted gene, 37725 |
| 3 | gene | 73.20770 | 73.20800 | negative | MGI_C57BL6J_5452315 | Gm22538 | ENSEMBL:ENSMUSG00000088794 | MGI:5452315 | unclassified non-coding RNA gene | predicted gene, 22538 |
| 3 | gene | 73.32943 | 73.47001 | positive | MGI_C57BL6J_5611581 | Gm38353 | ENSEMBL:ENSMUSG00000102820 | MGI:5611581 | lncRNA gene | predicted gene, 38353 |
| 3 | gene | 73.35121 | 73.48165 | negative | MGI_C57BL6J_1921943 | 4930509J09Rik | NCBI_Gene:74693,ENSEMBL:ENSMUSG00000102362 | MGI:1921943 | lncRNA gene | RIKEN cDNA 4930509J09 gene |
| 3 | gene | 73.63581 | 73.70844 | negative | MGI_C57BL6J_894278 | Bche | NCBI_Gene:12038,ENSEMBL:ENSMUSG00000027792 | MGI:894278 | protein coding gene | butyrylcholinesterase |
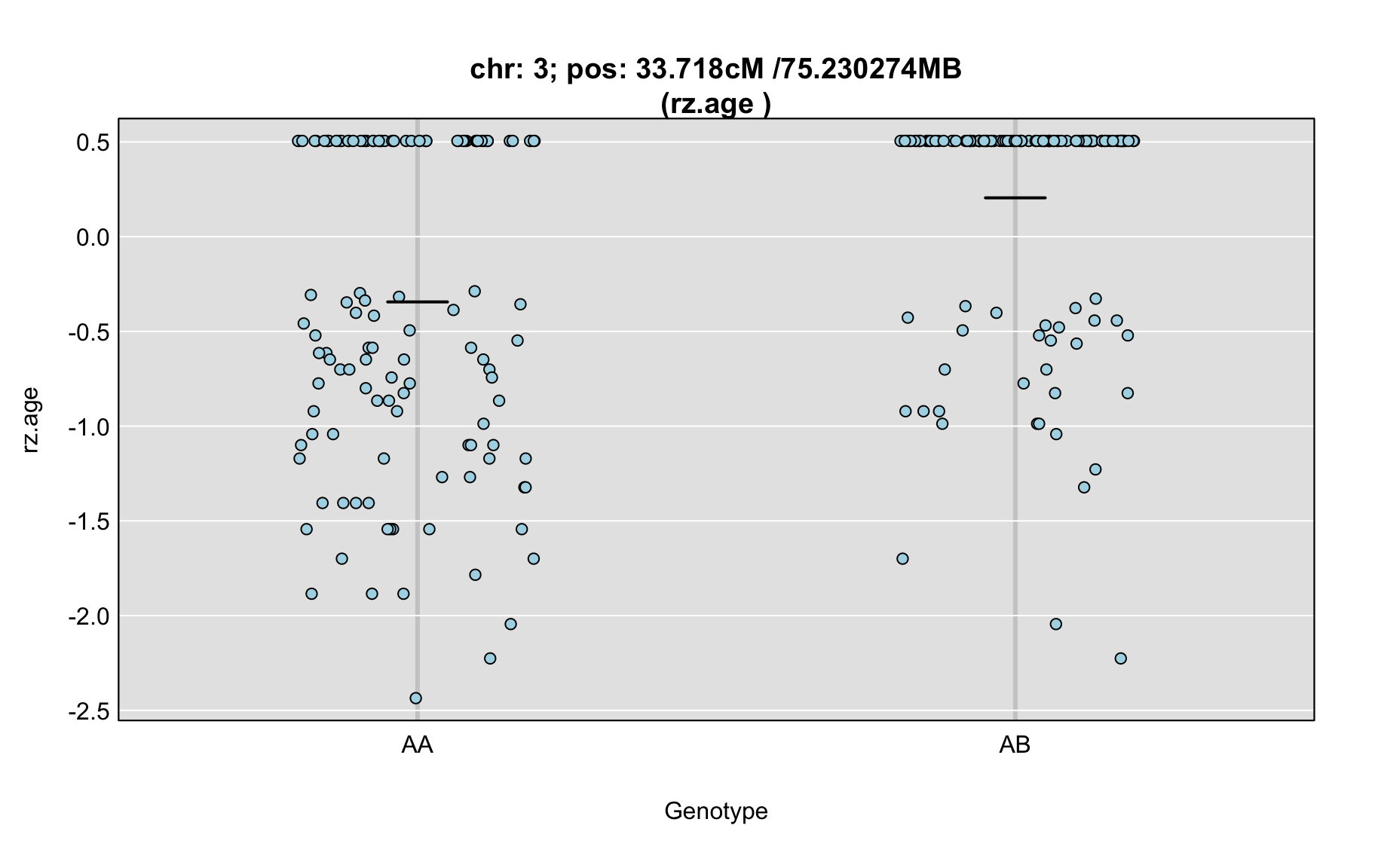
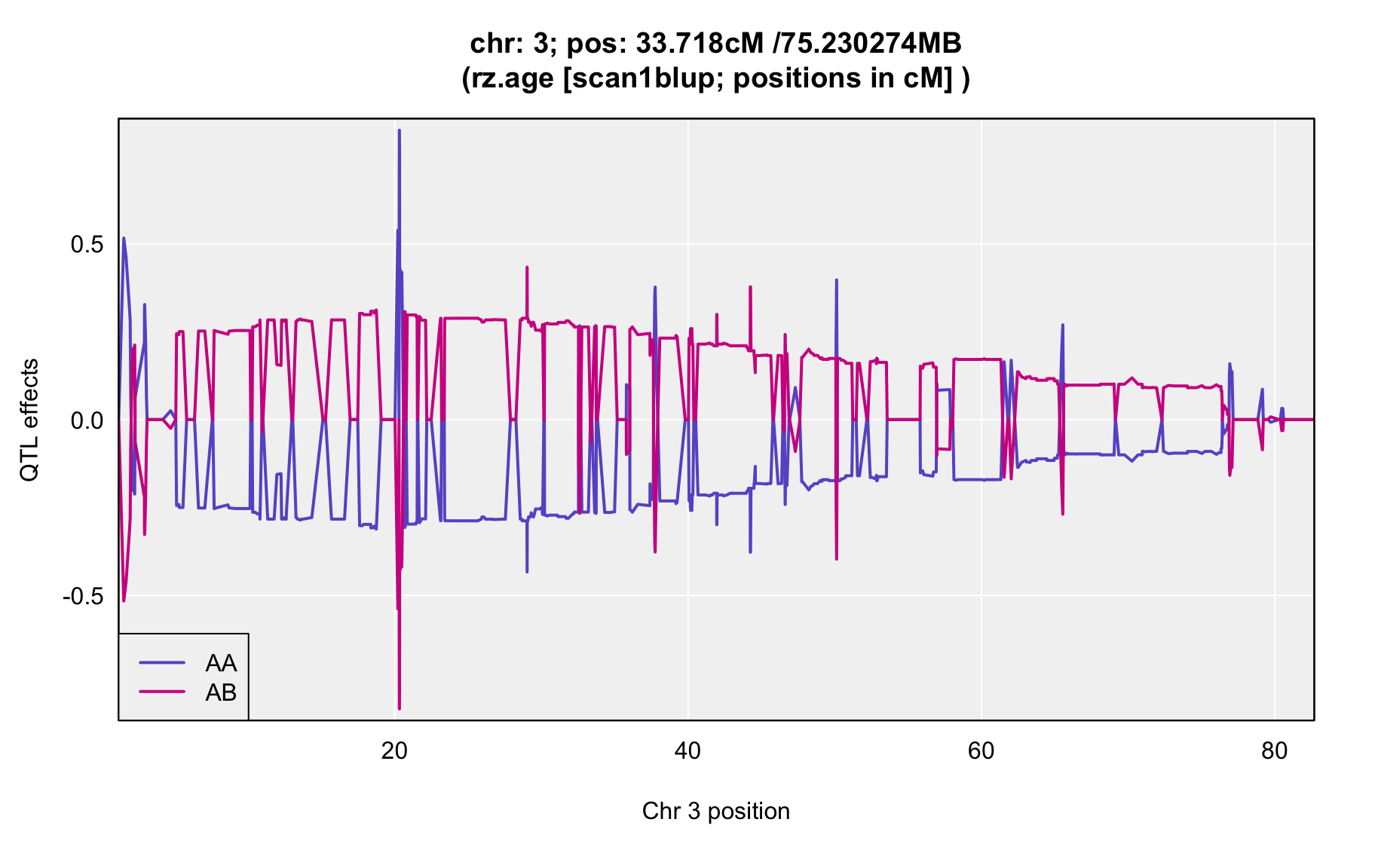
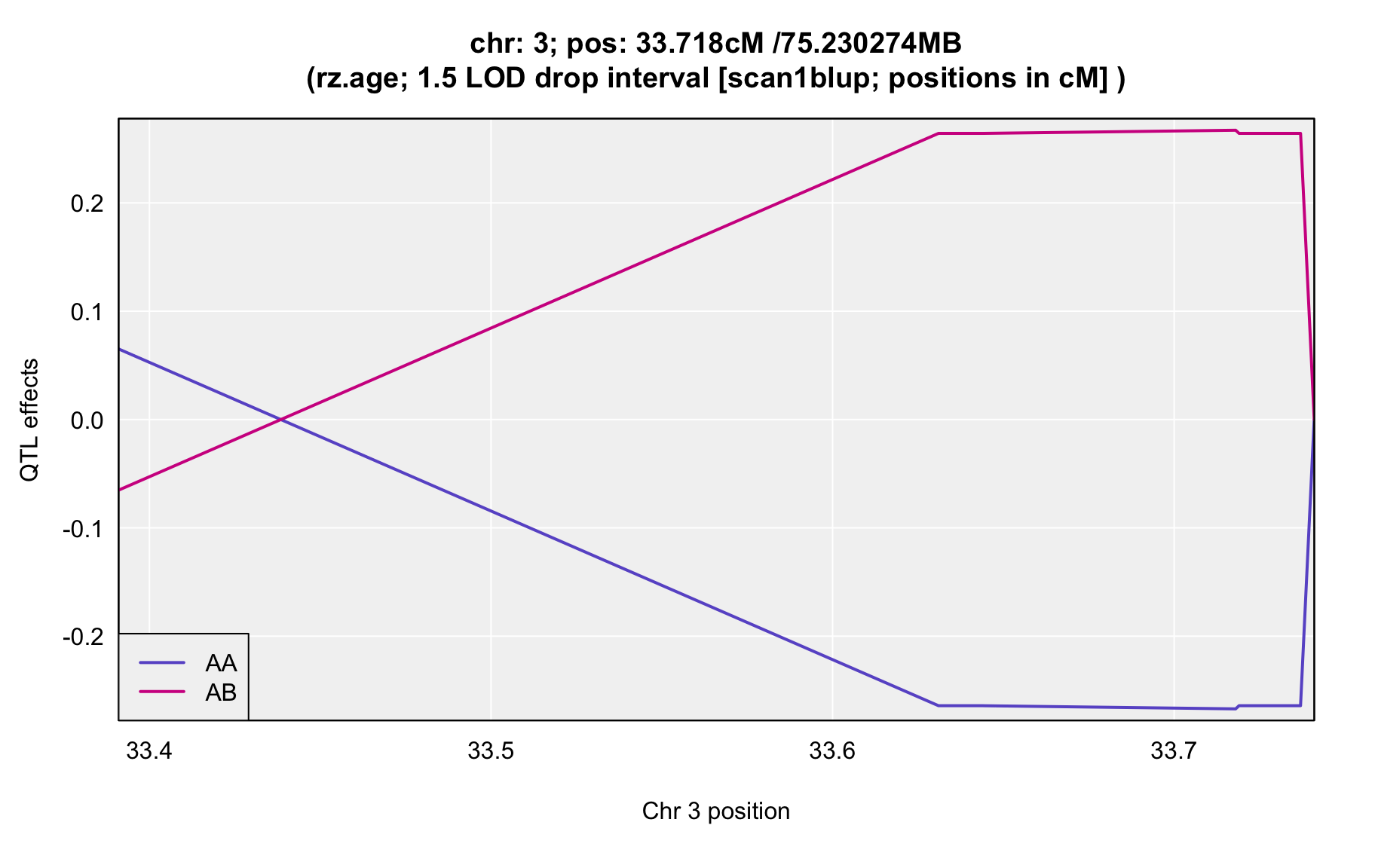
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | pseudogene | 73.93305 | 73.93412 | positive | MGI_C57BL6J_3644686 | Gm8625 | NCBI_Gene:667425,ENSEMBL:ENSMUSG00000102859 | MGI:3644686 | pseudogene | predicted gene 8625 |
| 3 | gene | 74.00922 | 74.01728 | negative | MGI_C57BL6J_5012541 | Gm20356 | NCBI_Gene:100504692,ENSEMBL:ENSMUSG00000104096 | MGI:5012541 | lncRNA gene | predicted gene, 20356 |
| 3 | gene | 74.10541 | 74.12645 | negative | MGI_C57BL6J_5595228 | Gm36069 | NCBI_Gene:102639855 | MGI:5595228 | lncRNA gene | predicted gene, 36069 |
| 3 | pseudogene | 74.24226 | 74.29687 | negative | MGI_C57BL6J_3779552 | Gm6098 | NCBI_Gene:100861784,ENSEMBL:ENSMUSG00000096400 | MGI:3779552 | pseudogene | predicted gene 6098 |
| 3 | pseudogene | 74.32353 | 74.32399 | negative | MGI_C57BL6J_5610278 | Gm37050 | ENSEMBL:ENSMUSG00000104227 | MGI:5610278 | pseudogene | predicted gene, 37050 |
| 3 | pseudogene | 74.81472 | 74.81534 | positive | MGI_C57BL6J_3647458 | Gm8643 | NCBI_Gene:108168917,ENSEMBL:ENSMUSG00000102809 | MGI:3647458 | pseudogene | predicted gene 8643 |
| 3 | gene | 75.02240 | 75.02461 | negative | MGI_C57BL6J_5610692 | Gm37464 | ENSEMBL:ENSMUSG00000102420 | MGI:5610692 | unclassified gene | predicted gene, 37464 |
| 3 | gene | 75.03790 | 75.16505 | negative | MGI_C57BL6J_2674085 | Zbbx | NCBI_Gene:213234,ENSEMBL:ENSMUSG00000034151 | MGI:2674085 | protein coding gene | zinc finger, B-box domain containing |
| 3 | gene | 75.16506 | 75.20808 | positive | MGI_C57BL6J_5622956 | Gm40071 | NCBI_Gene:105244469 | MGI:5622956 | lncRNA gene | predicted gene, 40071 |
| 3 | gene | 75.24236 | 75.27008 | negative | MGI_C57BL6J_1915181 | Serpini2 | NCBI_Gene:67931,ENSEMBL:ENSMUSG00000034139 | MGI:1915181 | protein coding gene | serine (or cysteine) peptidase inhibitor, clade I, member 2 |
| 3 | gene | 75.27393 | 75.48216 | negative | MGI_C57BL6J_3645287 | Wdr49 | NCBI_Gene:213248,ENSEMBL:ENSMUSG00000104301 | MGI:3645287 | protein coding gene | WD repeat domain 49 |
| 3 | pseudogene | 75.39132 | 75.39264 | positive | MGI_C57BL6J_5010021 | Gm17836 | NCBI_Gene:100415953,ENSEMBL:ENSMUSG00000103444 | MGI:5010021 | pseudogene | predicted gene, 17836 |
| 3 | gene | 75.51649 | 75.55686 | negative | MGI_C57BL6J_1928396 | Pdcd10 | NCBI_Gene:56426,ENSEMBL:ENSMUSG00000027835 | MGI:1928396 | protein coding gene | programmed cell death 10 |
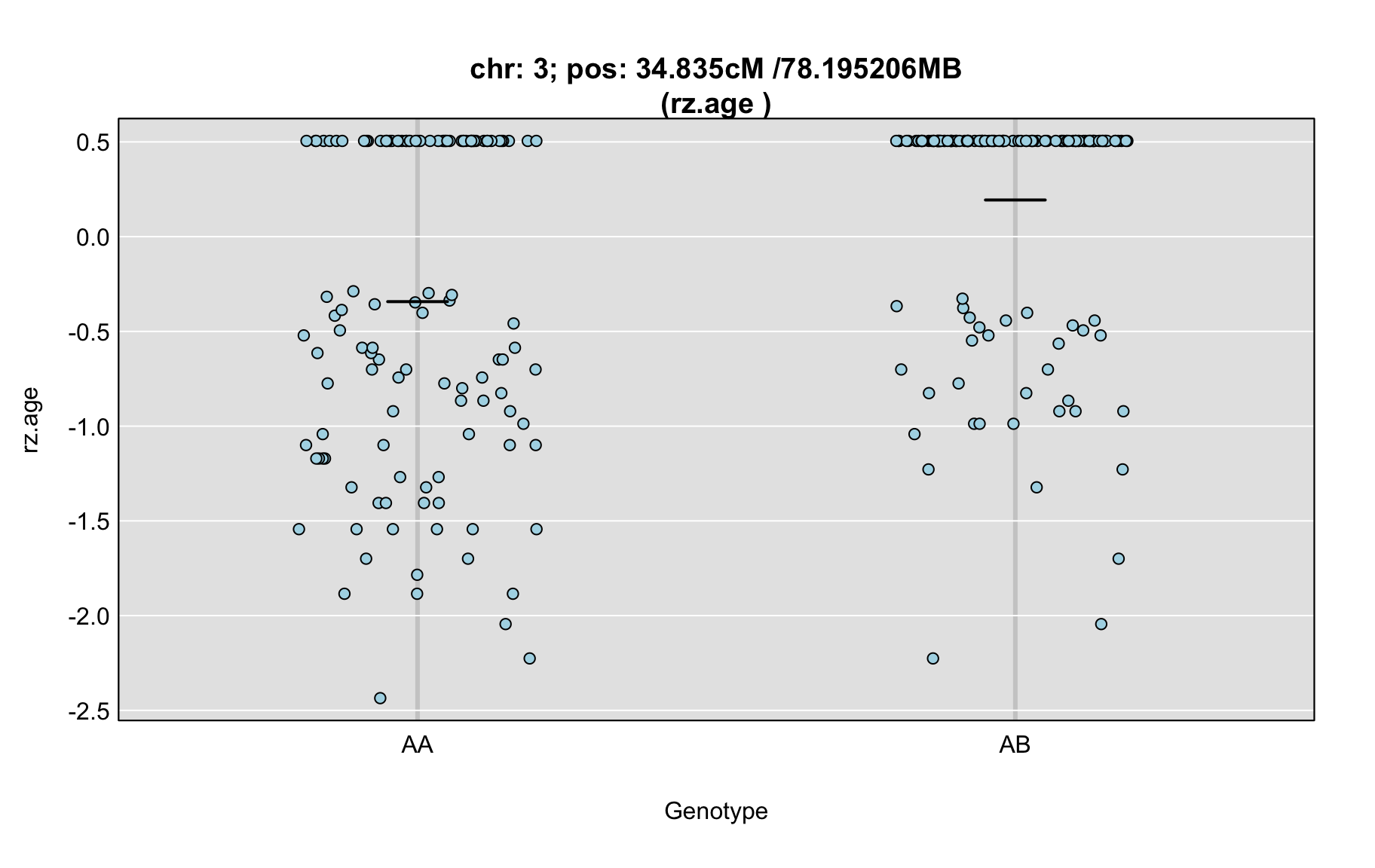
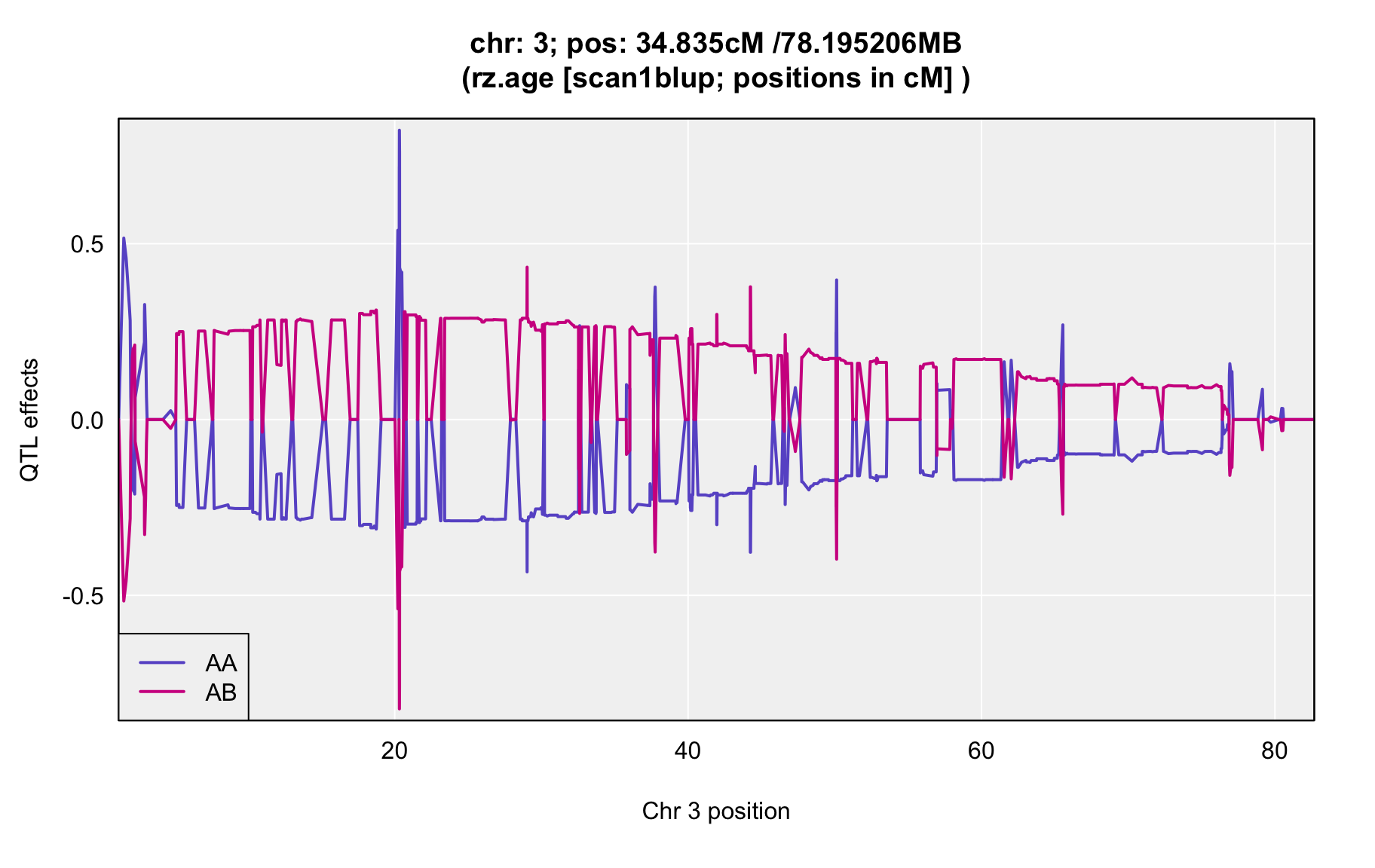
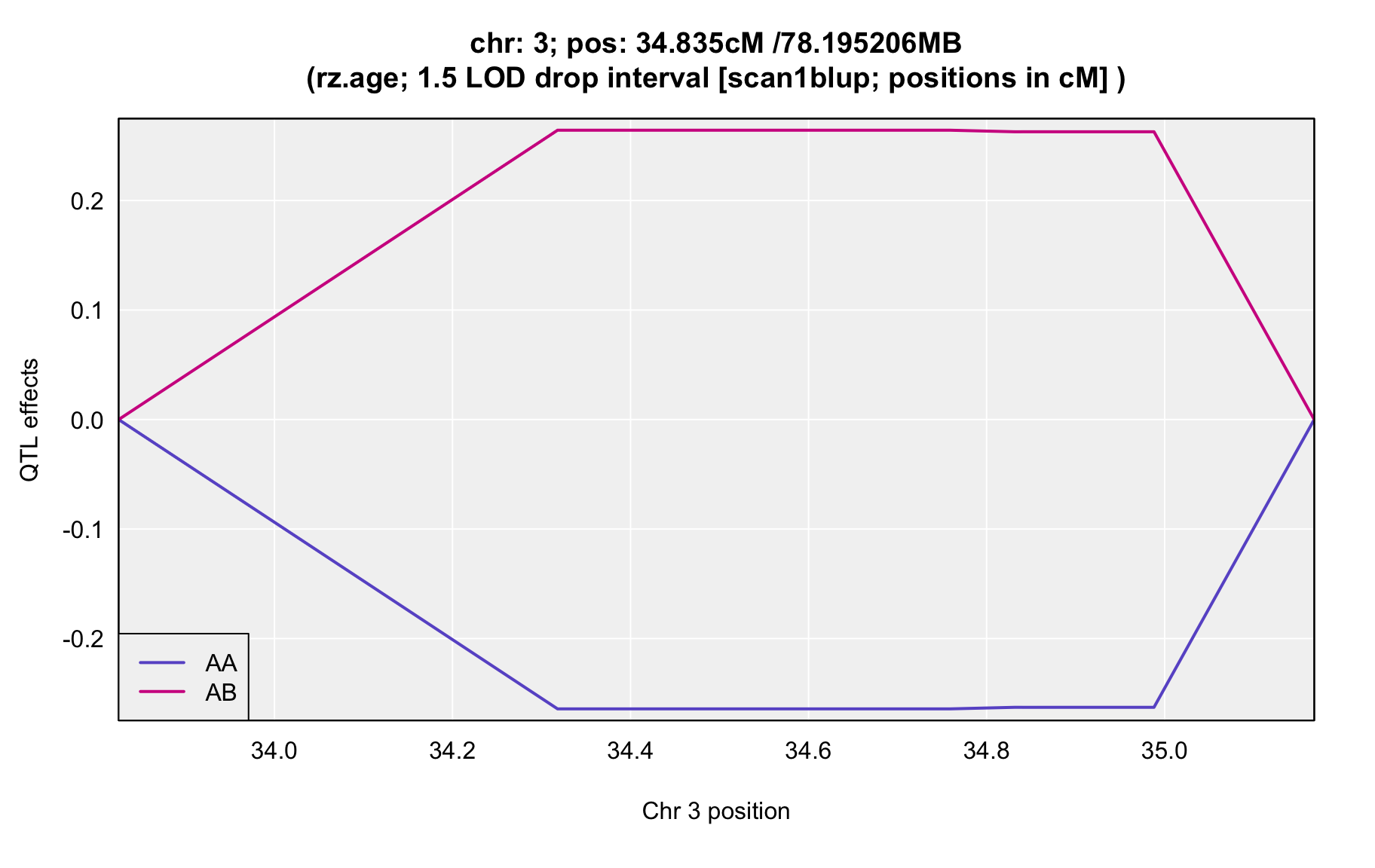
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 75.87508 | 75.95695 | negative | MGI_C57BL6J_1920374 | Golim4 | NCBI_Gene:73124,ENSEMBL:ENSMUSG00000034109 | MGI:1920374 | protein coding gene | golgi integral membrane protein 4 |
| 3 | gene | 75.95212 | 75.95430 | positive | MGI_C57BL6J_5610913 | Gm37685 | ENSEMBL:ENSMUSG00000103024 | MGI:5610913 | unclassified gene | predicted gene, 37685 |
| 3 | gene | 75.95610 | 76.00114 | positive | MGI_C57BL6J_5610484 | Gm37256 | ENSEMBL:ENSMUSG00000104448 | MGI:5610484 | lncRNA gene | predicted gene, 37256 |
| 3 | pseudogene | 76.04092 | 76.04172 | negative | MGI_C57BL6J_5010613 | Gm18428 | NCBI_Gene:100417155,ENSEMBL:ENSMUSG00000104223 | MGI:5010613 | pseudogene | predicted gene, 18428 |
| 3 | gene | 76.07427 | 76.71002 | positive | MGI_C57BL6J_2442179 | Fstl5 | NCBI_Gene:213262,ENSEMBL:ENSMUSG00000034098 | MGI:2442179 | protein coding gene | follistatin-like 5 |
| 3 | gene | 76.54669 | 76.55025 | positive | MGI_C57BL6J_5611125 | Gm37897 | ENSEMBL:ENSMUSG00000103920 | MGI:5611125 | unclassified gene | predicted gene, 37897 |
| 3 | pseudogene | 76.82206 | 76.82240 | positive | MGI_C57BL6J_5610518 | Gm37290 | ENSEMBL:ENSMUSG00000103702 | MGI:5610518 | pseudogene | predicted gene, 37290 |
| 3 | pseudogene | 76.88450 | 76.88533 | positive | MGI_C57BL6J_5011416 | Gm19231 | NCBI_Gene:100418471,ENSEMBL:ENSMUSG00000102216 | MGI:5011416 | pseudogene | predicted gene, 19231 |
| 3 | gene | 77.10521 | 77.10528 | negative | MGI_C57BL6J_5453421 | Gm23644 | ENSEMBL:ENSMUSG00000088481 | MGI:5453421 | snoRNA gene | predicted gene, 23644 |
| 3 | pseudogene | 77.56081 | 77.56394 | negative | MGI_C57BL6J_2674154 | Dnmt3a-ps1 | NCBI_Gene:445261,ENSEMBL:ENSMUSG00000102489 | MGI:2674154 | pseudogene | DNA methyltransferase 3A, pseudogene 1 |
| 3 | gene | 77.56496 | 77.57134 | negative | MGI_C57BL6J_5611034 | Gm37806 | ENSEMBL:ENSMUSG00000103849 | MGI:5611034 | unclassified gene | predicted gene, 37806 |
| 3 | pseudogene | 77.69137 | 77.69223 | negative | MGI_C57BL6J_5010480 | Gm18295 | NCBI_Gene:100416866 | MGI:5010480 | pseudogene | predicted gene, 18295 |
| 3 | gene | 77.70289 | 77.70439 | negative | MGI_C57BL6J_5611423 | Gm38195 | ENSEMBL:ENSMUSG00000102872 | MGI:5611423 | unclassified gene | predicted gene, 38195 |
| 3 | pseudogene | 77.87278 | 77.87315 | negative | MGI_C57BL6J_5610804 | Gm37576 | ENSEMBL:ENSMUSG00000102393 | MGI:5610804 | pseudogene | predicted gene, 37576 |
| 3 | pseudogene | 77.96138 | 77.96280 | positive | MGI_C57BL6J_5010904 | Gm18719 | NCBI_Gene:100417620,ENSEMBL:ENSMUSG00000105026 | MGI:5010904 | pseudogene | predicted gene, 18719 |
| 3 | pseudogene | 78.00646 | 78.00698 | positive | MGI_C57BL6J_3648953 | Gm6677 | NCBI_Gene:626461,ENSEMBL:ENSMUSG00000103312 | MGI:3648953 | pseudogene | predicted gene 6677 |
| 3 | pseudogene | 78.08583 | 78.08740 | negative | MGI_C57BL6J_3643677 | Gm8684 | NCBI_Gene:667526,ENSEMBL:ENSMUSG00000102637 | MGI:3643677 | pseudogene | predicted gene 8684 |
| 3 | pseudogene | 78.13446 | 78.13558 | positive | MGI_C57BL6J_3645908 | Gm6680 | NCBI_Gene:626479,ENSEMBL:ENSMUSG00000103817 | MGI:3645908 | pseudogene | predicted gene 6680 |
| 3 | gene | 78.40430 | 78.40594 | negative | MGI_C57BL6J_5663463 | Gm43326 | ENSEMBL:ENSMUSG00000104782 | MGI:5663463 | unclassified gene | predicted gene 43326 |
| 3 | gene | 78.40616 | 78.41077 | negative | MGI_C57BL6J_5595447 | Gm36288 | NCBI_Gene:102640153 | MGI:5595447 | lncRNA gene | predicted gene, 36288 |
| 3 | pseudogene | 78.48658 | 78.48868 | negative | MGI_C57BL6J_3705708 | Gm15442 | NCBI_Gene:100041691,ENSEMBL:ENSMUSG00000084370 | MGI:3705708 | pseudogene | predicted gene 15442 |
| 3 | pseudogene | 78.75862 | 78.75946 | positive | MGI_C57BL6J_5011137 | Gm18952 | NCBI_Gene:100418022,ENSEMBL:ENSMUSG00000102341 | MGI:5011137 | pseudogene | predicted gene, 18952 |
| 3 | gene | 78.78973 | 78.79223 | negative | MGI_C57BL6J_5595500 | Gm36341 | NCBI_Gene:102640221 | MGI:5595500 | lncRNA gene | predicted gene, 36341 |
| 3 | gene | 78.84222 | 78.84257 | positive | MGI_C57BL6J_5454526 | Gm24749 | ENSEMBL:ENSMUSG00000088463 | MGI:5454526 | unclassified non-coding RNA gene | predicted gene, 24749 |
| 3 | pseudogene | 78.86489 | 78.86557 | negative | MGI_C57BL6J_5010138 | Gm17953 | NCBI_Gene:100416167,ENSEMBL:ENSMUSG00000102438 | MGI:5010138 | pseudogene | predicted gene, 17953 |
| 3 | pseudogene | 78.89183 | 78.89247 | negative | MGI_C57BL6J_3646590 | Gm5277 | NCBI_Gene:383883,ENSEMBL:ENSMUSG00000091361 | MGI:3646590 | pseudogene | predicted gene 5277 |
| 3 | pseudogene | 78.91724 | 78.91846 | positive | MGI_C57BL6J_3641638 | Gm10291 | NCBI_Gene:100041748,ENSEMBL:ENSMUSG00000070443 | MGI:3641638 | pseudogene | predicted pseudogene 10291 |
| 3 | gene | 78.91846 | 78.93631 | negative | MGI_C57BL6J_5621427 | Gm38542 | NCBI_Gene:102641233 | MGI:5621427 | lncRNA gene | predicted gene, 38542 |
| 3 | pseudogene | 78.96549 | 78.96715 | negative | MGI_C57BL6J_3704220 | Gm9762 | NCBI_Gene:100041766,ENSEMBL:ENSMUSG00000037096 | MGI:3704220 | pseudogene | predicted pseudogene 9762 |
| 3 | gene | 78.97992 | 78.98005 | positive | MGI_C57BL6J_5453217 | Gm23440 | ENSEMBL:ENSMUSG00000077487 | MGI:5453217 | snoRNA gene | predicted gene, 23440 |
| 3 | gene | 79.06251 | 79.28724 | negative | MGI_C57BL6J_2659071 | Rapgef2 | NCBI_Gene:76089,ENSEMBL:ENSMUSG00000062232 | MGI:2659071 | protein coding gene | Rap guanine nucleotide exchange factor (GEF) 2 |
| 3 | gene | 79.19637 | 79.19648 | positive | MGI_C57BL6J_5453927 | Gm24150 | ENSEMBL:ENSMUSG00000064693 | MGI:5453927 | snRNA gene | predicted gene, 24150 |
| 3 | gene | 79.21292 | 79.25391 | positive | MGI_C57BL6J_1918174 | 4921511C10Rik | NCBI_Gene:70924,ENSEMBL:ENSMUSG00000104585 | MGI:1918174 | lncRNA gene | RIKEN cDNA 4921511C10 gene |
| 3 | gene | 79.23228 | 79.23413 | negative | MGI_C57BL6J_2443462 | B930012P20Rik | NA | NA | unclassified gene | RIKEN cDNA B930012P20 gene |
| 3 | gene | 79.28717 | 79.28985 | positive | MGI_C57BL6J_2686494 | 6430573P05Rik | ENSEMBL:ENSMUSG00000102545 | MGI:2686494 | lncRNA gene | RIKEN cDNA 6430573P05 gene |
| 3 | gene | 79.33420 | 79.33658 | negative | MGI_C57BL6J_5622957 | Gm40072 | NCBI_Gene:105244470 | MGI:5622957 | lncRNA gene | predicted gene, 40072 |
| 3 | gene | 79.33666 | 79.46413 | positive | MGI_C57BL6J_4936993 | Gm17359 | NCBI_Gene:100233207,ENSEMBL:ENSMUSG00000091685 | MGI:4936993 | protein coding gene | predicted gene, 17359 |
| 3 | gene | 79.45597 | 79.56780 | negative | MGI_C57BL6J_2683054 | Fnip2 | NCBI_Gene:329679,ENSEMBL:ENSMUSG00000061175 | MGI:2683054 | protein coding gene | folliculin interacting protein 2 |
| 3 | gene | 79.50979 | 79.51210 | negative | MGI_C57BL6J_5595601 | Gm36442 | NCBI_Gene:102640362 | MGI:5595601 | lncRNA gene | predicted gene, 36442 |
| 3 | gene | 79.55131 | 79.55219 | negative | MGI_C57BL6J_3781690 | Gm3513 | ENSEMBL:ENSMUSG00000089746 | MGI:3781690 | lncRNA gene | predicted gene 3513 |
| 3 | gene | 79.56819 | 79.57332 | positive | MGI_C57BL6J_1923129 | 4930589L23Rik | NCBI_Gene:75879,ENSEMBL:ENSMUSG00000068957 | MGI:1923129 | lncRNA gene | RIKEN cDNA 4930589L23 gene |
| 3 | pseudogene | 79.58259 | 79.58323 | positive | MGI_C57BL6J_5594226 | Gm35067 | ENSEMBL:ENSMUSG00000103564 | MGI:5594226 | pseudogene | predicted gene, 35067 |
| 3 | gene | 79.59134 | 79.60365 | positive | MGI_C57BL6J_1914988 | Ppid | NCBI_Gene:67738,ENSEMBL:ENSMUSG00000027804 | MGI:1914988 | protein coding gene | peptidylprolyl isomerase D (cyclophilin D) |
| 3 | gene | 79.60379 | 79.62950 | negative | MGI_C57BL6J_106100 | Etfdh | NCBI_Gene:66841,ENSEMBL:ENSMUSG00000027809 | MGI:106100 | protein coding gene | electron transferring flavoprotein, dehydrogenase |
| 3 | gene | 79.61054 | 79.61183 | negative | MGI_C57BL6J_5611201 | Gm37973 | ENSEMBL:ENSMUSG00000102370 | MGI:5611201 | unclassified gene | predicted gene, 37973 |
| 3 | gene | 79.62908 | 79.63282 | positive | MGI_C57BL6J_1923189 | 4930579G24Rik | NCBI_Gene:75939,ENSEMBL:ENSMUSG00000027811 | MGI:1923189 | protein coding gene | RIKEN cDNA 4930579G24 gene |
| 3 | gene | 79.64160 | 79.73795 | negative | MGI_C57BL6J_2682211 | Rxfp1 | NCBI_Gene:381489,ENSEMBL:ENSMUSG00000034009 | MGI:2682211 | protein coding gene | relaxin/insulin-like family peptide receptor 1 |
| 3 | gene | 79.67337 | 79.67688 | negative | MGI_C57BL6J_5504099 | Gm26984 | ENSEMBL:ENSMUSG00000098204 | MGI:5504099 | lncRNA gene | predicted gene, 26984 |
| 3 | gene | 79.81256 | 79.85279 | negative | MGI_C57BL6J_1917902 | Tmem144 | NCBI_Gene:70652,ENSEMBL:ENSMUSG00000027956 | MGI:1917902 | protein coding gene | transmembrane protein 144 |
| 3 | gene | 79.83493 | 79.83504 | positive | MGI_C57BL6J_5456197 | Gm26420 | ENSEMBL:ENSMUSG00000077461 | MGI:5456197 | snRNA gene | predicted gene, 26420 |
| 3 | gene | 79.85287 | 79.85666 | positive | MGI_C57BL6J_5622958 | Gm40073 | NCBI_Gene:105244471 | MGI:5622958 | lncRNA gene | predicted gene, 40073 |
| 3 | gene | 79.85929 | 79.88681 | negative | MGI_C57BL6J_5595728 | Gm36569 | NCBI_Gene:102640530,ENSEMBL:ENSMUSG00000102191 | MGI:5595728 | lncRNA gene | predicted gene, 36569 |
| 3 | gene | 79.88447 | 79.94628 | positive | MGI_C57BL6J_1915909 | Gask1b | NCBI_Gene:68659,ENSEMBL:ENSMUSG00000027955 | MGI:1915909 | protein coding gene | golgi associated kinase 1B |
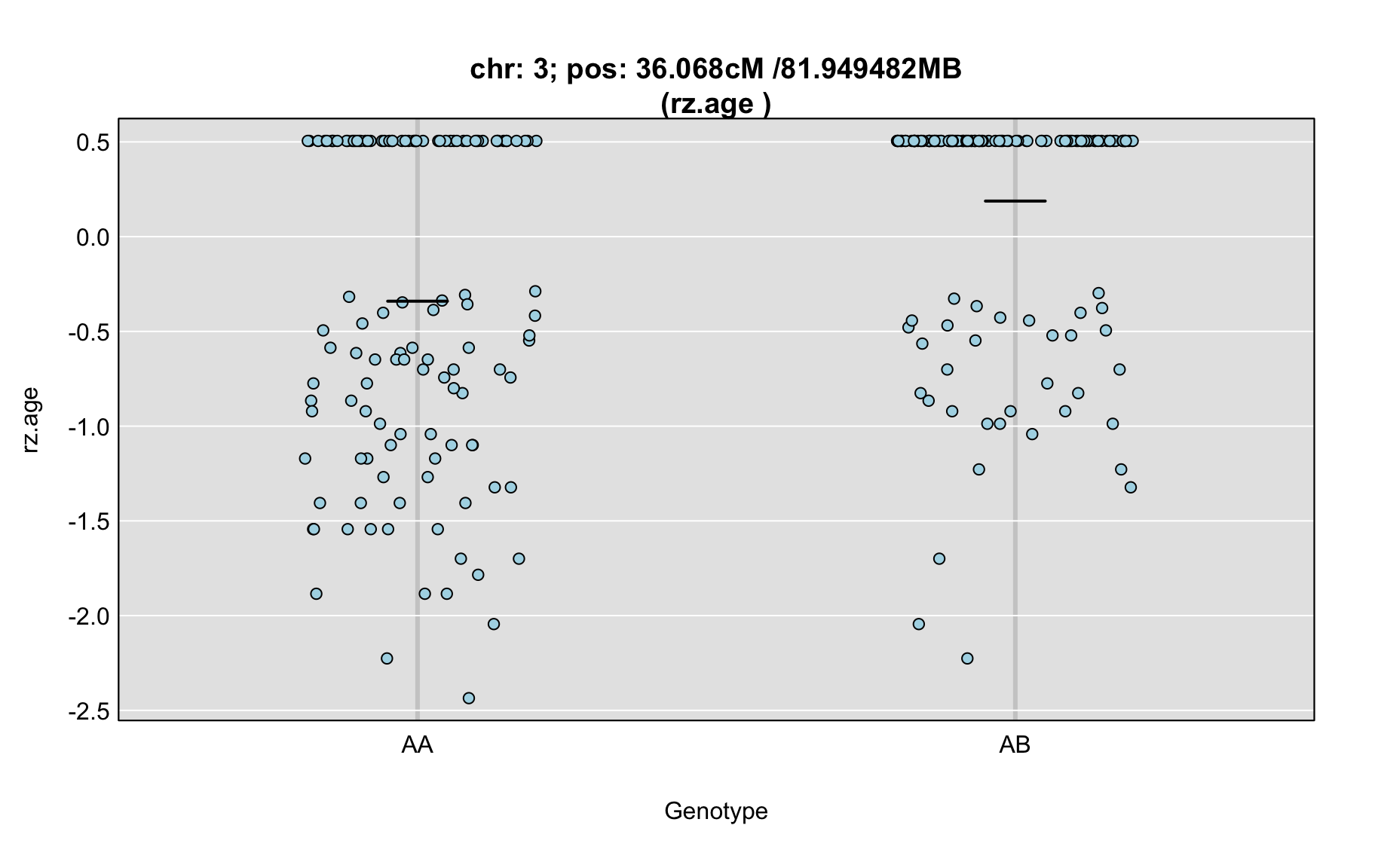
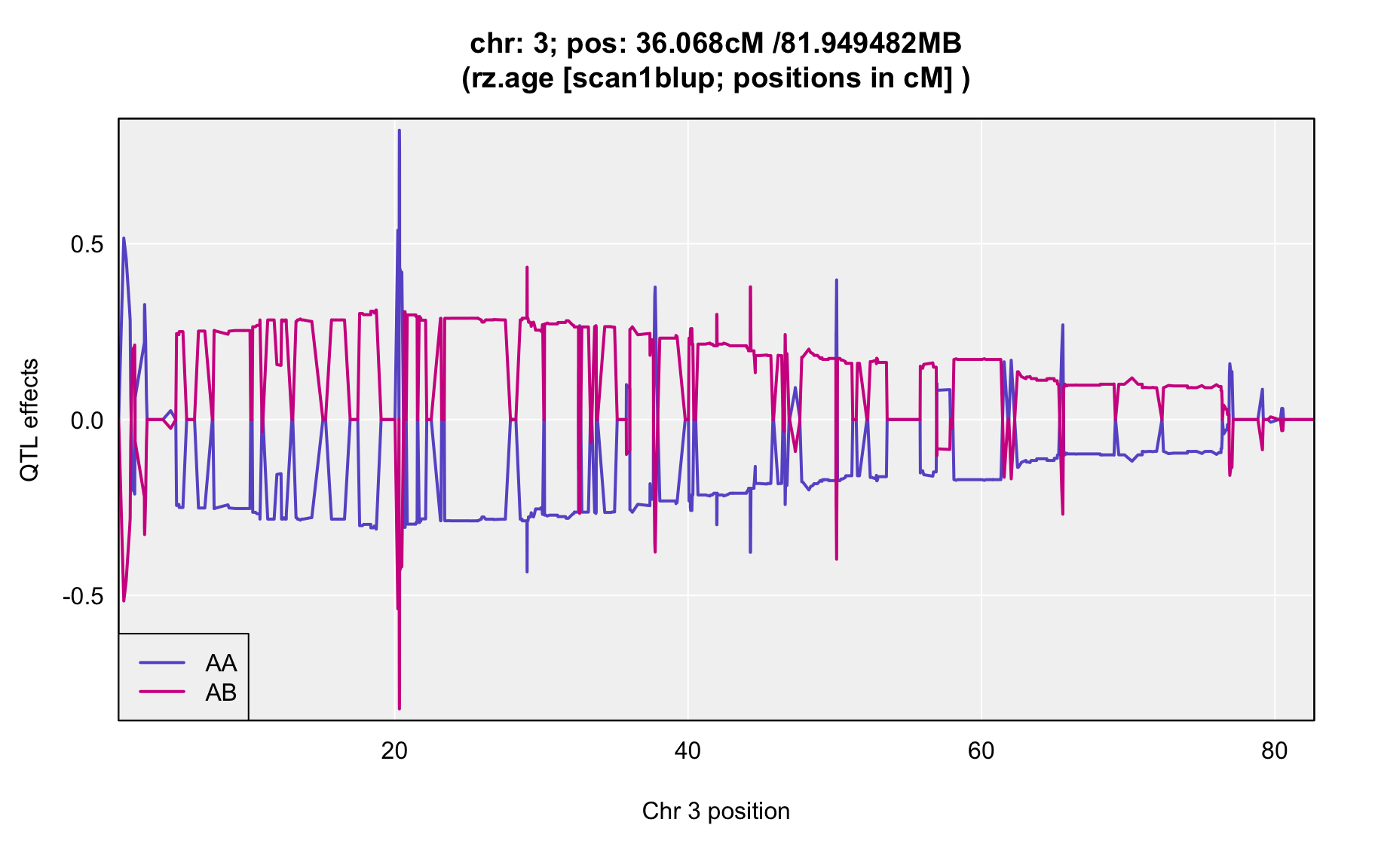
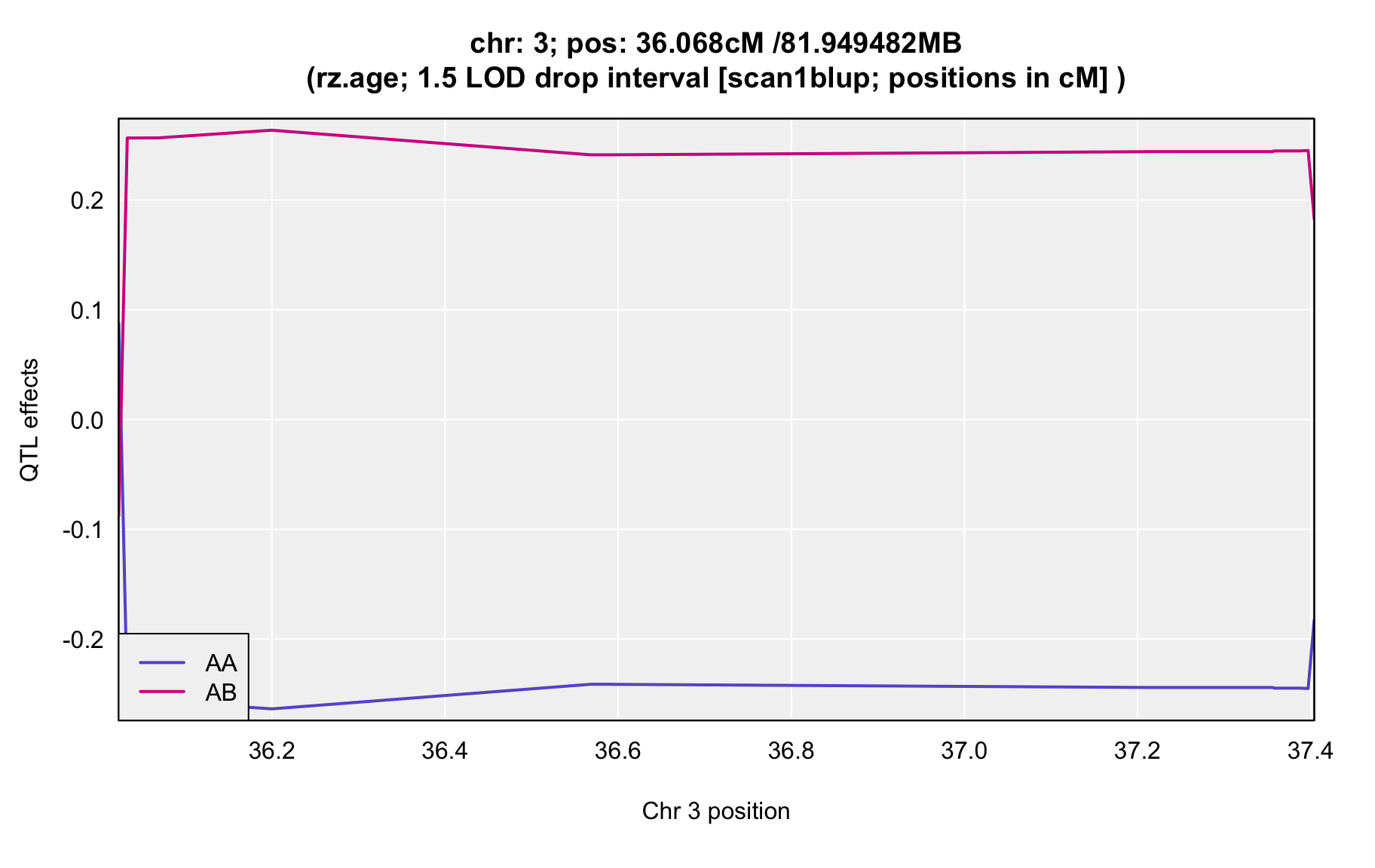
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 81.88073 | 81.88087 | positive | MGI_C57BL6J_5452308 | Gm22531 | ENSEMBL:ENSMUSG00000089444 | MGI:5452308 | snoRNA gene | predicted gene, 22531 |
| 3 | gene | 81.91659 | 81.92467 | negative | MGI_C57BL6J_3642361 | Gm9989 | NCBI_Gene:102640985,ENSEMBL:ENSMUSG00000056023 | MGI:3642361 | lncRNA gene | predicted gene 9989 |
| 3 | gene | 81.93260 | 81.95673 | positive | MGI_C57BL6J_2139628 | Ctso | NCBI_Gene:229445,ENSEMBL:ENSMUSG00000028015 | MGI:2139628 | protein coding gene | cathepsin O |
| 3 | gene | 81.95709 | 81.97620 | negative | MGI_C57BL6J_1928486 | Tdo2 | NCBI_Gene:56720,ENSEMBL:ENSMUSG00000028011 | MGI:1928486 | protein coding gene | tryptophan 2,3-dioxygenase |
| 3 | gene | 81.98229 | 82.02123 | positive | MGI_C57BL6J_1929259 | Asic5 | NCBI_Gene:58170,ENSEMBL:ENSMUSG00000028008 | MGI:1929259 | protein coding gene | acid-sensing (proton-gated) ion channel family member 5 |
| 3 | gene | 82.03200 | 82.07471 | negative | MGI_C57BL6J_1860604 | Gucy1b1 | NCBI_Gene:54195,ENSEMBL:ENSMUSG00000028005 | MGI:1860604 | protein coding gene | guanylate cyclase 1, soluble, beta 1 |
| 3 | gene | 82.09243 | 82.14588 | negative | MGI_C57BL6J_1926562 | Gucy1a1 | NCBI_Gene:60596,ENSEMBL:ENSMUSG00000033910 | MGI:1926562 | protein coding gene | guanylate cyclase 1, soluble, alpha 1 |
| 3 | gene | 82.10625 | 82.10631 | negative | MGI_C57BL6J_5531411 | Mir7010 | miRBase:MI0022859,NCBI_Gene:102466991,ENSEMBL:ENSMUSG00000098712 | MGI:5531411 | miRNA gene | microRNA 7010 |
| 3 | gene | 82.14640 | 82.23680 | negative | MGI_C57BL6J_5589080 | Gm29921 | NCBI_Gene:102631626 | MGI:5589080 | lncRNA gene | predicted gene, 29921 |
| 3 | gene | 82.35803 | 82.39527 | positive | MGI_C57BL6J_2442208 | Map9 | NCBI_Gene:213582,ENSEMBL:ENSMUSG00000033900 | MGI:2442208 | protein coding gene | microtubule-associated protein 9 |
| 3 | gene | 82.53838 | 82.54899 | negative | MGI_C57BL6J_108418 | Npy2r | NCBI_Gene:18167,ENSEMBL:ENSMUSG00000028004 | MGI:108418 | protein coding gene | neuropeptide Y receptor Y2 |
| 3 | pseudogene | 82.57740 | 82.57809 | positive | MGI_C57BL6J_5594482 | Gm35323 | NCBI_Gene:102638861,ENSEMBL:ENSMUSG00000102514 | MGI:5594482 | pseudogene | predicted gene, 35323 |
| 3 | gene | 82.66038 | 82.66490 | negative | MGI_C57BL6J_5589158 | Gm29999 | NCBI_Gene:102631733,ENSEMBL:ENSMUSG00000102629 | MGI:5589158 | lncRNA gene | predicted gene, 29999 |
| 3 | pseudogene | 82.67199 | 82.67257 | negative | MGI_C57BL6J_5663484 | Gm43347 | ENSEMBL:ENSMUSG00000104772 | MGI:5663484 | pseudogene | predicted gene 43347 |
| 3 | gene | 82.75452 | 82.75986 | positive | MGI_C57BL6J_5589202 | Gm30043 | ENSEMBL:ENSMUSG00000104898 | MGI:5589202 | lncRNA gene | predicted gene, 30043 |
| 3 | gene | 82.79366 | 82.79390 | negative | MGI_C57BL6J_5451815 | Gm22038 | ENSEMBL:ENSMUSG00000087902 | MGI:5451815 | unclassified non-coding RNA gene | predicted gene, 22038 |
| 3 | gene | 82.79497 | 82.81223 | negative | MGI_C57BL6J_5663485 | Gm43348 | ENSEMBL:ENSMUSG00000105034 | MGI:5663485 | lncRNA gene | predicted gene 43348 |
| 3 | gene | 82.83652 | 82.87648 | negative | MGI_C57BL6J_3645057 | Rbm46 | NCBI_Gene:633285,ENSEMBL:ENSMUSG00000033882 | MGI:3645057 | protein coding gene | RNA binding motif protein 46 |
| 3 | gene | 82.87671 | 82.94364 | positive | MGI_C57BL6J_1925462 | Rbm46os | NCBI_Gene:78212,ENSEMBL:ENSMUSG00000086273 | MGI:1925462 | antisense lncRNA gene | RNA binding motif protein 46, opposite strand |
| 3 | gene | 82.89258 | 82.90397 | negative | MGI_C57BL6J_1891259 | Lrat | NCBI_Gene:79235,ENSEMBL:ENSMUSG00000028003 | MGI:1891259 | protein coding gene | lecithin-retinol acyltransferase (phosphatidylcholine-retinol-O-acyltransferase) |
| 3 | gene | 82.92666 | 83.00849 | negative | MGI_C57BL6J_5589256 | Gm30097 | NCBI_Gene:102631873,ENSEMBL:ENSMUSG00000102595 | MGI:5589256 | lncRNA gene | predicted gene, 30097 |
| 3 | gene | 83.00772 | 83.01506 | positive | MGI_C57BL6J_95526 | Fgg | NCBI_Gene:99571,ENSEMBL:ENSMUSG00000033860 | MGI:95526 | protein coding gene | fibrinogen gamma chain |
| 3 | gene | 83.02608 | 83.03363 | positive | MGI_C57BL6J_1316726 | Fga | NCBI_Gene:14161,ENSEMBL:ENSMUSG00000028001 | MGI:1316726 | protein coding gene | fibrinogen alpha chain |
| 3 | gene | 83.03708 | 83.03897 | negative | MGI_C57BL6J_5622960 | Gm40075 | NCBI_Gene:105244473 | MGI:5622960 | lncRNA gene | predicted gene, 40075 |
| 3 | gene | 83.04014 | 83.04986 | negative | MGI_C57BL6J_99501 | Fgb | NCBI_Gene:110135,ENSEMBL:ENSMUSG00000033831 | MGI:99501 | protein coding gene | fibrinogen beta chain |
| 3 | gene | 83.05552 | 83.07236 | positive | MGI_C57BL6J_1858197 | Plrg1 | NCBI_Gene:53317,ENSEMBL:ENSMUSG00000027998 | MGI:1858197 | protein coding gene | pleiotropic regulator 1 |
| 3 | gene | 83.12528 | 83.12925 | negative | MGI_C57BL6J_3642896 | Gm10710 | NCBI_Gene:100038599,ENSEMBL:ENSMUSG00000074517 | MGI:3642896 | lncRNA gene | predicted gene 10710 |
| 3 | gene | 83.12795 | 83.35721 | positive | MGI_C57BL6J_2685263 | Dchs2 | NCBI_Gene:100534287,ENSEMBL:ENSMUSG00000102692 | MGI:2685263 | protein coding gene | dachsous cadherin related 2 |
| 3 | gene | 83.40340 | 83.40579 | positive | MGI_C57BL6J_5611324 | Gm38096 | ENSEMBL:ENSMUSG00000103459 | MGI:5611324 | lncRNA gene | predicted gene, 38096 |
| 3 | gene | 83.55536 | 83.57412 | negative | MGI_C57BL6J_1916710 | 1700028M03Rik | NCBI_Gene:69460,ENSEMBL:ENSMUSG00000101334 | MGI:1916710 | lncRNA gene | RIKEN cDNA 1700028M03 gene |
| 3 | gene | 83.59633 | 83.60109 | negative | MGI_C57BL6J_5622961 | Gm40076 | NCBI_Gene:105244474 | MGI:5622961 | lncRNA gene | predicted gene, 40076 |
| 3 | gene | 83.63458 | 83.63487 | negative | MGI_C57BL6J_5452119 | Gm22342 | ENSEMBL:ENSMUSG00000084596 | MGI:5452119 | unclassified non-coding RNA gene | predicted gene, 22342 |
| 3 | pseudogene | 83.68668 | 83.68727 | positive | MGI_C57BL6J_3781837 | Rpl28-ps2 | NCBI_Gene:100042091,ENSEMBL:ENSMUSG00000102137 | MGI:3781837 | pseudogene | ribosomal protein L28, pseudogene 2 |
| 3 | gene | 83.70225 | 83.70236 | positive | MGI_C57BL6J_5454361 | Gm24584 | ENSEMBL:ENSMUSG00000088597 | MGI:5454361 | snRNA gene | predicted gene, 24584 |
| 3 | gene | 83.71936 | 83.72737 | negative | MGI_C57BL6J_5589452 | Gm30293 | NCBI_Gene:102632140 | MGI:5589452 | lncRNA gene | predicted gene, 30293 |
| 3 | gene | 83.76616 | 83.77432 | positive | MGI_C57BL6J_108078 | Sfrp2 | NCBI_Gene:20319,ENSEMBL:ENSMUSG00000027996 | MGI:108078 | protein coding gene | secreted frizzled-related protein 2 |
| 3 | gene | 83.77385 | 83.78996 | negative | MGI_C57BL6J_5477265 | Gm26771 | ENSEMBL:ENSMUSG00000097250 | MGI:5477265 | lncRNA gene | predicted gene, 26771 |
| 3 | pseudogene | 83.78973 | 83.83222 | positive | MGI_C57BL6J_3646388 | Gm7115 | NCBI_Gene:633522,ENSEMBL:ENSMUSG00000104082 | MGI:3646388 | pseudogene | predicted gene 7115 |
| 3 | gene | 83.83627 | 83.84182 | negative | MGI_C57BL6J_1346060 | Tlr2 | NCBI_Gene:24088,ENSEMBL:ENSMUSG00000027995 | MGI:1346060 | protein coding gene | toll-like receptor 2 |
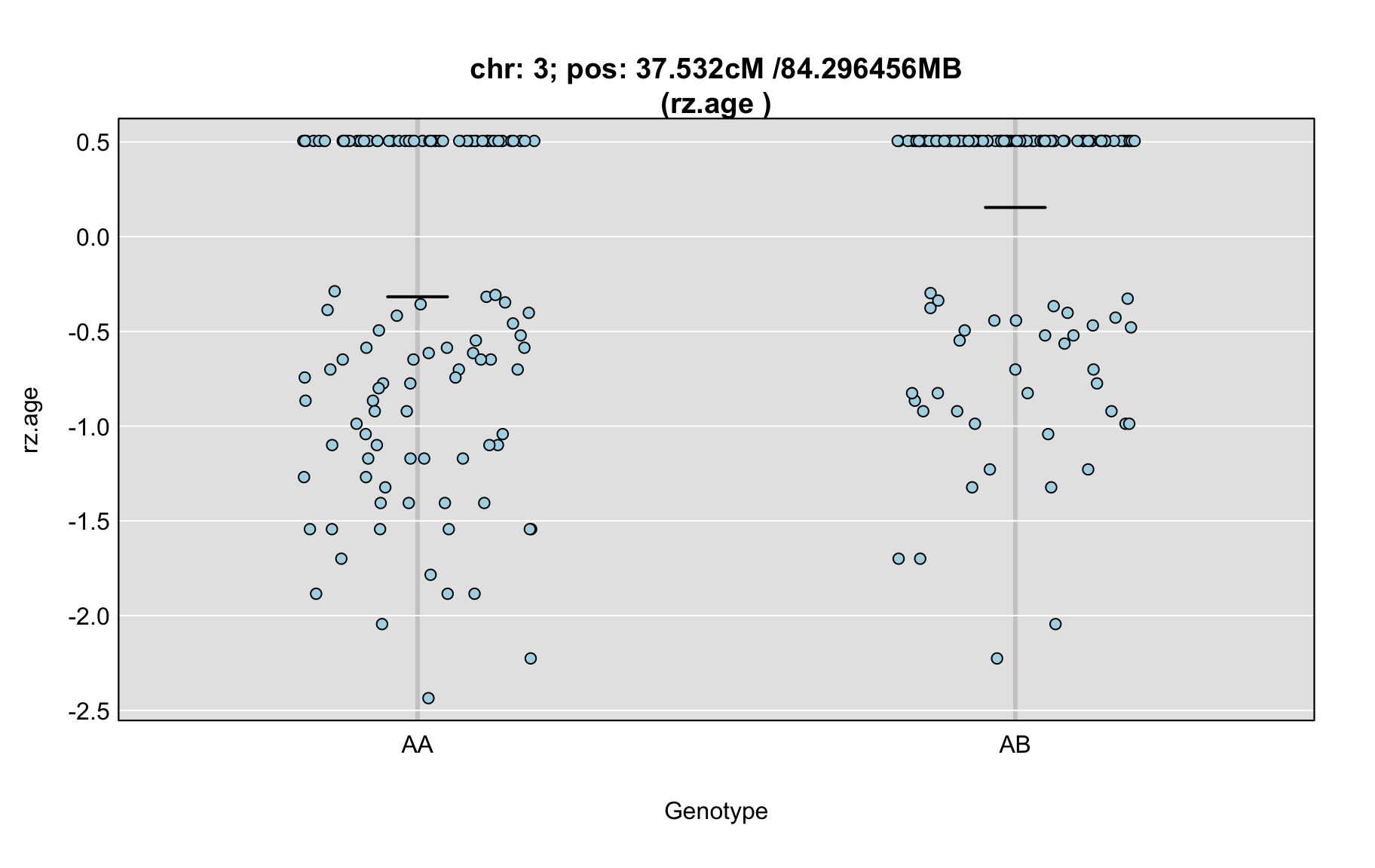
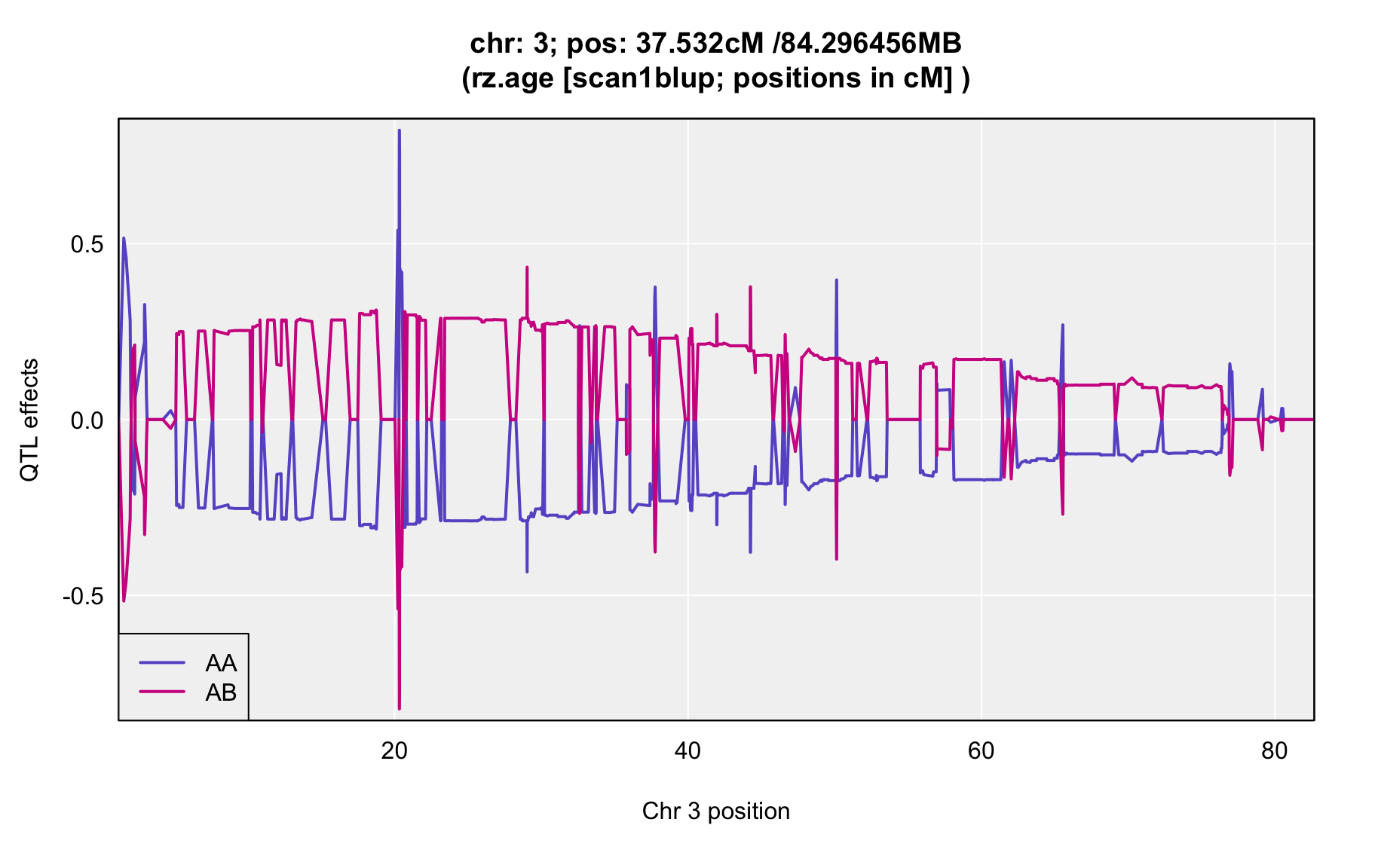
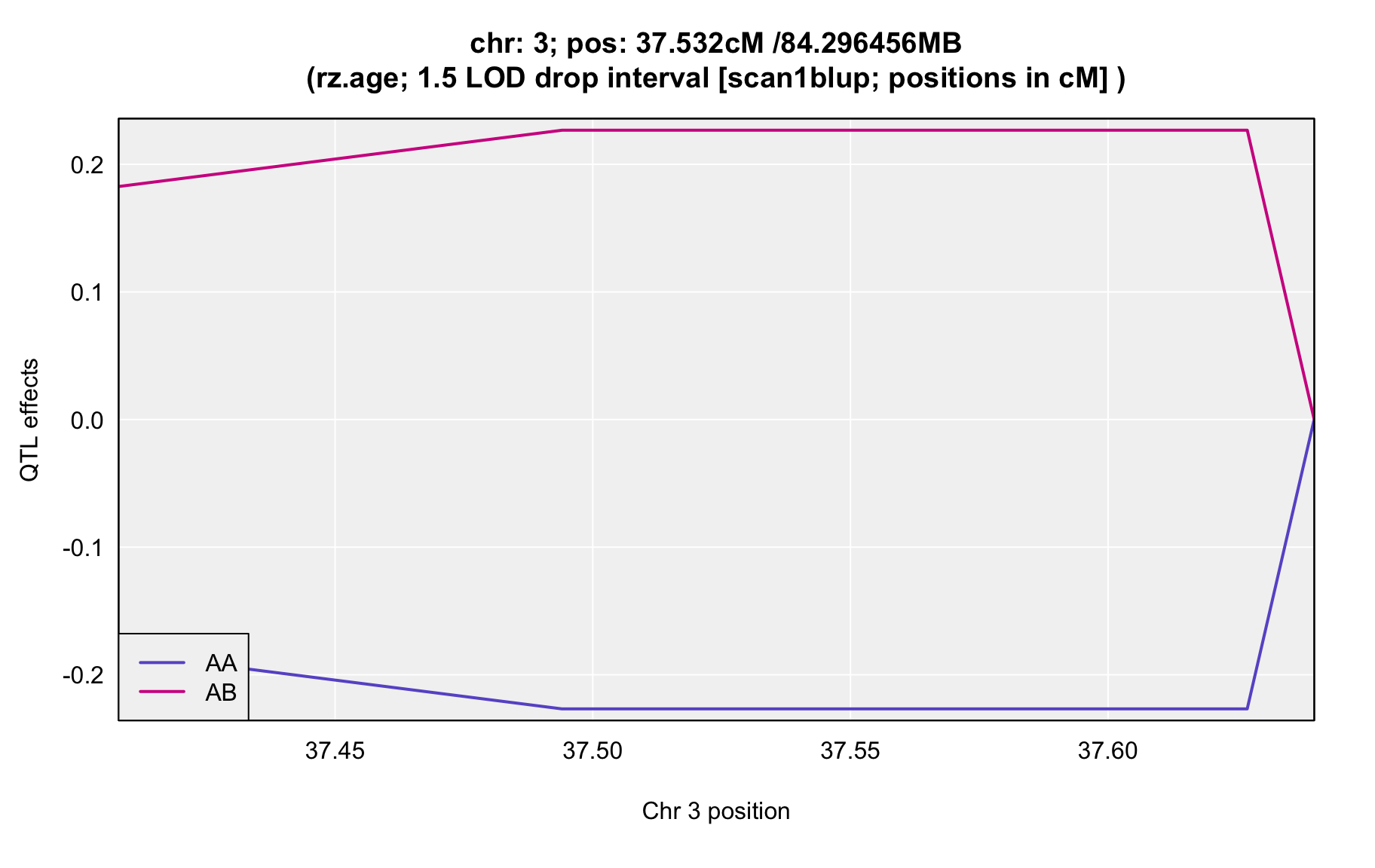
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 83.89766 | 84.04018 | negative | MGI_C57BL6J_2443399 | Tmem131l | NCBI_Gene:229473,ENSEMBL:ENSMUSG00000033767 | MGI:2443399 | protein coding gene | transmembrane 131 like |
| 3 | gene | 84.02844 | 84.02928 | positive | MGI_C57BL6J_1922302 | 4930509B17Rik | NA | NA | unclassified gene | RIKEN cDNA 4930509B17 gene |
| 3 | gene | 84.04022 | 84.05210 | positive | MGI_C57BL6J_5589511 | Gm30352 | NCBI_Gene:102632212 | MGI:5589511 | lncRNA gene | predicted gene, 30352 |
| 3 | gene | 84.05897 | 84.06345 | negative | MGI_C57BL6J_5610209 | Gm36981 | ENSEMBL:ENSMUSG00000103261 | MGI:5610209 | unclassified non-coding RNA gene | predicted gene, 36981 |
| 3 | gene | 84.06174 | 84.06277 | positive | MGI_C57BL6J_5589617 | Gm30458 | NCBI_Gene:102632366 | MGI:5589617 | lncRNA gene | predicted gene, 30458 |
| 3 | gene | 84.08793 | 84.15579 | negative | MGI_C57BL6J_1924165 | Mnd1 | NCBI_Gene:76915,ENSEMBL:ENSMUSG00000033752 | MGI:1924165 | protein coding gene | meiotic nuclear divisions 1 |
| 3 | gene | 84.16044 | 84.30688 | negative | MGI_C57BL6J_1933163 | Trim2 | NCBI_Gene:80890,ENSEMBL:ENSMUSG00000027993 | MGI:1933163 | protein coding gene | tripartite motif-containing 2 |
| 3 | gene | 84.17410 | 84.17519 | positive | MGI_C57BL6J_3648757 | Gm6525 | NCBI_Gene:624713,ENSEMBL:ENSMUSG00000104043 | MGI:3648757 | protein coding gene | predicted pseudogene 6525 |
| 3 | gene | 84.22869 | 84.24704 | positive | MGI_C57BL6J_5622963 | Gm40078 | NCBI_Gene:105244477 | MGI:5622963 | lncRNA gene | predicted gene, 40078 |
| 3 | gene | 84.22901 | 84.22908 | negative | MGI_C57BL6J_4413878 | n-TGgcc9 | NCBI_Gene:102467313 | MGI:4413878 | tRNA gene | nuclear encoded tRNA glycine 9 (anticodon GCC) |
| 3 | gene | 84.30298 | 84.30542 | negative | MGI_C57BL6J_2443303 | C630001G18Rik | NA | NA | unclassified gene | RIKEN cDNA C630001G18 gene |
| 3 | gene | 84.34938 | 84.37992 | negative | MGI_C57BL6J_1922559 | 4930565D16Rik | NCBI_Gene:75309,ENSEMBL:ENSMUSG00000104004 | MGI:1922559 | lncRNA gene | RIKEN cDNA 4930565D16 gene |
| 3 | gene | 84.44219 | 84.48044 | negative | MGI_C57BL6J_2684972 | Fhdc1 | NCBI_Gene:229474,ENSEMBL:ENSMUSG00000041842 | MGI:2684972 | protein coding gene | FH2 domain containing 1 |
| 3 | gene | 84.49609 | 84.58283 | negative | MGI_C57BL6J_1277120 | Arfip1 | NCBI_Gene:99889,ENSEMBL:ENSMUSG00000074513 | MGI:1277120 | protein coding gene | ADP-ribosylation factor interacting protein 1 |
| 3 | gene | 84.49643 | 85.88752 | negative | MGI_C57BL6J_5610468 | Gm37240 | ENSEMBL:ENSMUSG00000102805 | MGI:5610468 | protein coding gene | predicted gene, 37240 |
| 3 | pseudogene | 84.55266 | 84.55348 | negative | MGI_C57BL6J_3645282 | Gm8813 | NCBI_Gene:667788,ENSEMBL:ENSMUSG00000085625 | MGI:3645282 | pseudogene | predicted gene 8813 |
| 3 | pseudogene | 84.55689 | 84.55779 | negative | MGI_C57BL6J_5611469 | Gm38241 | ENSEMBL:ENSMUSG00000103741 | MGI:5611469 | pseudogene | predicted gene, 38241 |
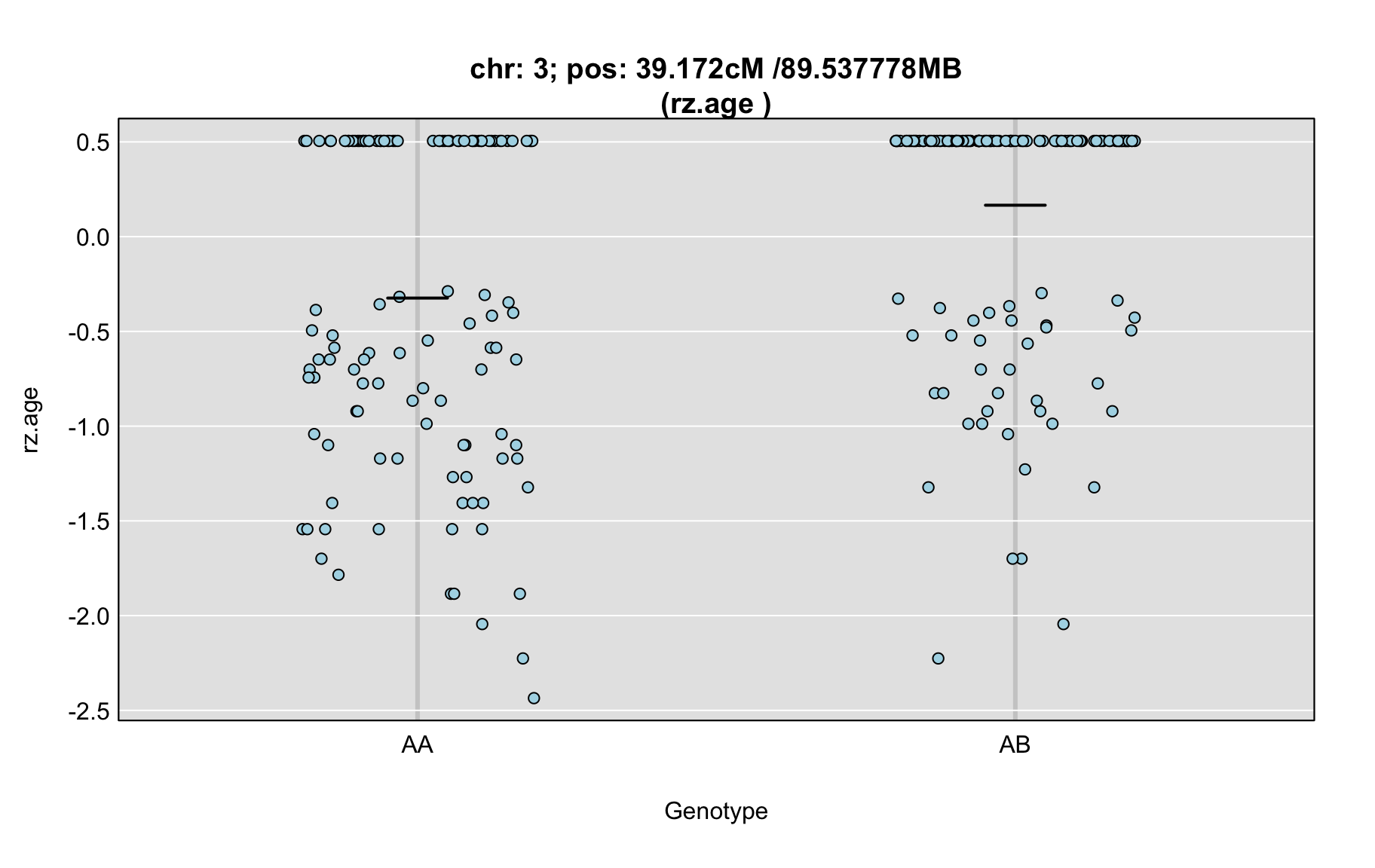
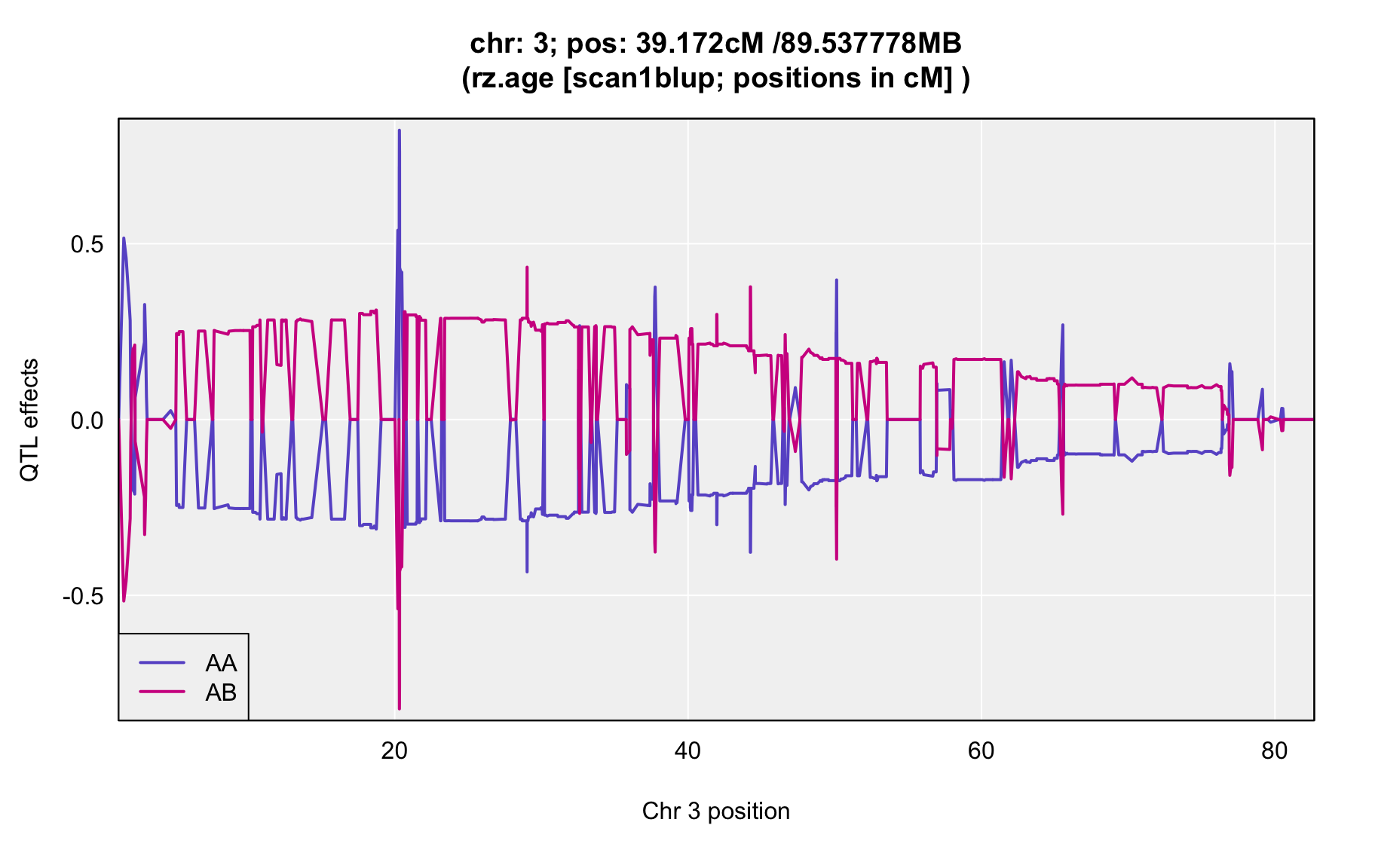
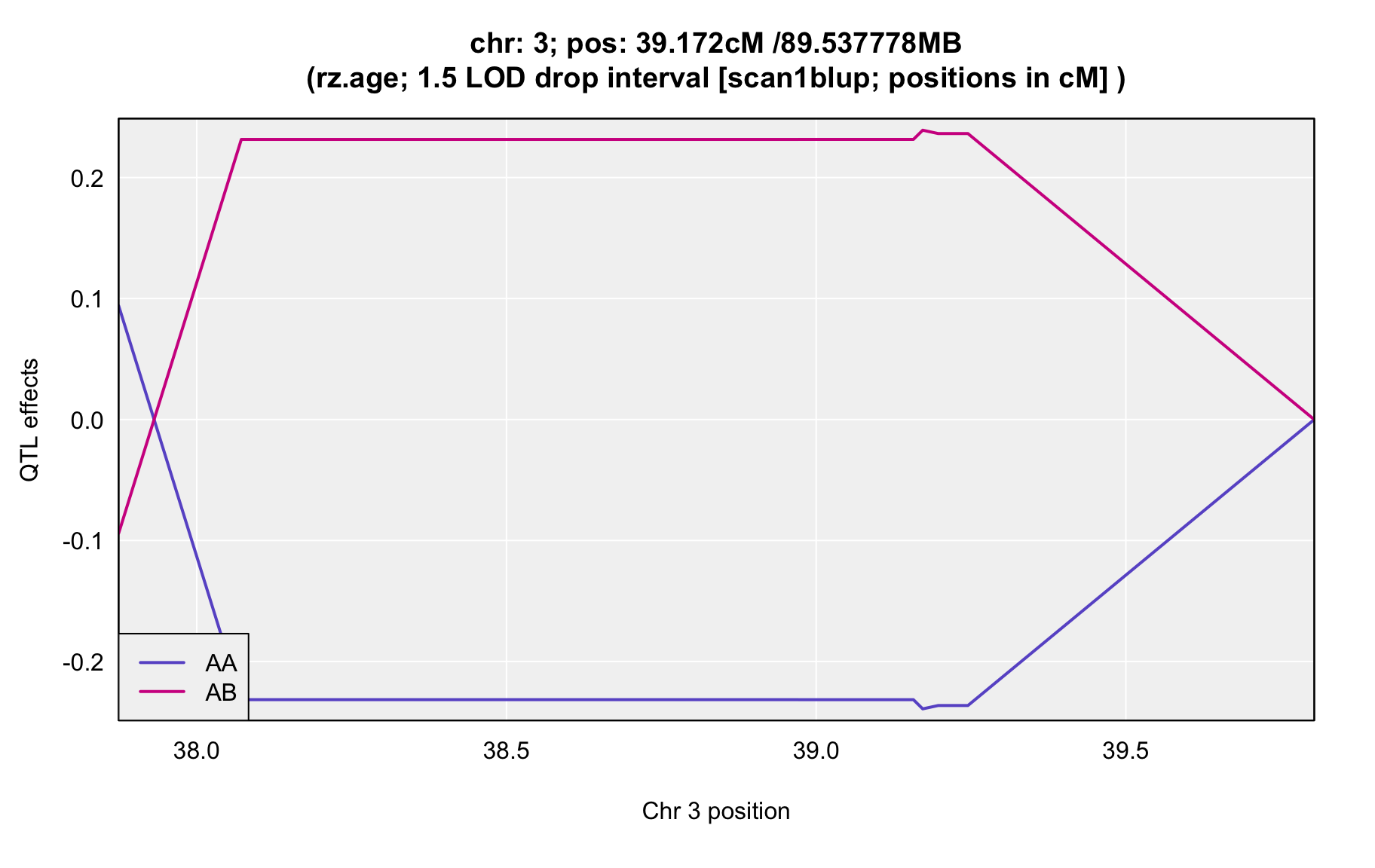
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 86.22390 | 86.78269 | positive | MGI_C57BL6J_1933162 | Lrba | NCBI_Gene:80877,ENSEMBL:ENSMUSG00000028080 | MGI:1933162 | protein coding gene | LPS-responsive beige-like anchor |
| 3 | pseudogene | 86.67177 | 86.67224 | negative | MGI_C57BL6J_3781961 | Gm3788 | NCBI_Gene:100042319,ENSEMBL:ENSMUSG00000094392 | MGI:3781961 | pseudogene | predicted gene 3788 |
| 3 | gene | 86.76807 | 86.77865 | negative | MGI_C57BL6J_5611104 | Gm37876 | ENSEMBL:ENSMUSG00000104108 | MGI:5611104 | lncRNA gene | predicted gene, 37876 |
| 3 | gene | 86.78615 | 86.92088 | negative | MGI_C57BL6J_1918012 | Dclk2 | NCBI_Gene:70762,ENSEMBL:ENSMUSG00000028078 | MGI:1918012 | protein coding gene | doublecortin-like kinase 2 |
| 3 | gene | 86.82565 | 86.82710 | negative | MGI_C57BL6J_5826447 | Gm46810 | NCBI_Gene:108168877 | MGI:5826447 | lncRNA gene | predicted gene, 46810 |
| 3 | pseudogene | 86.86449 | 86.86532 | positive | MGI_C57BL6J_5610253 | Gm37025 | ENSEMBL:ENSMUSG00000104074 | MGI:5610253 | pseudogene | predicted gene, 37025 |
| 3 | gene | 86.93074 | 86.93090 | positive | MGI_C57BL6J_5453153 | Gm23376 | ENSEMBL:ENSMUSG00000064872 | MGI:5453153 | snRNA gene | predicted gene, 23376 |
| 3 | pseudogene | 86.93337 | 86.93366 | negative | MGI_C57BL6J_5610435 | Gm37207 | ENSEMBL:ENSMUSG00000103650 | MGI:5610435 | pseudogene | predicted gene, 37207 |
| 3 | pseudogene | 86.98655 | 86.98978 | positive | MGI_C57BL6J_107675 | Cd1d2 | NCBI_Gene:12480,ENSEMBL:ENSMUSG00000041750 | MGI:107675 | polymorphic pseudogene | CD1d2 antigen |
| 3 | gene | 86.99583 | 86.99944 | negative | MGI_C57BL6J_107674 | Cd1d1 | NCBI_Gene:12479,ENSEMBL:ENSMUSG00000028076 | MGI:107674 | protein coding gene | CD1d1 antigen |
| 3 | pseudogene | 87.02286 | 87.02396 | negative | MGI_C57BL6J_3644826 | Gm8869 | NCBI_Gene:667897,ENSEMBL:ENSMUSG00000104265 | MGI:3644826 | pseudogene | predicted gene 8869 |
| 3 | pseudogene | 87.05436 | 87.05500 | positive | MGI_C57BL6J_5010588 | Gm18403 | NCBI_Gene:100417112,ENSEMBL:ENSMUSG00000103913 | MGI:5010588 | pseudogene | predicted gene, 18403 |
| 3 | pseudogene | 87.06012 | 87.06043 | positive | MGI_C57BL6J_5610803 | Gm37575 | ENSEMBL:ENSMUSG00000102394 | MGI:5610803 | pseudogene | predicted gene, 37575 |
| 3 | gene | 87.06689 | 87.06702 | positive | MGI_C57BL6J_5454249 | Gm24472 | ENSEMBL:ENSMUSG00000077672 | MGI:5454249 | snoRNA gene | predicted gene, 24472 |
| 3 | gene | 87.07859 | 87.17475 | negative | MGI_C57BL6J_1891396 | Kirrel | NCBI_Gene:170643,ENSEMBL:ENSMUSG00000041734 | MGI:1891396 | protein coding gene | kirre like nephrin family adhesion molecule 1 |
| 3 | gene | 87.11060 | 87.11222 | negative | MGI_C57BL6J_5011567 | Gm19382 | NA | NA | unclassified gene | predicted gene%2c 19382 |
| 3 | gene | 87.15769 | 87.16245 | positive | MGI_C57BL6J_5611083 | Gm37855 | ENSEMBL:ENSMUSG00000102893 | MGI:5611083 | lncRNA gene | predicted gene, 37855 |
| 3 | pseudogene | 87.20734 | 87.20803 | negative | MGI_C57BL6J_3708678 | Gm10705 | NCBI_Gene:100042355,ENSEMBL:ENSMUSG00000074506 | MGI:3708678 | pseudogene | predicted gene 10705 |
| 3 | gene | 87.25076 | 87.26377 | negative | MGI_C57BL6J_1933397 | Fcrls | NCBI_Gene:80891,ENSEMBL:ENSMUSG00000015852 | MGI:1933397 | protein coding gene | Fc receptor-like S, scavenger receptor |
| 3 | gene | 87.29034 | 87.30091 | positive | MGI_C57BL6J_5591038 | Gm31879 | NCBI_Gene:102634249 | MGI:5591038 | lncRNA gene | predicted gene, 31879 |
| 3 | gene | 87.35400 | 87.37760 | negative | MGI_C57BL6J_5591099 | Gm31940 | NCBI_Gene:102634332 | MGI:5591099 | lncRNA gene | predicted gene, 31940 |
| 3 | gene | 87.35788 | 87.37107 | positive | MGI_C57BL6J_1334419 | Cd5l | NCBI_Gene:11801,ENSEMBL:ENSMUSG00000015854 | MGI:1334419 | protein coding gene | CD5 antigen-like |
| 3 | gene | 87.37634 | 87.40397 | positive | MGI_C57BL6J_2442862 | Fcrl1 | NCBI_Gene:229499,ENSEMBL:ENSMUSG00000059994 | MGI:2442862 | protein coding gene | Fc receptor-like 1 |
| 3 | gene | 87.41141 | 87.41479 | positive | MGI_C57BL6J_5622969 | Gm40084 | NCBI_Gene:105244483 | MGI:5622969 | lncRNA gene | predicted gene, 40084 |
| 3 | gene | 87.43577 | 87.50068 | positive | MGI_C57BL6J_3053558 | Fcrl5 | NCBI_Gene:329693,ENSEMBL:ENSMUSG00000048031 | MGI:3053558 | protein coding gene | Fc receptor-like 5 |
| 3 | gene | 87.48702 | 87.51219 | negative | MGI_C57BL6J_5622970 | Gm40085 | NCBI_Gene:105244484 | MGI:5622970 | lncRNA gene | predicted gene, 40085 |
| 3 | gene | 87.50853 | 87.51318 | positive | MGI_C57BL6J_5591157 | Gm31998 | NCBI_Gene:102634410 | MGI:5591157 | lncRNA gene | predicted gene, 31998 |
| 3 | gene | 87.52541 | 87.54016 | positive | MGI_C57BL6J_1350926 | Etv3 | NCBI_Gene:27049,ENSEMBL:ENSMUSG00000003382 | MGI:1350926 | protein coding gene | ets variant 3 |
| 3 | gene | 87.54988 | 87.56110 | positive | MGI_C57BL6J_3646099 | Etv3l | NCBI_Gene:546801,ENSEMBL:ENSMUSG00000104058 | MGI:3646099 | protein coding gene | ets variant 3-like |
| 3 | gene | 87.58267 | 87.58279 | negative | MGI_C57BL6J_5454944 | Gm25167 | ENSEMBL:ENSMUSG00000077376 | MGI:5454944 | snoRNA gene | predicted gene, 25167 |
| 3 | gene | 87.61740 | 87.61750 | positive | MGI_C57BL6J_5453886 | Gm24109 | ENSEMBL:ENSMUSG00000093230 | MGI:5453886 | miRNA gene | predicted gene, 24109 |
| 3 | gene | 87.61755 | 87.73803 | positive | MGI_C57BL6J_2441869 | Arhgef11 | NCBI_Gene:213498,ENSEMBL:ENSMUSG00000041977 | MGI:2441869 | protein coding gene | Rho guanine nucleotide exchange factor (GEF) 11 |
| 3 | pseudogene | 87.64843 | 87.64867 | negative | MGI_C57BL6J_5011427 | Gm19242 | NCBI_Gene:100462734 | MGI:5011427 | pseudogene | predicted gene, 19242 |
| 3 | gene | 87.67964 | 87.68003 | negative | MGI_C57BL6J_3644100 | Gm6570 | NCBI_Gene:625281 | MGI:3644100 | protein coding gene | predicted gene 6570 |
| 3 | gene | 87.73692 | 87.74863 | negative | MGI_C57BL6J_1921735 | Lrrc71 | NCBI_Gene:74485,ENSEMBL:ENSMUSG00000023084 | MGI:1921735 | protein coding gene | leucine rich repeat containing 71 |
| 3 | gene | 87.74789 | 87.82431 | negative | MGI_C57BL6J_1920432 | Pear1 | NCBI_Gene:73182,ENSEMBL:ENSMUSG00000028073 | MGI:1920432 | protein coding gene | platelet endothelial aggregation receptor 1 |
| 3 | gene | 87.77824 | 87.79524 | negative | MGI_C57BL6J_97383 | Ntrk1 | NCBI_Gene:18211,ENSEMBL:ENSMUSG00000028072 | MGI:97383 | protein coding gene | neurotrophic tyrosine kinase, receptor, type 1 |
| 3 | gene | 87.79691 | 87.81610 | positive | MGI_C57BL6J_1346037 | Insrr | NCBI_Gene:23920,ENSEMBL:ENSMUSG00000005640 | MGI:1346037 | protein coding gene | insulin receptor-related receptor |
| 3 | gene | 87.82054 | 87.82307 | positive | MGI_C57BL6J_5622971 | Gm40086 | NCBI_Gene:105244485 | MGI:5622971 | lncRNA gene | predicted gene, 40086 |
| 3 | gene | 87.82475 | 87.83127 | positive | MGI_C57BL6J_5591629 | Gm32470 | NCBI_Gene:105244486 | MGI:5591629 | lncRNA gene | predicted gene, 32470 |
| 3 | gene | 87.84676 | 87.85572 | positive | MGI_C57BL6J_1351596 | Sh2d2a | NCBI_Gene:27371,ENSEMBL:ENSMUSG00000028071 | MGI:1351596 | protein coding gene | SH2 domain containing 2A |
| 3 | gene | 87.85890 | 87.88561 | negative | MGI_C57BL6J_2137738 | Prcc | NCBI_Gene:94315,ENSEMBL:ENSMUSG00000004895 | MGI:2137738 | protein coding gene | papillary renal cell carcinoma (translocation-associated) |
| 3 | gene | 87.90632 | 87.91613 | positive | MGI_C57BL6J_1194494 | Hdgf | NCBI_Gene:15191,ENSEMBL:ENSMUSG00000004897 | MGI:1194494 | protein coding gene | heparin binding growth factor |
| 3 | gene | 87.91813 | 87.92367 | positive | MGI_C57BL6J_1914957 | Mrpl24 | NCBI_Gene:67707,ENSEMBL:ENSMUSG00000019710 | MGI:1914957 | protein coding gene | mitochondrial ribosomal protein L24 |
| 3 | gene | 87.92260 | 87.93072 | negative | MGI_C57BL6J_2387197 | Rrnad1 | NCBI_Gene:229503,ENSEMBL:ENSMUSG00000004896 | MGI:2387197 | protein coding gene | ribosomal RNA adenine dimethylase domain containing 1 |
| 3 | gene | 87.93031 | 87.94069 | positive | MGI_C57BL6J_2140076 | Isg20l2 | NCBI_Gene:229504,ENSEMBL:ENSMUSG00000048039 | MGI:2140076 | protein coding gene | interferon stimulated exonuclease gene 20-like 2 |
| 3 | gene | 87.94867 | 87.95338 | positive | MGI_C57BL6J_88491 | Crabp2 | NCBI_Gene:12904,ENSEMBL:ENSMUSG00000004885 | MGI:88491 | protein coding gene | cellular retinoic acid binding protein II |
| 3 | gene | 87.96008 | 87.96573 | positive | MGI_C57BL6J_5591677 | Gm32518 | NCBI_Gene:102635095 | MGI:5591677 | lncRNA gene | predicted gene, 32518 |
| 3 | pseudogene | 87.96676 | 87.96869 | positive | MGI_C57BL6J_3781920 | Gm3745 | NCBI_Gene:100042242,ENSEMBL:ENSMUSG00000103792 | MGI:3781920 | pseudogene | predicted gene 3745 |
| 3 | gene | 87.97108 | 87.98045 | positive | MGI_C57BL6J_101784 | Nes | NCBI_Gene:18008,ENSEMBL:ENSMUSG00000004891 | MGI:101784 | protein coding gene | nestin |
| 3 | gene | 87.98753 | 88.00089 | negative | MGI_C57BL6J_1096385 | Bcan | NCBI_Gene:12032,ENSEMBL:ENSMUSG00000004892 | MGI:1096385 | protein coding gene | brevican |
| 3 | gene | 88.02056 | 88.02758 | negative | MGI_C57BL6J_2137300 | Hapln2 | NCBI_Gene:73940,ENSEMBL:ENSMUSG00000004894 | MGI:2137300 | protein coding gene | hyaluronan and proteoglycan link protein 2 |
| 3 | gene | 88.03078 | 88.03621 | positive | MGI_C57BL6J_5011624 | Gm19439 | NCBI_Gene:105244487,ENSEMBL:ENSMUSG00000105843 | MGI:5011624 | lncRNA gene | predicted gene, 19439 |
| 3 | gene | 88.04311 | 88.05599 | positive | MGI_C57BL6J_1913864 | Gpatch4 | NCBI_Gene:66614,ENSEMBL:ENSMUSG00000028069 | MGI:1913864 | protein coding gene | G patch domain containing 4 |
| 3 | gene | 88.05652 | 88.05854 | negative | MGI_C57BL6J_2180167 | Naxe | NCBI_Gene:246703,ENSEMBL:ENSMUSG00000028070 | MGI:2180167 | protein coding gene | NAD(P)HX epimerase |
| 3 | gene | 88.06941 | 88.07830 | negative | MGI_C57BL6J_2443841 | Ttc24 | NCBI_Gene:214191,ENSEMBL:ENSMUSG00000051036 | MGI:2443841 | protein coding gene | tetratricopeptide repeat domain 24 |
| 3 | gene | 88.08197 | 88.12105 | positive | MGI_C57BL6J_3028642 | Iqgap3 | NCBI_Gene:404710,ENSEMBL:ENSMUSG00000028068 | MGI:3028642 | protein coding gene | IQ motif containing GTPase activating protein 3 |
| 3 | gene | 88.13352 | 88.13842 | positive | MGI_C57BL6J_5622972 | Gm40087 | NCBI_Gene:105244489 | MGI:5622972 | lncRNA gene | predicted gene, 40087 |
| 3 | gene | 88.14197 | 88.17209 | positive | MGI_C57BL6J_99533 | Mef2d | NCBI_Gene:17261,ENSEMBL:ENSMUSG00000001419 | MGI:99533 | protein coding gene | myocyte enhancer factor 2D |
| 3 | gene | 88.17156 | 88.17779 | negative | MGI_C57BL6J_1923892 | 1700113A16Rik | NCBI_Gene:76642,ENSEMBL:ENSMUSG00000096935 | MGI:1923892 | lncRNA gene | RIKEN cDNA 1700113A16 gene |
| 3 | gene | 88.19910 | 88.20459 | negative | MGI_C57BL6J_2139818 | AI849053 | ENSEMBL:ENSMUSG00000097280 | MGI:2139818 | lncRNA gene | expressed sequence AI849053 |
| 3 | gene | 88.20020 | 88.20348 | positive | MGI_C57BL6J_2139951 | AW047730 | NCBI_Gene:99870,ENSEMBL:ENSMUSG00000097428 | MGI:2139951 | lncRNA gene | expressed sequence AW047730 |
| 3 | gene | 88.20577 | 88.22996 | positive | MGI_C57BL6J_3781938 | Gm3764 | NCBI_Gene:100042277,ENSEMBL:ENSMUSG00000097156 | MGI:3781938 | lncRNA gene | predicted gene 3764 |
| 3 | gene | 88.21517 | 88.21526 | positive | MGI_C57BL6J_4834307 | Mir3093 | miRBase:MI0014086,NCBI_Gene:100526540,ENSEMBL:ENSMUSG00000092988 | MGI:4834307 | miRNA gene | microRNA 3093 |
| 3 | gene | 88.21559 | 88.21570 | negative | MGI_C57BL6J_5455418 | Gm25641 | ENSEMBL:ENSMUSG00000092876 | MGI:5455418 | miRNA gene | predicted gene, 25641 |
| 3 | gene | 88.21560 | 88.21569 | positive | MGI_C57BL6J_2676911 | Mir9-1 | miRBase:MI0000720,NCBI_Gene:387133,ENSEMBL:ENSMUSG00000106493 | MGI:2676911 | miRNA gene | microRNA 9-1 |
| 3 | gene | 88.22741 | 88.23234 | negative | MGI_C57BL6J_1918213 | 4931402H11Rik | NA | NA | unclassified non-coding RNA gene | RIKEN cDNA 4931402H11 gene |
| 3 | gene | 88.24287 | 88.25474 | negative | MGI_C57BL6J_1927379 | Rhbg | NCBI_Gene:58176,ENSEMBL:ENSMUSG00000104445 | MGI:1927379 | protein coding gene | Rhesus blood group-associated B glycoprotein |
| 3 | gene | 88.24338 | 88.28721 | negative | MGI_C57BL6J_5613632 | Gm38392 | ENSEMBL:ENSMUSG00000103766 | MGI:5613632 | protein coding gene | predicted gene, 38392 |
| 3 | gene | 88.28275 | 88.29701 | negative | MGI_C57BL6J_1924177 | Tsacc | NCBI_Gene:76927,ENSEMBL:ENSMUSG00000010538 | MGI:1924177 | protein coding gene | TSSK6 activating co-chaperone |
| 3 | pseudogene | 88.28461 | 88.28475 | negative | MGI_C57BL6J_5610344 | Gm37116 | ENSEMBL:ENSMUSG00000103574 | MGI:5610344 | pseudogene | predicted gene, 37116 |
| 3 | gene | 88.29712 | 88.32177 | positive | MGI_C57BL6J_104708 | Cct3 | NCBI_Gene:12462,ENSEMBL:ENSMUSG00000001416 | MGI:104708 | protein coding gene | chaperonin containing Tcp1, subunit 3 (gamma) |
| 3 | gene | 88.32502 | 88.33131 | positive | MGI_C57BL6J_1913318 | Glmp | NCBI_Gene:56700,ENSEMBL:ENSMUSG00000001418 | MGI:1913318 | protein coding gene | glycosylated lysosomal membrane protein |
| 3 | gene | 88.32865 | 88.33615 | negative | MGI_C57BL6J_1919163 | Tmem79 | NCBI_Gene:71913,ENSEMBL:ENSMUSG00000001420 | MGI:1919163 | protein coding gene | transmembrane protein 79 |
| 3 | gene | 88.33474 | 88.33618 | negative | MGI_C57BL6J_3040675 | BC038331 | NA | NA | unclassified gene | cDNA sequence BC038331 |
| 3 | gene | 88.33626 | 88.36234 | positive | MGI_C57BL6J_2447364 | Smg5 | NCBI_Gene:229512,ENSEMBL:ENSMUSG00000001415 | MGI:2447364 | protein coding gene | Smg-5 homolog, nonsense mediated mRNA decay factor (C. elegans) |
| 3 | gene | 88.36407 | 88.36854 | positive | MGI_C57BL6J_1916207 | Paqr6 | NCBI_Gene:68957,ENSEMBL:ENSMUSG00000041423 | MGI:1916207 | protein coding gene | progestin and adipoQ receptor family member VI |
| 3 | gene | 88.36862 | 88.37274 | negative | MGI_C57BL6J_88155 | Bglap3 | NCBI_Gene:12095,ENSEMBL:ENSMUSG00000074489 | MGI:88155 | protein coding gene | bone gamma-carboxyglutamate protein 3 |
| 3 | pseudogene | 88.37581 | 88.38293 | positive | MGI_C57BL6J_5439425 | Gm21956 | NCBI_Gene:101055651,NCBI_Gene:108168878,ENSEMBL:ENSMUSG00000096792 | MGI:5439425 | pseudogene | predicted gene, 21956 |
| 3 | gene | 88.37774 | 88.37870 | negative | MGI_C57BL6J_88157 | Bglap2 | NCBI_Gene:12097,ENSEMBL:ENSMUSG00000074486 | MGI:88157 | protein coding gene | bone gamma-carboxyglutamate protein 2 |
| 3 | pseudogene | 88.38163 | 88.38293 | positive | MGI_C57BL6J_3779634 | Gm6821 | NCBI_Gene:101055651,ENSEMBL:ENSMUSG00000078691 | MGI:3779634 | pseudogene | predicted gene 6821 |
| 3 | gene | 88.38349 | 88.38447 | negative | MGI_C57BL6J_88156 | Bglap | NCBI_Gene:12096,ENSEMBL:ENSMUSG00000074483 | MGI:88156 | protein coding gene | bone gamma carboxyglutamate protein |
| 3 | gene | 88.39414 | 88.41040 | negative | MGI_C57BL6J_1914287 | Pmf1 | NCBI_Gene:67037,ENSEMBL:ENSMUSG00000028066 | MGI:1914287 | protein coding gene | polyamine-modulated factor 1 |
| 3 | gene | 88.41049 | 88.42514 | negative | MGI_C57BL6J_2444391 | Slc25a44 | NCBI_Gene:229517,ENSEMBL:ENSMUSG00000050144 | MGI:2444391 | protein coding gene | solute carrier family 25, member 44 |
| 3 | gene | 88.42508 | 88.42867 | positive | MGI_C57BL6J_5592672 | Gm33513 | NCBI_Gene:102636453 | MGI:5592672 | lncRNA gene | predicted gene, 33513 |
| 3 | pseudogene | 88.42894 | 88.42939 | positive | MGI_C57BL6J_5662951 | Gm42814 | ENSEMBL:ENSMUSG00000106568 | MGI:5662951 | pseudogene | predicted gene 42814 |
| 3 | gene | 88.43596 | 88.46118 | negative | MGI_C57BL6J_107560 | Sema4a | NCBI_Gene:20351,ENSEMBL:ENSMUSG00000028064 | MGI:107560 | protein coding gene | sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| 3 | gene | 88.44088 | 88.44095 | negative | MGI_C57BL6J_5531059 | Mir7011 | miRBase:MI0022860,NCBI_Gene:102465611,ENSEMBL:ENSMUSG00000098655 | MGI:5531059 | miRNA gene | microRNA 7011 |
| 3 | gene | 88.46209 | 88.46613 | negative | MGI_C57BL6J_5622974 | Gm40089 | NCBI_Gene:105244492 | MGI:5622974 | lncRNA gene | predicted gene, 40089 |
| 3 | pseudogene | 88.47454 | 88.47535 | positive | MGI_C57BL6J_3643641 | Gm5979 | NCBI_Gene:546802,ENSEMBL:ENSMUSG00000103304 | MGI:3643641 | pseudogene | predicted gene 5979 |
| 3 | gene | 88.48015 | 88.50996 | negative | MGI_C57BL6J_96794 | Lmna | NCBI_Gene:16905,ENSEMBL:ENSMUSG00000028063 | MGI:96794 | protein coding gene | lamin A |
| 3 | gene | 88.52607 | 88.52736 | negative | MGI_C57BL6J_5610812 | Gm37584 | ENSEMBL:ENSMUSG00000103115 | MGI:5610812 | lncRNA gene | predicted gene, 37584 |
| 3 | gene | 88.52927 | 88.53179 | negative | MGI_C57BL6J_5621503 | Gm38618 | NCBI_Gene:102642612 | MGI:5621503 | lncRNA gene | predicted gene, 38618 |
| 3 | gene | 88.53239 | 88.54140 | positive | MGI_C57BL6J_1919890 | Mex3a | NCBI_Gene:72640,ENSEMBL:ENSMUSG00000074480 | MGI:1919890 | protein coding gene | mex3 RNA binding family member A |
| 3 | gene | 88.53630 | 88.53638 | negative | MGI_C57BL6J_3811408 | Mir1905 | miRBase:MI0008315,NCBI_Gene:100316808,ENSEMBL:ENSMUSG00000084728 | MGI:3811408 | miRNA gene | microRNA 1905 |
| 3 | gene | 88.54203 | 88.54830 | negative | MGI_C57BL6J_1858203 | Rab25 | NCBI_Gene:53868,ENSEMBL:ENSMUSG00000008601 | MGI:1858203 | protein coding gene | RAB25, member RAS oncogene family |
| 3 | gene | 88.54982 | 88.55307 | negative | MGI_C57BL6J_1932697 | Lamtor2 | NCBI_Gene:83409,ENSEMBL:ENSMUSG00000028062 | MGI:1932697 | protein coding gene | late endosomal/lysosomal adaptor, MAPK and MTOR activator 2 |
| 3 | gene | 88.55368 | 88.56973 | positive | MGI_C57BL6J_2150152 | Ubqln4 | NCBI_Gene:94232,ENSEMBL:ENSMUSG00000008604 | MGI:2150152 | protein coding gene | ubiquilin 4 |
| 3 | gene | 88.57588 | 88.58842 | positive | MGI_C57BL6J_1913506 | Ssr2 | NCBI_Gene:66256,ENSEMBL:ENSMUSG00000041355 | MGI:1913506 | protein coding gene | signal sequence receptor, beta |
| 3 | pseudogene | 88.57624 | 88.57720 | negative | MGI_C57BL6J_3642446 | Rtraf-ps | NCBI_Gene:545536,ENSEMBL:ENSMUSG00000074479 | MGI:3642446 | pseudogene | RNA transcription, translation and transport factor, pseudogene |
| 3 | gene | 88.59311 | 88.64953 | positive | MGI_C57BL6J_103264 | Arhgef2 | NCBI_Gene:16800,ENSEMBL:ENSMUSG00000028059 | MGI:103264 | protein coding gene | rho/rac guanine nucleotide exchange factor (GEF) 2 |
| 3 | gene | 88.60667 | 88.60722 | negative | MGI_C57BL6J_5622975 | Gm40090 | NCBI_Gene:105244493 | MGI:5622975 | lncRNA gene | predicted gene, 40090 |
| 3 | gene | 88.61646 | 88.62111 | negative | MGI_C57BL6J_5313099 | Gm20652 | NCBI_Gene:102636610,ENSEMBL:ENSMUSG00000093390 | MGI:5313099 | lncRNA gene | predicted gene 20652 |
| 3 | gene | 88.65190 | 88.65314 | negative | MGI_C57BL6J_2182926 | Rxfp4 | NCBI_Gene:242093,ENSEMBL:ENSMUSG00000049741 | MGI:2182926 | protein coding gene | relaxin family peptide receptor 4 |
| 3 | gene | 88.68502 | 88.68640 | negative | MGI_C57BL6J_5663851 | Gm43714 | ENSEMBL:ENSMUSG00000105512 | MGI:5663851 | lncRNA gene | predicted gene 43714 |
| 3 | gene | 88.68578 | 88.71293 | positive | MGI_C57BL6J_1921450 | Khdc4 | NCBI_Gene:74200,ENSEMBL:ENSMUSG00000028060 | MGI:1921450 | protein coding gene | KH domain containing 4, pre-mRNA splicing factor |
| 3 | gene | 88.69393 | 88.69406 | positive | MGI_C57BL6J_5455597 | Gm25820 | ENSEMBL:ENSMUSG00000105288,ENSEMBL:ENSMUSG00000105448 | MGI:5455597 | snoRNA gene | predicted gene, 25820 |
| 3 | gene | 88.70716 | 88.70729 | positive | MGI_C57BL6J_5455722 | Gm25945 | ENSEMBL:ENSMUSG00000084433 | MGI:5455722 | snoRNA gene | predicted gene, 25945 |
| 3 | gene | 88.71684 | 88.73105 | positive | MGI_C57BL6J_108053 | Rit1 | NCBI_Gene:19769,ENSEMBL:ENSMUSG00000028057 | MGI:108053 | protein coding gene | Ras-like without CAAX 1 |
| 3 | gene | 88.72209 | 88.72267 | positive | MGI_C57BL6J_4937973 | Gm17146 | ENSEMBL:ENSMUSG00000091881 | MGI:4937973 | lncRNA gene | predicted gene 17146 |
| 3 | gene | 88.73875 | 88.73969 | negative | MGI_C57BL6J_3641796 | Gm10253 | ENSEMBL:ENSMUSG00000104333 | MGI:3641796 | unclassified gene | predicted gene 10253 |
| 3 | gene | 88.74470 | 88.77516 | negative | MGI_C57BL6J_1859547 | Syt11 | NCBI_Gene:229521,ENSEMBL:ENSMUSG00000068923 | MGI:1859547 | protein coding gene | synaptotagmin XI |
| 3 | pseudogene | 88.77522 | 88.82937 | positive | MGI_C57BL6J_1923272 | 5830417I10Rik | NCBI_Gene:100302730,ENSEMBL:ENSMUSG00000078684 | MGI:1923272 | pseudogene | RIKEN cDNA 5830417I10 gene |
| 3 | gene | 88.78294 | 88.83255 | negative | MGI_C57BL6J_2442808 | 1500004A13Rik | NCBI_Gene:319830,ENSEMBL:ENSMUSG00000098912 | MGI:2442808 | lncRNA gene | RIKEN cDNA 1500004A13 gene |
| 3 | gene | 88.83097 | 88.91010 | positive | MGI_C57BL6J_1917579 | Gon4l | NCBI_Gene:76022,ENSEMBL:ENSMUSG00000054199 | MGI:1917579 | protein coding gene | gon-4-like (C.elegans) |
| 3 | gene | 88.84503 | 88.90000 | negative | MGI_C57BL6J_5663850 | Gm43713 | ENSEMBL:ENSMUSG00000106218 | MGI:5663850 | lncRNA gene | predicted gene 43713 |
| 3 | gene | 88.86292 | 88.86308 | positive | MGI_C57BL6J_5452831 | Gm23054 | ENSEMBL:ENSMUSG00000064738 | MGI:5452831 | snRNA gene | predicted gene, 23054 |
| 3 | gene | 88.88092 | 88.89994 | negative | MGI_C57BL6J_5622976 | Gm40091 | NCBI_Gene:105244494 | MGI:5622976 | lncRNA gene | predicted gene, 40091 |
| 3 | gene | 88.90511 | 88.91400 | negative | MGI_C57BL6J_2385175 | Msto1 | NCBI_Gene:229524,ENSEMBL:ENSMUSG00000068922 | MGI:2385175 | protein coding gene | misato 1, mitochondrial distribution and morphology regulator |
| 3 | gene | 88.90651 | 88.90663 | positive | MGI_C57BL6J_4422062 | n-R5s197 | ENSEMBL:ENSMUSG00000065236 | MGI:4422062 | rRNA gene | nuclear encoded rRNA 5S 197 |
| 3 | gene | 88.92080 | 88.95120 | negative | MGI_C57BL6J_1929538 | Dap3 | NCBI_Gene:65111,ENSEMBL:ENSMUSG00000068921 | MGI:1929538 | protein coding gene | death associated protein 3 |
| 3 | gene | 88.95049 | 89.07937 | positive | MGI_C57BL6J_2183158 | Ash1l | NCBI_Gene:192195,ENSEMBL:ENSMUSG00000028053 | MGI:2183158 | protein coding gene | ASH1 like histone lysine methyltransferase |
| 3 | gene | 88.97062 | 88.97073 | negative | MGI_C57BL6J_5456242 | Gm26465 | ENSEMBL:ENSMUSG00000095712 | MGI:5456242 | snRNA gene | predicted gene, 26465 |
| 3 | gene | 89.04900 | 89.04913 | negative | MGI_C57BL6J_5454180 | Gm24403 | ENSEMBL:ENSMUSG00000077804 | MGI:5454180 | snoRNA gene | predicted gene, 24403 |
| 3 | gene | 89.08194 | 89.10110 | negative | MGI_C57BL6J_5663875 | Gm43738 | ENSEMBL:ENSMUSG00000105204 | MGI:5663875 | protein coding gene | predicted gene 43738 |
| 3 | gene | 89.08397 | 89.09336 | negative | MGI_C57BL6J_1919546 | Rusc1 | NCBI_Gene:72296,ENSEMBL:ENSMUSG00000041263 | MGI:1919546 | protein coding gene | RUN and SH3 domain containing 1 |
| 3 | gene | 89.09359 | 89.10197 | negative | MGI_C57BL6J_104888 | Fdps | NCBI_Gene:110196,ENSEMBL:ENSMUSG00000059743 | MGI:104888 | protein coding gene | farnesyl diphosphate synthetase |
| 3 | gene | 89.09791 | 89.09801 | negative | MGI_C57BL6J_5452712 | Gm22935 | ENSEMBL:ENSMUSG00000077148 | MGI:5452712 | snRNA gene | predicted gene, 22935 |
| 3 | pseudogene | 89.11669 | 89.11702 | positive | MGI_C57BL6J_5588863 | Gm29704 | NCBI_Gene:101055641,ENSEMBL:ENSMUSG00000105654 | MGI:5588863 | pseudogene | predicted gene, 29704 |
| 3 | gene | 89.13613 | 89.14678 | positive | MGI_C57BL6J_97604 | Pklr | NCBI_Gene:18770,ENSEMBL:ENSMUSG00000041237 | MGI:97604 | protein coding gene | pyruvate kinase liver and red blood cell |
| 3 | gene | 89.14607 | 89.16023 | negative | MGI_C57BL6J_1298211 | Hcn3 | NCBI_Gene:15168,ENSEMBL:ENSMUSG00000028051 | MGI:1298211 | protein coding gene | hyperpolarization-activated, cyclic nucleotide-gated K+ 3 |
| 3 | gene | 89.16479 | 89.17709 | positive | MGI_C57BL6J_1098669 | Clk2 | NCBI_Gene:12748,ENSEMBL:ENSMUSG00000068917 | MGI:1098669 | protein coding gene | CDC-like kinase 2 |
| 3 | gene | 89.16479 | 89.18277 | positive | MGI_C57BL6J_5825564 | Gm45927 | NCBI_Gene:105943584 | MGI:5825564 | protein coding gene | predicted gene, 45927 |
| 3 | gene | 89.17739 | 89.18277 | positive | MGI_C57BL6J_1346346 | Scamp3 | NCBI_Gene:24045,ENSEMBL:ENSMUSG00000028049 | MGI:1346346 | protein coding gene | secretory carrier membrane protein 3 |
| 3 | gene | 89.18076 | 89.18410 | negative | MGI_C57BL6J_3801823 | Gm16069 | NCBI_Gene:102636781,ENSEMBL:ENSMUSG00000086718 | MGI:3801823 | lncRNA gene | predicted gene 16069 |
| 3 | gene | 89.18314 | 89.18930 | positive | MGI_C57BL6J_1915771 | Fam189b | NCBI_Gene:68521,ENSEMBL:ENSMUSG00000032657 | MGI:1915771 | protein coding gene | family with sequence similarity 189, member B |
| 3 | gene | 89.20117 | 89.20127 | negative | MGI_C57BL6J_5453046 | Gm23269 | ENSEMBL:ENSMUSG00000094489 | MGI:5453046 | snRNA gene | predicted gene, 23269 |
| 3 | gene | 89.20291 | 89.20897 | positive | MGI_C57BL6J_95665 | Gba | NCBI_Gene:14466,ENSEMBL:ENSMUSG00000028048 | MGI:95665 | protein coding gene | glucosidase, beta, acid |
| 3 | gene | 89.20908 | 89.22709 | negative | MGI_C57BL6J_103025 | Mtx1 | NCBI_Gene:17827,ENSEMBL:ENSMUSG00000064068 | MGI:103025 | protein coding gene | metaxin 1 |
| 3 | gene | 89.21043 | 89.21316 | positive | MGI_C57BL6J_5663874 | Gm43737 | ENSEMBL:ENSMUSG00000105692 | MGI:5663874 | unclassified gene | predicted gene 43737 |
| 3 | gene | 89.21481 | 89.22684 | positive | MGI_C57BL6J_98739 | Thbs3 | NCBI_Gene:21827,ENSEMBL:ENSMUSG00000028047 | MGI:98739 | protein coding gene | thrombospondin 3 |
| 3 | gene | 89.22712 | 89.22720 | negative | MGI_C57BL6J_3718577 | Mir92b | miRBase:MI0005521,NCBI_Gene:100124470,ENSEMBL:ENSMUSG00000076255 | MGI:3718577 | miRNA gene | microRNA 92b |
| 3 | gene | 89.22906 | 89.23338 | positive | MGI_C57BL6J_97231 | Muc1 | NCBI_Gene:17829,ENSEMBL:ENSMUSG00000042784 | MGI:97231 | protein coding gene | mucin 1, transmembrane |
| 3 | gene | 89.23417 | 89.24631 | negative | MGI_C57BL6J_2673000 | Trim46 | NCBI_Gene:360213,ENSEMBL:ENSMUSG00000042766 | MGI:2673000 | protein coding gene | tripartite motif-containing 46 |
| 3 | gene | 89.24597 | 89.24991 | positive | MGI_C57BL6J_1913309 | Krtcap2 | NCBI_Gene:66059,ENSEMBL:ENSMUSG00000042747 | MGI:1913309 | protein coding gene | keratinocyte associated protein 2 |
| 3 | gene | 89.25926 | 89.26708 | positive | MGI_C57BL6J_1915813 | Dpm3 | NCBI_Gene:68563,ENSEMBL:ENSMUSG00000042737 | MGI:1915813 | protein coding gene | dolichyl-phosphate mannosyltransferase polypeptide 3 |
| 3 | gene | 89.26825 | 89.27057 | negative | MGI_C57BL6J_107417 | Slc50a1 | NCBI_Gene:19729,ENSEMBL:ENSMUSG00000027953 | MGI:107417 | protein coding gene | solute carrier family 50 (sugar transporter), member 1 |
| 3 | gene | 89.27173 | 89.28114 | negative | MGI_C57BL6J_103236 | Efna1 | NCBI_Gene:13636,ENSEMBL:ENSMUSG00000027954 | MGI:103236 | protein coding gene | ephrin A1 |
| 3 | gene | 89.29054 | 89.30290 | negative | MGI_C57BL6J_5593004 | Gm33845 | NCBI_Gene:102636898 | MGI:5593004 | lncRNA gene | predicted gene, 33845 |
| 3 | gene | 89.30248 | 89.30273 | positive | MGI_C57BL6J_5452419 | Gm22642 | ENSEMBL:ENSMUSG00000093093 | MGI:5452419 | unclassified non-coding RNA gene | predicted gene, 22642 |
| 3 | gene | 89.30324 | 89.31422 | positive | MGI_C57BL6J_5593173 | Gm34014 | NCBI_Gene:102637122 | MGI:5593173 | lncRNA gene | predicted gene, 34014 |
| 3 | gene | 89.31390 | 89.32296 | negative | MGI_C57BL6J_106644 | Efna3 | NCBI_Gene:13638,ENSEMBL:ENSMUSG00000028039 | MGI:106644 | protein coding gene | ephrin A3 |
| 3 | gene | 89.31544 | 89.31891 | positive | MGI_C57BL6J_3802094 | Gm15998 | ENSEMBL:ENSMUSG00000086389 | MGI:3802094 | lncRNA gene | predicted gene 15998 |
| 3 | gene | 89.32590 | 89.33117 | negative | MGI_C57BL6J_5593277 | Gm34118 | NCBI_Gene:102637253 | MGI:5593277 | lncRNA gene | predicted gene, 34118 |
| 3 | gene | 89.33339 | 89.33803 | negative | MGI_C57BL6J_106643 | Efna4 | NCBI_Gene:13639,ENSEMBL:ENSMUSG00000028040 | MGI:106643 | protein coding gene | ephrin A4 |
| 3 | gene | 89.33785 | 89.34783 | positive | MGI_C57BL6J_3704222 | 4731419I09Rik | ENSEMBL:ENSMUSG00000097004 | MGI:3704222 | lncRNA gene | RIKEN cDNA 4731419I09 gene |
| 3 | gene | 89.33854 | 89.35003 | negative | MGI_C57BL6J_1333882 | Adam15 | NCBI_Gene:11490,ENSEMBL:ENSMUSG00000028041 | MGI:1333882 | protein coding gene | a disintegrin and metallopeptidase domain 15 (metargidin) |
| 3 | gene | 89.35022 | 89.36525 | negative | MGI_C57BL6J_1925022 | Dcst1 | NCBI_Gene:77772,ENSEMBL:ENSMUSG00000042672 | MGI:1925022 | protein coding gene | DC-STAMP domain containing 1 |
| 3 | gene | 89.35042 | 89.37764 | positive | MGI_C57BL6J_2685606 | Dcst2 | NCBI_Gene:329702,ENSEMBL:ENSMUSG00000109293 | MGI:2685606 | protein coding gene | DC-STAMP domain containing 2 |
| 3 | pseudogene | 89.35363 | 89.35490 | positive | MGI_C57BL6J_3708679 | Gm10702 | ENSEMBL:ENSMUSG00000074467 | MGI:3708679 | pseudogene | predicted gene 10702 |
| 3 | gene | 89.37764 | 89.39481 | negative | MGI_C57BL6J_102755 | Zbtb7b | NCBI_Gene:22724,ENSEMBL:ENSMUSG00000028042 | MGI:102755 | protein coding gene | zinc finger and BTB domain containing 7B |
| 3 | gene | 89.39185 | 89.39878 | positive | MGI_C57BL6J_3642531 | Gm15417 | NCBI_Gene:545539,ENSEMBL:ENSMUSG00000074466 | MGI:3642531 | lncRNA gene | predicted gene 15417 |
| 3 | gene | 89.40099 | 89.40279 | negative | MGI_C57BL6J_1930020 | Lenep | NCBI_Gene:57275,ENSEMBL:ENSMUSG00000078173 | MGI:1930020 | protein coding gene | lens epithelial protein |
| 3 | gene | 89.40100 | 89.41187 | negative | MGI_C57BL6J_2443030 | Flad1 | NCBI_Gene:319945,ENSEMBL:ENSMUSG00000042642 | MGI:2443030 | protein coding gene | flavin adenine dinucleotide synthetase 1 |
| 3 | gene | 89.41547 | 89.41838 | negative | MGI_C57BL6J_1889208 | Cks1b | NCBI_Gene:54124,ENSEMBL:ENSMUSG00000028044 | MGI:1889208 | protein coding gene | CDC28 protein kinase 1b |
| 3 | gene | 89.41840 | 89.43003 | positive | MGI_C57BL6J_98296 | Shc1 | NCBI_Gene:20416,ENSEMBL:ENSMUSG00000042626 | MGI:98296 | protein coding gene | src homology 2 domain-containing transforming protein C1 |
| 3 | gene | 89.43012 | 89.43513 | positive | MGI_C57BL6J_1916161 | Pygo2 | NCBI_Gene:68911,ENSEMBL:ENSMUSG00000047824 | MGI:1916161 | protein coding gene | pygopus 2 |
| 3 | gene | 89.43667 | 89.45095 | positive | MGI_C57BL6J_2441670 | Pbxip1 | NCBI_Gene:229534,ENSEMBL:ENSMUSG00000042613 | MGI:2441670 | protein coding gene | pre B cell leukemia transcription factor interacting protein 1 |
| 3 | gene | 89.45077 | 89.45094 | positive | MGI_C57BL6J_1916805 | 2310033F14Rik | NA | NA | unclassified gene | RIKEN cDNA 2310033F14 gene |
| 3 | gene | 89.45454 | 89.46901 | positive | MGI_C57BL6J_1915853 | Pmvk | NCBI_Gene:68603,ENSEMBL:ENSMUSG00000027952 | MGI:1915853 | protein coding gene | phosphomevalonate kinase |
| 3 | gene | 89.52016 | 89.67513 | positive | MGI_C57BL6J_2153183 | Kcnn3 | NCBI_Gene:140493,ENSEMBL:ENSMUSG00000000794 | MGI:2153183 | protein coding gene | potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3 |
| 3 | gene | 89.71502 | 89.75346 | positive | MGI_C57BL6J_1889575 | Adar | NCBI_Gene:56417,ENSEMBL:ENSMUSG00000027951 | MGI:1889575 | protein coding gene | adenosine deaminase, RNA-specific |
| 3 | gene | 89.74620 | 89.76463 | negative | MGI_C57BL6J_87891 | Chrnb2 | NCBI_Gene:11444,ENSEMBL:ENSMUSG00000027950 | MGI:87891 | protein coding gene | cholinergic receptor, nicotinic, beta polypeptide 2 (neuronal) |
| 3 | gene | 89.76751 | 89.77340 | negative | MGI_C57BL6J_1921284 | 4632404H12Rik | NCBI_Gene:74034,ENSEMBL:ENSMUSG00000042579 | MGI:1921284 | lncRNA gene | RIKEN cDNA 4632404H12 gene |
| 3 | gene | 89.77361 | 89.78400 | positive | MGI_C57BL6J_1917343 | Ube2q1 | NCBI_Gene:70093,ENSEMBL:ENSMUSG00000042572 | MGI:1917343 | protein coding gene | ubiquitin-conjugating enzyme E2Q family member 1 |
| 3 | pseudogene | 89.81948 | 89.82168 | positive | MGI_C57BL6J_3645290 | Gm8969 | NCBI_Gene:668087,ENSEMBL:ENSMUSG00000106564 | MGI:3645290 | pseudogene | predicted gene 8969 |
| 3 | gene | 89.83137 | 89.85885 | positive | MGI_C57BL6J_1099462 | She | NCBI_Gene:214547,ENSEMBL:ENSMUSG00000046280 | MGI:1099462 | protein coding gene | src homology 2 domain-containing transforming protein E |
| 3 | gene | 89.86406 | 89.91320 | negative | MGI_C57BL6J_105304 | Il6ra | NCBI_Gene:16194,ENSEMBL:ENSMUSG00000027947 | MGI:105304 | protein coding gene | interleukin 6 receptor, alpha |
| 3 | gene | 89.87208 | 89.89024 | positive | MGI_C57BL6J_5826448 | Gm46811 | NCBI_Gene:108168879 | MGI:5826448 | lncRNA gene | predicted gene, 46811 |
| 3 | gene | 89.91292 | 89.91405 | positive | MGI_C57BL6J_5662946 | Gm42809 | ENSEMBL:ENSMUSG00000105941 | MGI:5662946 | lncRNA gene | predicted gene 42809 |
| 3 | gene | 89.93948 | 89.96351 | negative | MGI_C57BL6J_1859660 | Atp8b2 | NCBI_Gene:54667,ENSEMBL:ENSMUSG00000060671 | MGI:1859660 | protein coding gene | ATPase, class I, type 8B, member 2 |
| 3 | pseudogene | 89.96344 | 89.96973 | negative | MGI_C57BL6J_4356425 | Aqp10-ps | NCBI_Gene:435743,ENSEMBL:ENSMUSG00000105082 | MGI:4356425 | pseudogene | aquaporin 10, pseudogene |
| 3 | gene | 89.96800 | 89.96816 | negative | MGI_C57BL6J_5453823 | Gm24046 | ENSEMBL:ENSMUSG00000065045 | MGI:5453823 | snRNA gene | predicted gene, 24046 |
| 3 | gene | 89.99545 | 89.99878 | negative | MGI_C57BL6J_1346319 | Hax1 | NCBI_Gene:23897,ENSEMBL:ENSMUSG00000027944 | MGI:1346319 | protein coding gene | HCLS1 associated X-1 |
| 3 | gene | 89.99876 | 90.02144 | positive | MGI_C57BL6J_5011895 | Gm19710 | NCBI_Gene:100503469,ENSEMBL:ENSMUSG00000105552 | MGI:5011895 | lncRNA gene | predicted gene, 19710 |
| 3 | gene | 89.99959 | 90.05263 | negative | MGI_C57BL6J_1921633 | Ubap2l | NCBI_Gene:74383,ENSEMBL:ENSMUSG00000042520 | MGI:1921633 | protein coding gene | ubiquitin-associated protein 2-like |
| 3 | gene | 90.01059 | 90.01073 | negative | MGI_C57BL6J_5454385 | Gm24608 | ENSEMBL:ENSMUSG00000077569 | MGI:5454385 | snoRNA gene | predicted gene, 24608 |
| 3 | gene | 90.01115 | 90.01121 | negative | MGI_C57BL6J_5531378 | Mir7669 | miRBase:MI0025009,NCBI_Gene:102465771,ENSEMBL:ENSMUSG00000098397 | MGI:5531378 | miRNA gene | microRNA 7669 |
| 3 | gene | 90.05164 | 90.06835 | positive | MGI_C57BL6J_1914027 | 4933434E20Rik | NCBI_Gene:99650,ENSEMBL:ENSMUSG00000027942 | MGI:1914027 | protein coding gene | RIKEN cDNA 4933434E20 gene |
| 3 | gene | 90.05726 | 90.05838 | positive | MGI_C57BL6J_4414960 | Gm16540 | ENSEMBL:ENSMUSG00000090005 | MGI:4414960 | lncRNA gene | predicted gene 16540 |
| 3 | gene | 90.06280 | 90.06835 | positive | MGI_C57BL6J_1920795 | 1700094D03Rik | NCBI_Gene:73545 | MGI:1920795 | protein coding gene | RIKEN cDNA 1700094D03 gene |
| 3 | gene | 90.07002 | 90.07010 | positive | MGI_C57BL6J_3718457 | Mir190b | miRBase:MI0005478,NCBI_Gene:100124481,ENSEMBL:ENSMUSG00000077996 | MGI:3718457 | miRNA gene | microRNA 190b |
| 3 | gene | 90.07265 | 90.10090 | positive | MGI_C57BL6J_1890149 | Tpm3 | NCBI_Gene:59069,ENSEMBL:ENSMUSG00000027940 | MGI:1890149 | protein coding gene | tropomyosin 3, gamma |
| 3 | gene | 90.10157 | 90.21215 | positive | MGI_C57BL6J_1924845 | Nup210l | NCBI_Gene:77595,ENSEMBL:ENSMUSG00000027939 | MGI:1924845 | protein coding gene | nucleoporin 210-like |
| 3 | gene | 90.12465 | 90.12712 | positive | MGI_C57BL6J_5594024 | Gm34865 | NCBI_Gene:102638261 | MGI:5594024 | lncRNA gene | predicted gene, 34865 |
| 3 | gene | 90.12654 | 90.12667 | positive | MGI_C57BL6J_5453500 | Gm23723 | ENSEMBL:ENSMUSG00000064930 | MGI:5453500 | snoRNA gene | predicted gene, 23723 |
| 3 | pseudogene | 90.12677 | 90.12706 | positive | MGI_C57BL6J_5663849 | Gm43712 | ENSEMBL:ENSMUSG00000104649 | MGI:5663849 | pseudogene | predicted gene 43712 |
| 3 | gene | 90.19868 | 90.19879 | positive | MGI_C57BL6J_5453962 | Gm24185 | ENSEMBL:ENSMUSG00000084527 | MGI:5453962 | snRNA gene | predicted gene, 24185 |
| 3 | gene | 90.21252 | 90.21365 | negative | MGI_C57BL6J_1888676 | Rps27 | NCBI_Gene:57294,ENSEMBL:ENSMUSG00000090733 | MGI:1888676 | protein coding gene | ribosomal protein S27 |
| 3 | gene | 90.21370 | 90.22639 | positive | MGI_C57BL6J_1927232 | Rab13 | NCBI_Gene:68328,ENSEMBL:ENSMUSG00000027935 | MGI:1927232 | protein coding gene | RAB13, member RAS oncogene family |
| 3 | gene | 90.23160 | 90.23584 | positive | MGI_C57BL6J_1346082 | Jtb | NCBI_Gene:23922,ENSEMBL:ENSMUSG00000027937 | MGI:1346082 | protein coding gene | jumping translocation breakpoint |
| 3 | gene | 90.23750 | 90.24445 | negative | MGI_C57BL6J_1916603 | Creb3l4 | NCBI_Gene:78284,ENSEMBL:ENSMUSG00000027938 | MGI:1916603 | protein coding gene | cAMP responsive element binding protein 3-like 4 |
| 3 | gene | 90.24817 | 90.25361 | positive | MGI_C57BL6J_1353474 | Slc39a1 | NCBI_Gene:30791,ENSEMBL:ENSMUSG00000052310 | MGI:1353474 | protein coding gene | solute carrier family 39 (zinc transporter), member 1 |
| 3 | gene | 90.25386 | 90.26413 | positive | MGI_C57BL6J_1921593 | Crtc2 | NCBI_Gene:74343,ENSEMBL:ENSMUSG00000027936 | MGI:1921593 | protein coding gene | CREB regulated transcription coactivator 2 |
| 3 | gene | 90.26492 | 90.28067 | positive | MGI_C57BL6J_2446201 | Dennd4b | NCBI_Gene:229541,ENSEMBL:ENSMUSG00000042404 | MGI:2446201 | protein coding gene | DENN/MADD domain containing 4B |
| 3 | gene | 90.27015 | 90.27021 | positive | MGI_C57BL6J_5530752 | Mir7012 | miRBase:MI0022861,NCBI_Gene:102465612,ENSEMBL:ENSMUSG00000099199 | MGI:5530752 | miRNA gene | microRNA 7012 |
| 3 | gene | 90.28999 | 90.29073 | positive | MGI_C57BL6J_1918721 | 8430422J01Rik | NA | NA | unclassified gene | RIKEN cDNA 8430422J01 gene |
| 3 | gene | 90.29230 | 90.29398 | negative | MGI_C57BL6J_1923075 | 4930537H20Rik | ENSEMBL:ENSMUSG00000105350 | MGI:1923075 | lncRNA gene | RIKEN cDNA 4930537H20 gene |
| 3 | gene | 90.29318 | 90.36341 | positive | MGI_C57BL6J_2443225 | Gatad2b | NCBI_Gene:229542,ENSEMBL:ENSMUSG00000042390 | MGI:2443225 | protein coding gene | GATA zinc finger domain containing 2B |
| 3 | gene | 90.31152 | 90.31701 | positive | MGI_C57BL6J_5791313 | Gm45477 | ENSEMBL:ENSMUSG00000108394 | MGI:5791313 | unclassified gene | predicted gene 45477 |
| 3 | gene | 90.33540 | 90.33795 | positive | MGI_C57BL6J_5791190 | Gm45354 | ENSEMBL:ENSMUSG00000108689 | MGI:5791190 | unclassified gene | predicted gene 45354 |
| 3 | gene | 90.38523 | 90.39005 | negative | MGI_C57BL6J_1347358 | Slc27a3 | NCBI_Gene:26568,ENSEMBL:ENSMUSG00000027932 | MGI:1347358 | protein coding gene | solute carrier family 27 (fatty acid transporter), member 3 |
| 3 | gene | 90.39138 | 90.43364 | negative | MGI_C57BL6J_2140050 | Ints3 | NCBI_Gene:229543,ENSEMBL:ENSMUSG00000027933 | MGI:2140050 | protein coding gene | integrator complex subunit 3 |
| 3 | gene | 90.45059 | 90.46593 | negative | MGI_C57BL6J_97371 | Npr1 | NCBI_Gene:18160,ENSEMBL:ENSMUSG00000027931 | MGI:97371 | protein coding gene | natriuretic peptide receptor 1 |
| 3 | gene | 90.45158 | 90.45709 | positive | MGI_C57BL6J_3801855 | Gm16048 | ENSEMBL:ENSMUSG00000087411 | MGI:3801855 | lncRNA gene | predicted gene 16048 |
| 3 | pseudogene | 90.46890 | 90.47096 | negative | MGI_C57BL6J_5663732 | Gm43595 | ENSEMBL:ENSMUSG00000105746 | MGI:5663732 | pseudogene | predicted gene 43595 |
| 3 | gene | 90.47586 | 90.47593 | negative | MGI_C57BL6J_4413988 | n-TMcat11 | NCBI_Gene:102467358 | MGI:4413988 | tRNA gene | nuclear encoded tRNA methionine 11 (anticodon CAT) |
| 3 | gene | 90.47613 | 90.48838 | positive | MGI_C57BL6J_1915031 | Ilf2 | NCBI_Gene:67781,ENSEMBL:ENSMUSG00000001016 | MGI:1915031 | protein coding gene | interleukin enhancer binding factor 2 |
| 3 | gene | 90.48803 | 90.49103 | negative | MGI_C57BL6J_1333745 | Snapin | NCBI_Gene:20615,ENSEMBL:ENSMUSG00000001018 | MGI:1333745 | protein coding gene | SNAP-associated protein |
| 3 | gene | 90.49854 | 90.50954 | negative | MGI_C57BL6J_1913761 | Chtop | NCBI_Gene:66511,ENSEMBL:ENSMUSG00000001017 | MGI:1913761 | protein coding gene | chromatin target of PRMT1 |
| 3 | gene | 90.51103 | 90.51439 | negative | MGI_C57BL6J_1338917 | S100a1 | NCBI_Gene:20193,ENSEMBL:ENSMUSG00000044080 | MGI:1338917 | protein coding gene | S100 calcium binding protein A1 |
| 3 | gene | 90.51444 | 90.52458 | positive | MGI_C57BL6J_109581 | S100a13 | NCBI_Gene:20196,ENSEMBL:ENSMUSG00000042312 | MGI:109581 | protein coding gene | S100 calcium binding protein A13 |
| 3 | gene | 90.52685 | 90.52884 | positive | MGI_C57BL6J_1913416 | S100a14 | NCBI_Gene:66166,ENSEMBL:ENSMUSG00000042306 | MGI:1913416 | protein coding gene | S100 calcium binding protein A14 |
| 3 | gene | 90.53725 | 90.54315 | positive | MGI_C57BL6J_1915110 | S100a16 | NCBI_Gene:67860,ENSEMBL:ENSMUSG00000074457 | MGI:1915110 | protein coding gene | S100 calcium binding protein A16 |
| 3 | gene | 90.54862 | 90.54872 | positive | MGI_C57BL6J_5454370 | Gm24593 | ENSEMBL:ENSMUSG00000084456 | MGI:5454370 | snRNA gene | predicted gene, 24593 |
| 3 | gene | 90.56036 | 90.60271 | positive | MGI_C57BL6J_1338849 | S100a3 | NCBI_Gene:20197,ENSEMBL:ENSMUSG00000001021 | MGI:1338849 | protein coding gene | S100 calcium binding protein A3 |
| 3 | gene | 90.56036 | 90.59153 | positive | MGI_C57BL6J_3510999 | S100a2 | NCBI_Gene:628324,ENSEMBL:ENSMUSG00000094018 | MGI:3510999 | protein coding gene | S100 calcium binding protein A2 |
| 3 | gene | 90.56056 | 90.56070 | negative | MGI_C57BL6J_5453241 | Gm23464 | ENSEMBL:ENSMUSG00000065079 | MGI:5453241 | snRNA gene | predicted gene, 23464 |
| 3 | gene | 90.60190 | 90.60589 | positive | MGI_C57BL6J_5662811 | Gm42674 | ENSEMBL:ENSMUSG00000105518 | MGI:5662811 | protein coding gene | predicted gene 42674 |
| 3 | gene | 90.60377 | 90.60604 | positive | MGI_C57BL6J_1330282 | S100a4 | NCBI_Gene:20198,ENSEMBL:ENSMUSG00000001020 | MGI:1330282 | protein coding gene | S100 calcium binding protein A4 |
| 3 | gene | 90.60648 | 90.61178 | positive | MGI_C57BL6J_1338915 | S100a5 | NCBI_Gene:20199,ENSEMBL:ENSMUSG00000001023 | MGI:1338915 | protein coding gene | S100 calcium binding protein A5 |
| 3 | gene | 90.61288 | 90.62418 | positive | MGI_C57BL6J_1339467 | S100a6 | NCBI_Gene:20200,ENSEMBL:ENSMUSG00000001025 | MGI:1339467 | protein coding gene | S100 calcium binding protein A6 (calcyclin) |
| 3 | pseudogene | 90.65165 | 90.65187 | negative | MGI_C57BL6J_5011924 | Gm19739 | NCBI_Gene:102638335,ENSEMBL:ENSMUSG00000105644 | MGI:5011924 | pseudogene | predicted gene, 19739 |
| 3 | gene | 90.65183 | 90.65867 | positive | MGI_C57BL6J_2687194 | S100a7a | NCBI_Gene:381493,ENSEMBL:ENSMUSG00000063767 | MGI:2687194 | protein coding gene | S100 calcium binding protein A7A |
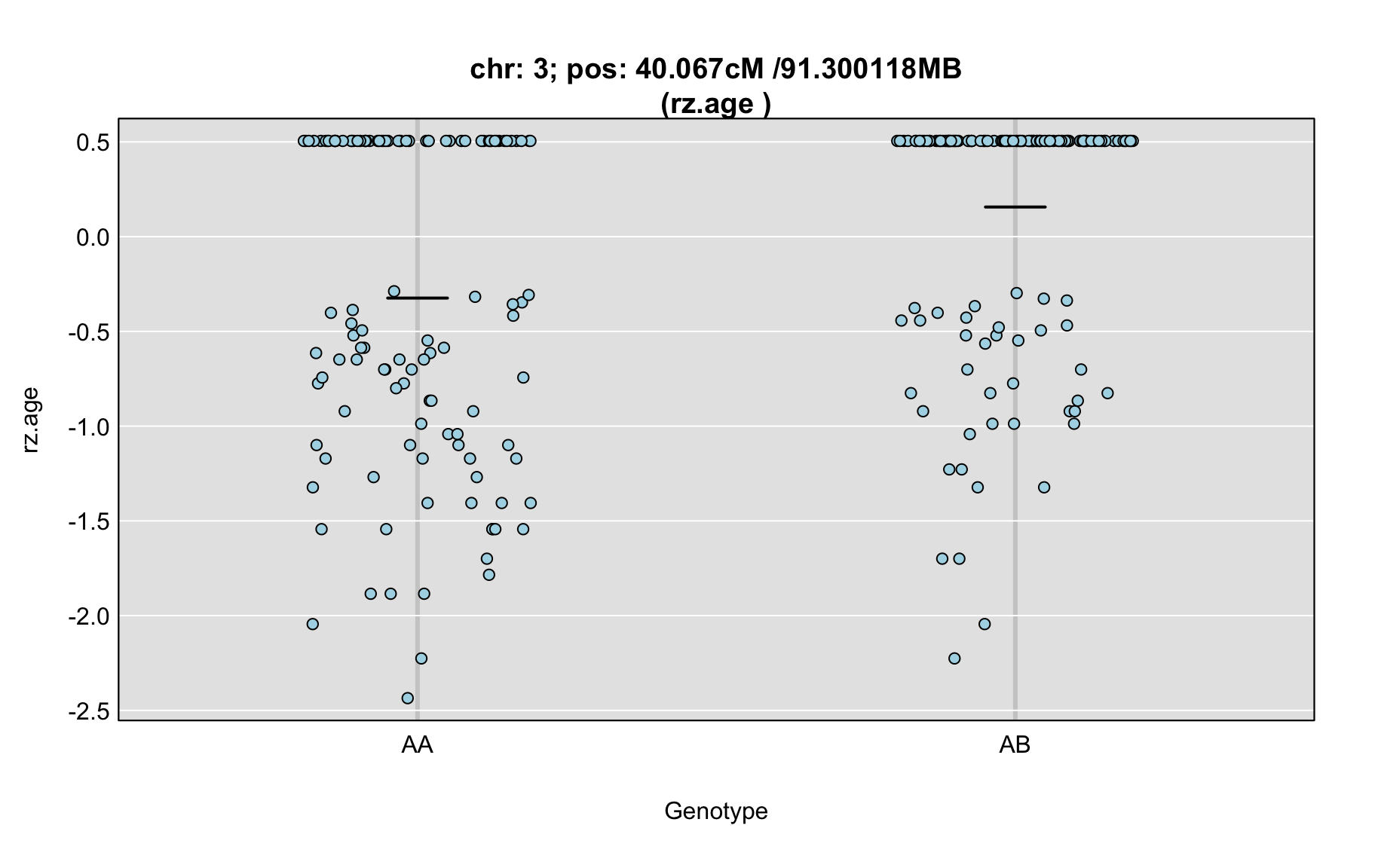
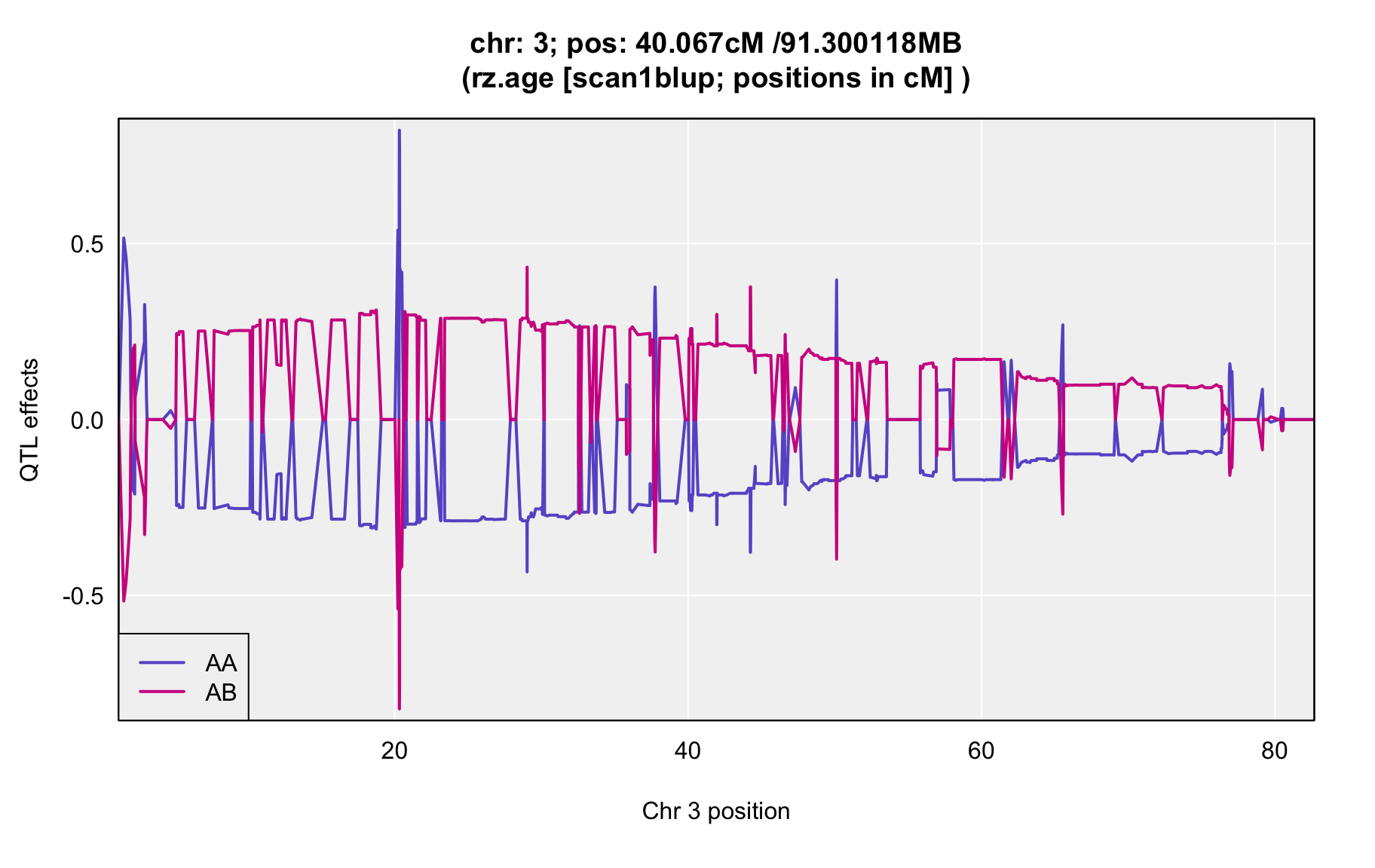
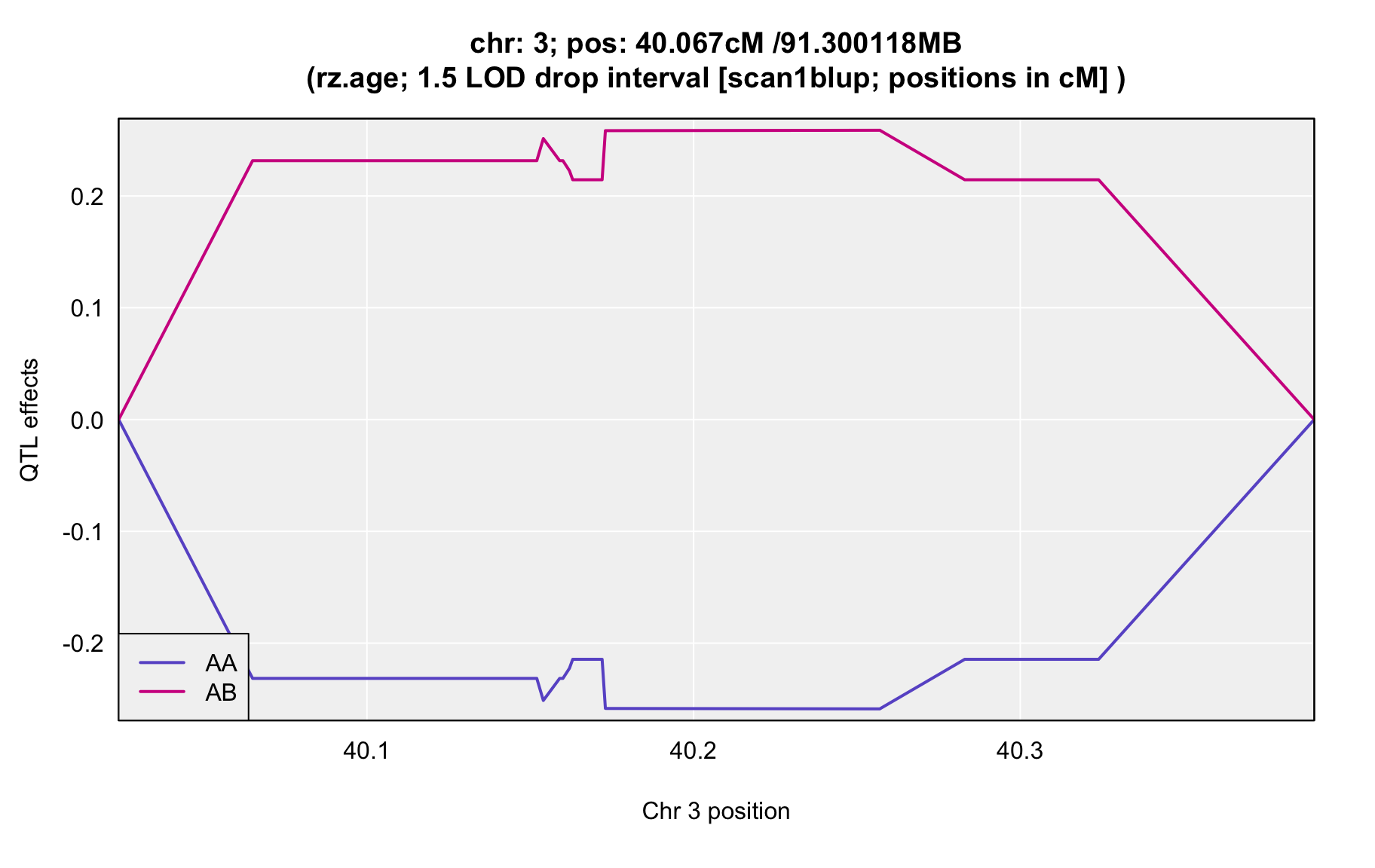
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | pseudogene | 90.84637 | 90.84825 | negative | MGI_C57BL6J_5663372 | Gm43235 | ENSEMBL:ENSMUSG00000106174 | MGI:5663372 | pseudogene | predicted gene 43235 |
| 3 | pseudogene | 91.00976 | 91.01718 | negative | MGI_C57BL6J_3643784 | Gm9023 | NCBI_Gene:668179,ENSEMBL:ENSMUSG00000106257 | MGI:3643784 | pseudogene | predicted gene 9023 |
| 3 | pseudogene | 91.03957 | 91.03987 | positive | MGI_C57BL6J_3779688 | Gm7166 | ENSEMBL:ENSMUSG00000105919 | MGI:3779688 | pseudogene | predicted gene 7166 |
| 3 | pseudogene | 91.07166 | 91.07519 | positive | MGI_C57BL6J_1922392 | 4930529C04Rik | NCBI_Gene:619677,ENSEMBL:ENSMUSG00000086377 | MGI:1922392 | pseudogene | RIKEN cDNA 4930529C04 gene |
| 3 | gene | 91.08814 | 91.10743 | negative | MGI_C57BL6J_2684973 | S100a7l2 | NCBI_Gene:229550,ENSEMBL:ENSMUSG00000091175 | MGI:2684973 | protein coding gene | S100 calcium binding protein A7 like 2 |
| 3 | gene | 91.62856 | 91.63070 | negative | MGI_C57BL6J_5595482 | Gm36323 | NCBI_Gene:102640196 | MGI:5595482 | protein coding gene | predicted gene, 36323 |
| 3 | gene | 91.73835 | 91.74049 | negative | MGI_C57BL6J_5594344 | Gm35185 | NCBI_Gene:102638680 | MGI:5594344 | protein coding gene | predicted gene, 35185 |
| 3 | pseudogene | 91.91438 | 91.91653 | positive | MGI_C57BL6J_5826474 | Gm46837 | NCBI_Gene:108168919 | MGI:5826474 | pseudogene | predicted gene, 46837 |
| 3 | pseudogene | 91.99611 | 91.99725 | positive | MGI_C57BL6J_5009818 | Gm17742 | NCBI_Gene:383887 | MGI:5009818 | pseudogene | predicted gene, 17742 |
| 3 | gene | 92.01076 | 92.01086 | negative | MGI_C57BL6J_5453474 | Gm23697 | ENSEMBL:ENSMUSG00000077085 | MGI:5453474 | miRNA gene | predicted gene, 23697 |
| 3 | gene | 92.01248 | 92.03267 | positive | MGI_C57BL6J_2685266 | Pglyrp3 | NCBI_Gene:242100,ENSEMBL:ENSMUSG00000042244 | MGI:2685266 | protein coding gene | peptidoglycan recognition protein 3 |
| 3 | pseudogene | 92.02449 | 92.02503 | positive | MGI_C57BL6J_103164 | Mif-ps3 | NCBI_Gene:619750,ENSEMBL:ENSMUSG00000082826 | MGI:103164 | pseudogene | macrophage migration inhibitory factor, pseudogene 3 |
| 3 | gene | 92.06140 | 92.06151 | negative | MGI_C57BL6J_5454922 | Gm25145 | ENSEMBL:ENSMUSG00000096133 | MGI:5454922 | miRNA gene | predicted gene, 25145 |
| 3 | gene | 92.08027 | 92.08314 | negative | MGI_C57BL6J_96816 | Lor | NCBI_Gene:16939,ENSEMBL:ENSMUSG00000043165 | MGI:96816 | protein coding gene | loricrin |
| 3 | gene | 92.09735 | 92.09963 | positive | MGI_C57BL6J_5663873 | Gm43736 | ENSEMBL:ENSMUSG00000104981 | MGI:5663873 | lncRNA gene | predicted gene 43736 |
| 3 | gene | 92.10258 | 92.14274 | negative | MGI_C57BL6J_1922313 | 4930511M18Rik | ENSEMBL:ENSMUSG00000104791 | MGI:1922313 | lncRNA gene | RIKEN cDNA 4930511M18 gene |
| 3 | gene | 92.11319 | 92.11932 | positive | MGI_C57BL6J_5663872 | Gm43735 | ENSEMBL:ENSMUSG00000106143 | MGI:5663872 | lncRNA gene | predicted gene 43735 |
| 3 | gene | 92.12220 | 92.12395 | negative | MGI_C57BL6J_1925680 | Prr9 | NCBI_Gene:109314,ENSEMBL:ENSMUSG00000056270 | MGI:1925680 | protein coding gene | proline rich 9 |
| 3 | gene | 92.13499 | 92.14275 | negative | MGI_C57BL6J_1916582 | Lelp1 | NCBI_Gene:69332,ENSEMBL:ENSMUSG00000027927 | MGI:1916582 | protein coding gene | late cornified envelope-like proline-rich 1 |
| 3 | gene | 92.17019 | 92.18323 | positive | MGI_C57BL6J_5594598 | Gm35439 | NCBI_Gene:102639022,ENSEMBL:ENSMUSG00000105476 | MGI:5594598 | lncRNA gene | predicted gene, 35439 |
| 3 | gene | 92.21580 | 92.22249 | positive | MGI_C57BL6J_1330350 | Sprr2a1 | NCBI_Gene:20755,ENSEMBL:ENSMUSG00000078664 | MGI:1330350 | protein coding gene | small proline-rich protein 2A1 |
| 3 | gene | 92.21592 | 92.25730 | positive | MGI_C57BL6J_3845026 | Sprr2a2 | NCBI_Gene:100303744,ENSEMBL:ENSMUSG00000068893 | MGI:3845026 | protein coding gene | small proline-rich protein 2A2 |
| 3 | pseudogene | 92.22741 | 92.22870 | negative | MGI_C57BL6J_3645657 | Gm5850 | NCBI_Gene:545543,ENSEMBL:ENSMUSG00000103286 | MGI:3645657 | pseudogene | predicted gene 5850 |
| 3 | gene | 92.24293 | 92.24624 | negative | MGI_C57BL6J_5610315 | Gm37087 | ENSEMBL:ENSMUSG00000103187 | MGI:5610315 | unclassified gene | predicted gene, 37087 |
| 3 | pseudogene | 92.26222 | 92.26350 | negative | MGI_C57BL6J_3645652 | Gm5851 | NCBI_Gene:545545,ENSEMBL:ENSMUSG00000104028 | MGI:3645652 | pseudogene | predicted gene 5851 |
| 3 | gene | 92.27773 | 92.28103 | negative | MGI_C57BL6J_5610980 | Gm37752 | ENSEMBL:ENSMUSG00000103385 | MGI:5610980 | unclassified gene | predicted gene, 37752 |
| 3 | gene | 92.28530 | 92.29140 | positive | MGI_C57BL6J_3845028 | Sprr2a3 | NCBI_Gene:100042514,ENSEMBL:ENSMUSG00000074445 | MGI:3845028 | protein coding gene | small proline-rich protein 2A3 |
| 3 | gene | 92.31662 | 92.31808 | positive | MGI_C57BL6J_1330352 | Sprr2b | NCBI_Gene:20756,ENSEMBL:ENSMUSG00000050092 | MGI:1330352 | protein coding gene | small proline-rich protein 2B |
| 3 | pseudogene | 92.32091 | 92.32105 | positive | MGI_C57BL6J_5610656 | Gm37428 | ENSEMBL:ENSMUSG00000103518 | MGI:5610656 | pseudogene | predicted gene, 37428 |
| 3 | pseudogene | 92.32660 | 92.32690 | positive | MGI_C57BL6J_1330351 | Sprr2c-ps | NCBI_Gene:20757,ENSEMBL:ENSMUSG00000103221 | MGI:1330351 | pseudogene | small proline-rich protein 2C, pseudogene |
| 3 | gene | 92.33914 | 92.34087 | positive | MGI_C57BL6J_1330347 | Sprr2d | NCBI_Gene:20758,ENSEMBL:ENSMUSG00000042212 | MGI:1330347 | protein coding gene | small proline-rich protein 2D |
| 3 | gene | 92.35214 | 92.35345 | positive | MGI_C57BL6J_1330346 | Sprr2e | NCBI_Gene:20759,ENSEMBL:ENSMUSG00000055030 | MGI:1330346 | protein coding gene | small proline-rich protein 2E |
| 3 | gene | 92.36519 | 92.36644 | positive | MGI_C57BL6J_1330349 | Sprr2f | NCBI_Gene:20760,ENSEMBL:ENSMUSG00000050635 | MGI:1330349 | protein coding gene | small proline-rich protein 2F |
| 3 | pseudogene | 92.37391 | 92.37523 | positive | MGI_C57BL6J_1330348 | Sprr2g | NCBI_Gene:20761,ENSEMBL:ENSMUSG00000046203 | MGI:1330348 | polymorphic pseudogene | small proline-rich protein 2G |
| 3 | gene | 92.38568 | 92.38732 | positive | MGI_C57BL6J_1330343 | Sprr2h | NCBI_Gene:20762,ENSEMBL:ENSMUSG00000046259 | MGI:1330343 | protein coding gene | small proline-rich protein 2H |
| 3 | gene | 92.40799 | 92.40927 | positive | MGI_C57BL6J_1330309 | Sprr2i | NCBI_Gene:20763,ENSEMBL:ENSMUSG00000042157 | MGI:1330309 | protein coding gene | small proline-rich protein 2I |
| 3 | pseudogene | 92.41808 | 92.41940 | positive | MGI_C57BL6J_1330345 | Sprr2j-ps | NCBI_Gene:20764,ENSEMBL:ENSMUSG00000027925 | MGI:1330345 | pseudogene | small proline-rich protein 2J, pseudogene |
| 3 | gene | 92.42803 | 92.42942 | negative | MGI_C57BL6J_3642386 | Gm9774 | NCBI_Gene:100043022,ENSEMBL:ENSMUSG00000042165 | MGI:3642386 | protein coding gene | predicted pseudogene 9774 |
| 3 | gene | 92.43258 | 92.43393 | positive | MGI_C57BL6J_1330344 | Sprr2k | NCBI_Gene:20765,ENSEMBL:ENSMUSG00000054215 | MGI:1330344 | protein coding gene | small proline-rich protein 2K |
| 3 | gene | 92.43681 | 92.43882 | negative | MGI_C57BL6J_106659 | Sprr1b | NCBI_Gene:20754,ENSEMBL:ENSMUSG00000048455 | MGI:106659 | protein coding gene | small proline-rich protein 1B |
| 3 | gene | 92.45650 | 92.45872 | negative | MGI_C57BL6J_1330237 | Sprr3 | NCBI_Gene:20766,ENSEMBL:ENSMUSG00000045539 | MGI:1330237 | protein coding gene | small proline-rich protein 3 |
| 3 | pseudogene | 92.47324 | 92.47354 | negative | MGI_C57BL6J_5610605 | Gm37377 | ENSEMBL:ENSMUSG00000102345 | MGI:5610605 | pseudogene | predicted gene, 37377 |
| 3 | gene | 92.48395 | 92.48589 | negative | MGI_C57BL6J_106660 | Sprr1a | NCBI_Gene:20753,ENSEMBL:ENSMUSG00000050359 | MGI:106660 | protein coding gene | small proline-rich protein 1A |
| 3 | gene | 92.50026 | 92.50049 | negative | MGI_C57BL6J_2654508 | Sprr4 | NCBI_Gene:229562,ENSEMBL:ENSMUSG00000045566 | MGI:2654508 | protein coding gene | small proline-rich protein 4 |
| 3 | gene | 92.53251 | 92.53451 | negative | MGI_C57BL6J_1924218 | 2310046K23Rik | ENSEMBL:ENSMUSG00000102308 | MGI:1924218 | protein coding gene | RIKEN cDNA 2310046K23 gene |
| 3 | gene | 92.57090 | 92.57379 | negative | MGI_C57BL6J_96626 | Ivl | NCBI_Gene:16447,ENSEMBL:ENSMUSG00000049128 | MGI:96626 | protein coding gene | involucrin |
| 3 | gene | 92.57387 | 92.62033 | positive | MGI_C57BL6J_5594798 | Gm35639 | NCBI_Gene:102639293 | MGI:5594798 | lncRNA gene | predicted gene, 35639 |
| 3 | gene | 92.58387 | 92.58902 | negative | MGI_C57BL6J_96945 | Smcp | NCBI_Gene:17235,ENSEMBL:ENSMUSG00000074435 | MGI:96945 | protein coding gene | sperm mitochondria-associated cysteine-rich protein |
| 3 | gene | 92.60472 | 92.60749 | negative | MGI_C57BL6J_3588251 | 2210017I01Rik | ENSEMBL:ENSMUSG00000103523 | MGI:3588251 | protein coding gene | RIKEN cDNA 2210017I01 gene |
| 3 | gene | 92.60476 | 92.62166 | negative | MGI_C57BL6J_1925632 | Lce6a | NCBI_Gene:78382,ENSEMBL:ENSMUSG00000086848 | MGI:1925632 | protein coding gene | late cornified envelope 6A |
| 3 | gene | 92.62034 | 92.64644 | positive | MGI_C57BL6J_5826449 | Gm46812 | NCBI_Gene:108168880 | MGI:5826449 | lncRNA gene | predicted gene, 46812 |
| 3 | gene | 92.63906 | 92.64007 | negative | MGI_C57BL6J_5594858 | Gm35699 | NCBI_Gene:102639369 | MGI:5594858 | lncRNA gene | predicted gene, 35699 |
| 3 | gene | 92.64653 | 92.64831 | negative | MGI_C57BL6J_1914377 | Lce1a1 | NCBI_Gene:67127,ENSEMBL:ENSMUSG00000057609 | MGI:1914377 | protein coding gene | late cornified envelope 1A1 |
| 3 | gene | 92.65565 | 92.65693 | negative | MGI_C57BL6J_1915970 | Lce1b | NCBI_Gene:68720,ENSEMBL:ENSMUSG00000027923 | MGI:1915970 | protein coding gene | late cornified envelope 1B |
| 3 | pseudogene | 92.66419 | 92.66490 | positive | MGI_C57BL6J_3648077 | Rpl13-ps4 | NCBI_Gene:620325,ENSEMBL:ENSMUSG00000104215 | MGI:3648077 | pseudogene | ribosomal protein L13, pseudogene 4 |
| 3 | gene | 92.66861 | 92.67032 | negative | MGI_C57BL6J_1920972 | Lce1a2 | NCBI_Gene:73722,ENSEMBL:ENSMUSG00000068890 | MGI:1920972 | protein coding gene | late cornified envelope 1A2 |
| 3 | gene | 92.67925 | 92.68092 | positive | MGI_C57BL6J_1920969 | Lce1c | NCBI_Gene:73719,ENSEMBL:ENSMUSG00000042092 | MGI:1920969 | protein coding gene | late cornified envelope 1C |
| 3 | gene | 92.68550 | 92.68721 | negative | MGI_C57BL6J_1916861 | Lce1d | NCBI_Gene:69611,ENSEMBL:ENSMUSG00000103243 | MGI:1916861 | protein coding gene | late cornified envelope 1D |
| 3 | pseudogene | 92.68942 | 92.69016 | positive | MGI_C57BL6J_3782379 | Gm4202 | NCBI_Gene:100418460,ENSEMBL:ENSMUSG00000091421 | MGI:3782379 | pseudogene | predicted gene 4202 |
| 3 | gene | 92.70740 | 92.70907 | negative | MGI_C57BL6J_1915944 | Lce1e | NCBI_Gene:68694,ENSEMBL:ENSMUSG00000068889 | MGI:1915944 | protein coding gene | late cornified envelope 1E |
| 3 | gene | 92.71868 | 92.72035 | negative | MGI_C57BL6J_1915078 | Lce1f | NCBI_Gene:67828,ENSEMBL:ENSMUSG00000042124 | MGI:1915078 | protein coding gene | late cornified envelope 1F |
| 3 | gene | 92.73765 | 92.73910 | negative | MGI_C57BL6J_5611347 | Gm38119 | ENSEMBL:ENSMUSG00000103084 | MGI:5611347 | protein coding gene | predicted gene, 38119 |
| 3 | gene | 92.75015 | 92.75234 | negative | MGI_C57BL6J_1913445 | Lce1g | NCBI_Gene:66195,ENSEMBL:ENSMUSG00000027919 | MGI:1913445 | protein coding gene | late cornified envelope 1G |
| 3 | gene | 92.76322 | 92.76507 | negative | MGI_C57BL6J_1914968 | Lce1h | NCBI_Gene:67718,ENSEMBL:ENSMUSG00000049593 | MGI:1914968 | protein coding gene | late cornified envelope 1H |
| 3 | gene | 92.77721 | 92.77890 | negative | MGI_C57BL6J_1923835 | Lce1i | NCBI_Gene:76585,ENSEMBL:ENSMUSG00000068888 | MGI:1923835 | protein coding gene | late cornified envelope 1I |
| 3 | gene | 92.78884 | 92.79051 | negative | MGI_C57BL6J_3702532 | Lce1j | NCBI_Gene:545547,ENSEMBL:ENSMUSG00000068887 | MGI:3702532 | protein coding gene | late cornified envelope 1J |
| 3 | gene | 92.80629 | 92.80789 | negative | MGI_C57BL6J_3702534 | Lce1k | NCBI_Gene:631101,ENSEMBL:ENSMUSG00000095870 | MGI:3702534 | protein coding gene | late cornified envelope 1K |
| 3 | gene | 92.82307 | 92.82725 | negative | MGI_C57BL6J_1920981 | Kprp | NCBI_Gene:433619,ENSEMBL:ENSMUSG00000059832 | MGI:1920981 | protein coding gene | keratinocyte expressed, proline-rich |
| 3 | gene | 92.84330 | 92.84368 | negative | MGI_C57BL6J_5611429 | Gm38201 | ENSEMBL:ENSMUSG00000102469 | MGI:5611429 | lncRNA gene | predicted gene, 38201 |
| 3 | gene | 92.84995 | 92.85129 | negative | MGI_C57BL6J_1920980 | Lce1l | NCBI_Gene:73730,ENSEMBL:ENSMUSG00000046676 | MGI:1920980 | protein coding gene | late cornified envelope 1L |
| 3 | gene | 92.86836 | 92.87022 | negative | MGI_C57BL6J_1913783 | 2310050C09Rik | NCBI_Gene:66533,ENSEMBL:ENSMUSG00000090314 | MGI:1913783 | protein coding gene | RIKEN cDNA 2310050C09 gene |
| 3 | gene | 92.92529 | 92.92623 | negative | MGI_C57BL6J_3645650 | Lce3a | NCBI_Gene:545548,ENSEMBL:ENSMUSG00000054325 | MGI:3645650 | protein coding gene | late cornified envelope 3A |
| 3 | gene | 92.93298 | 92.93410 | positive | MGI_C57BL6J_1913594 | Lce3b | NCBI_Gene:66344,ENSEMBL:ENSMUSG00000042031 | MGI:1913594 | protein coding gene | late cornified envelope 3B |
| 3 | pseudogene | 92.93768 | 92.93807 | positive | MGI_C57BL6J_5611306 | Gm38078 | ENSEMBL:ENSMUSG00000104364 | MGI:5611306 | pseudogene | predicted gene, 38078 |
| 3 | gene | 92.94449 | 92.94573 | positive | MGI_C57BL6J_2135932 | Lce3c | NCBI_Gene:94060,ENSEMBL:ENSMUSG00000045475 | MGI:2135932 | protein coding gene | late cornified envelope 3C |
| 3 | gene | 92.94916 | 92.94926 | negative | MGI_C57BL6J_5454258 | Gm24481 | ENSEMBL:ENSMUSG00000087835 | MGI:5454258 | snRNA gene | predicted gene, 24481 |
| 3 | gene | 92.95739 | 92.95857 | positive | MGI_C57BL6J_3642919 | Lce3d | NCBI_Gene:630994,ENSEMBL:ENSMUSG00000043472 | MGI:3642919 | protein coding gene | late cornified envelope 3D |
| 3 | gene | 92.96201 | 92.96212 | negative | MGI_C57BL6J_5452354 | Gm22577 | ENSEMBL:ENSMUSG00000065665 | MGI:5452354 | snRNA gene | predicted gene, 22577 |
| 3 | gene | 92.96706 | 92.96828 | positive | MGI_C57BL6J_1916764 | Lce3e | NCBI_Gene:69514,ENSEMBL:ENSMUSG00000074433 | MGI:1916764 | protein coding gene | late cornified envelope 3E |
| 3 | gene | 92.99223 | 92.99343 | positive | MGI_C57BL6J_1916770 | Lce3f | NCBI_Gene:69520,ENSEMBL:ENSMUSG00000068885 | MGI:1916770 | protein coding gene | late cornified envelope 3F |
| 3 | gene | 93.01421 | 93.01569 | negative | MGI_C57BL6J_1921425 | Crct1 | NCBI_Gene:74175,ENSEMBL:ENSMUSG00000027913 | MGI:1921425 | protein coding gene | cysteine-rich C-terminal 1 |
| 3 | gene | 93.01781 | 93.01906 | negative | MGI_C57BL6J_1913453 | Lce1m | NCBI_Gene:66203,ENSEMBL:ENSMUSG00000027912 | MGI:1913453 | protein coding gene | late cornified envelope 1M |
| 3 | pseudogene | 93.06474 | 93.06496 | negative | MGI_C57BL6J_5611268 | Gm38040 | ENSEMBL:ENSMUSG00000104298 | MGI:5611268 | pseudogene | predicted gene, 38040 |
| 3 | gene | 93.06882 | 93.07551 | positive | MGI_C57BL6J_3645677 | Tdpoz8 | NCBI_Gene:229571,ENSEMBL:ENSMUSG00000096879 | MGI:3645677 | protein coding gene | TD and POZ domain containing 8 |
| 3 | gene | 93.09758 | 93.12825 | negative | MGI_C57BL6J_5594944 | Gm35785 | NCBI_Gene:102639481 | MGI:5594944 | lncRNA gene | predicted gene, 35785 |
| 3 | pseudogene | 93.10044 | 93.10060 | negative | MGI_C57BL6J_5610589 | Gm37361 | NCBI_Gene:108168920,ENSEMBL:ENSMUSG00000103628 | MGI:5610589 | pseudogene | predicted gene, 37361 |
| 3 | gene | 93.14479 | 93.14982 | positive | MGI_C57BL6J_2685861 | Crnn | NCBI_Gene:381457,ENSEMBL:ENSMUSG00000078657 | MGI:2685861 | protein coding gene | cornulin |
| 3 | pseudogene | 93.15218 | 93.15287 | positive | MGI_C57BL6J_5610729 | Gm37501 | ENSEMBL:ENSMUSG00000103902 | MGI:5610729 | pseudogene | predicted gene, 37501 |
| 3 | gene | 93.19727 | 93.22139 | positive | MGI_C57BL6J_3645678 | Flg2 | NCBI_Gene:229574,ENSEMBL:ENSMUSG00000049133 | MGI:3645678 | protein coding gene | filaggrin family member 2 |
| 3 | pseudogene | 93.23492 | 93.23647 | positive | MGI_C57BL6J_5010617 | Gm18432 | NCBI_Gene:100417160,ENSEMBL:ENSMUSG00000102609 | MGI:5010617 | pseudogene | predicted gene, 18432 |
| 3 | gene | 93.27352 | 93.29369 | positive | MGI_C57BL6J_95553 | Flg | NCBI_Gene:14246,ENSEMBL:ENSMUSG00000102439 | MGI:95553 | protein coding gene | filaggrin |
| 3 | pseudogene | 93.27807 | 93.27915 | positive | MGI_C57BL6J_3648599 | Rplp0-ps1 | NCBI_Gene:668300,ENSEMBL:ENSMUSG00000081640 | MGI:3648599 | pseudogene | ribosomal protein, large, P0, pseudogene 1 |
| 3 | pseudogene | 93.30501 | 93.30547 | positive | MGI_C57BL6J_5611150 | Gm37922 | ENSEMBL:ENSMUSG00000102485 | MGI:5611150 | pseudogene | predicted gene, 37922 |
| 3 | gene | 93.30647 | 93.30901 | positive | MGI_C57BL6J_5611141 | Gm37913 | ENSEMBL:ENSMUSG00000103273 | MGI:5611141 | unclassified gene | predicted gene, 37913 |
| 3 | gene | 93.30647 | 93.46536 | negative | MGI_C57BL6J_5610473 | Gm37245 | ENSEMBL:ENSMUSG00000102476 | MGI:5610473 | lncRNA gene | predicted gene, 37245 |
| 3 | gene | 93.31954 | 93.33357 | positive | MGI_C57BL6J_3046938 | Hrnr | NCBI_Gene:68723,ENSEMBL:ENSMUSG00000041991 | MGI:3046938 | protein coding gene | hornerin |
| 3 | gene | 93.38869 | 93.38883 | positive | MGI_C57BL6J_5690745 | Gm44353 | ENSEMBL:ENSMUSG00000106242 | MGI:5690745 | miRNA gene | predicted gene, 44353 |
| 3 | gene | 93.39370 | 93.39490 | positive | MGI_C57BL6J_5611085 | Gm37857 | ENSEMBL:ENSMUSG00000103765 | MGI:5611085 | unclassified gene | predicted gene, 37857 |
| 3 | gene | 93.39370 | 93.39944 | positive | MGI_C57BL6J_1099055 | Rptn | NCBI_Gene:20129,ENSEMBL:ENSMUSG00000041984 | MGI:1099055 | protein coding gene | repetin |
| 3 | pseudogene | 93.42065 | 93.42143 | negative | MGI_C57BL6J_5010481 | Gm18296 | NCBI_Gene:100416867,ENSEMBL:ENSMUSG00000104171 | MGI:5010481 | pseudogene | predicted gene, 18296 |
| 3 | pseudogene | 93.43306 | 93.43396 | negative | MGI_C57BL6J_3643701 | Gm5278 | NCBI_Gene:383891,ENSEMBL:ENSMUSG00000057829 | MGI:3643701 | pseudogene | predicted pseudogene 5278 |
| 3 | gene | 93.43320 | 93.43331 | negative | MGI_C57BL6J_5452529 | Gm22752 | ENSEMBL:ENSMUSG00000076037 | MGI:5452529 | miRNA gene | predicted gene, 22752 |
| 3 | gene | 93.44233 | 93.44908 | positive | MGI_C57BL6J_2177944 | Tchh | NCBI_Gene:99681,ENSEMBL:ENSMUSG00000052415 | MGI:2177944 | protein coding gene | trichohyalin |
| 3 | gene | 93.46875 | 93.47198 | positive | MGI_C57BL6J_1918575 | Tchhl1 | NCBI_Gene:71325,ENSEMBL:ENSMUSG00000027908 | MGI:1918575 | protein coding gene | trichohyalin-like 1 |
| 3 | pseudogene | 93.49981 | 93.50030 | negative | MGI_C57BL6J_5610181 | Gm36953 | ENSEMBL:ENSMUSG00000104135 | MGI:5610181 | pseudogene | predicted gene, 36953 |
| 3 | gene | 93.52049 | 93.52629 | positive | MGI_C57BL6J_1338798 | S100a11 | NCBI_Gene:20195,ENSEMBL:ENSMUSG00000027907 | MGI:1338798 | protein coding gene | S100 calcium binding protein A11 |
| 3 | pseudogene | 93.53051 | 93.53079 | negative | MGI_C57BL6J_5589109 | Gm29950 | NCBI_Gene:102631667,ENSEMBL:ENSMUSG00000103031 | MGI:5589109 | pseudogene | predicted gene, 29950 |
| 3 | gene | 93.55507 | 93.56464 | positive | MGI_C57BL6J_1339468 | S100a10 | NCBI_Gene:20194,ENSEMBL:ENSMUSG00000041959 | MGI:1339468 | protein coding gene | S100 calcium binding protein A10 (calpactin) |
| 3 | gene | 93.59536 | 93.59750 | negative | MGI_C57BL6J_5622978 | Gm40093 | NCBI_Gene:105244496 | MGI:5622978 | lncRNA gene | predicted gene, 40093 |
| 3 | pseudogene | 93.60517 | 93.60571 | positive | MGI_C57BL6J_5010516 | Gm18331 | NCBI_Gene:100416948 | MGI:5010516 | pseudogene | predicted gene, 18331 |
| 3 | pseudogene | 93.62864 | 93.62975 | negative | MGI_C57BL6J_3645005 | Gm5541 | NCBI_Gene:433623,ENSEMBL:ENSMUSG00000103406 | MGI:3645005 | pseudogene | predicted gene 5541 |
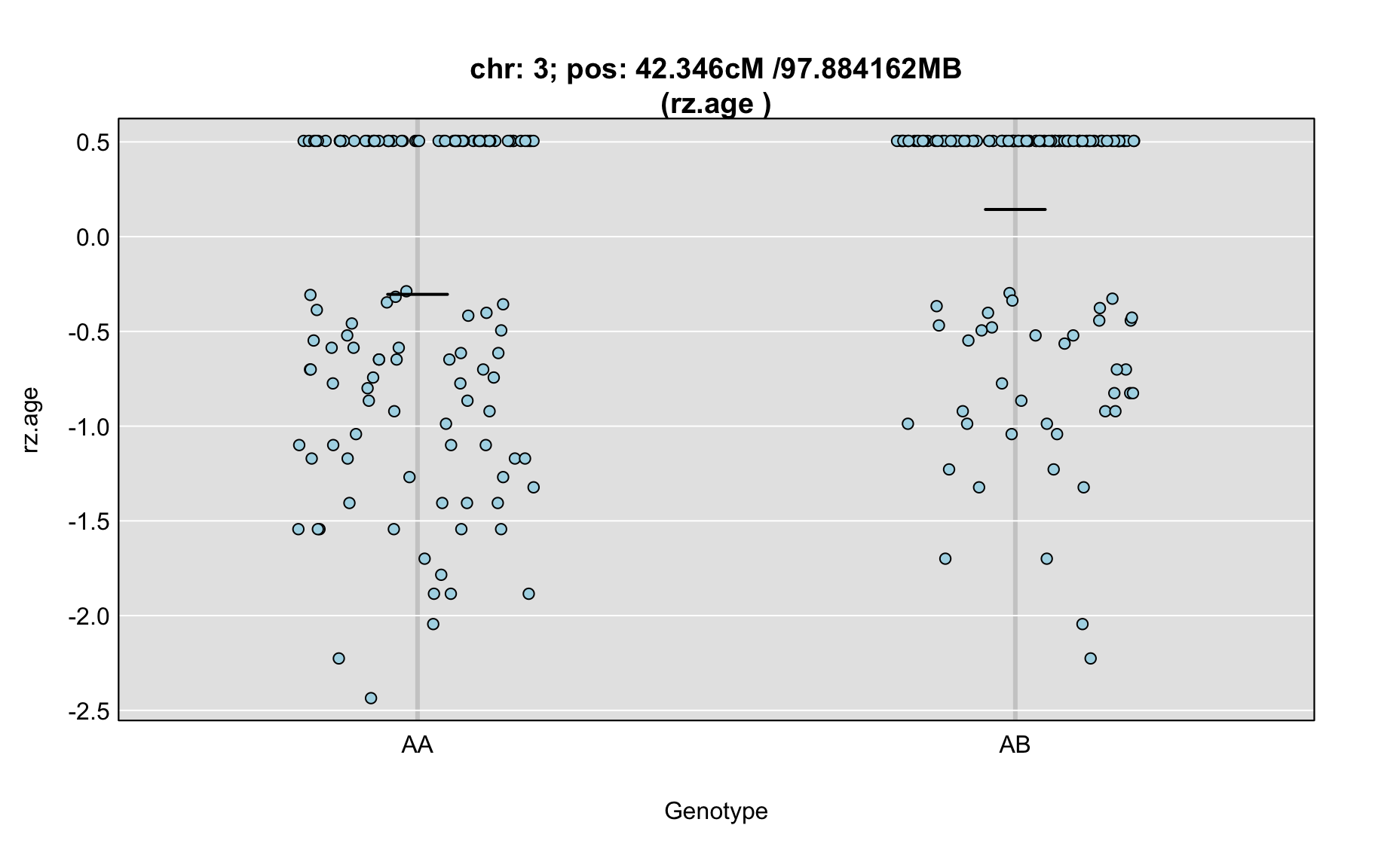
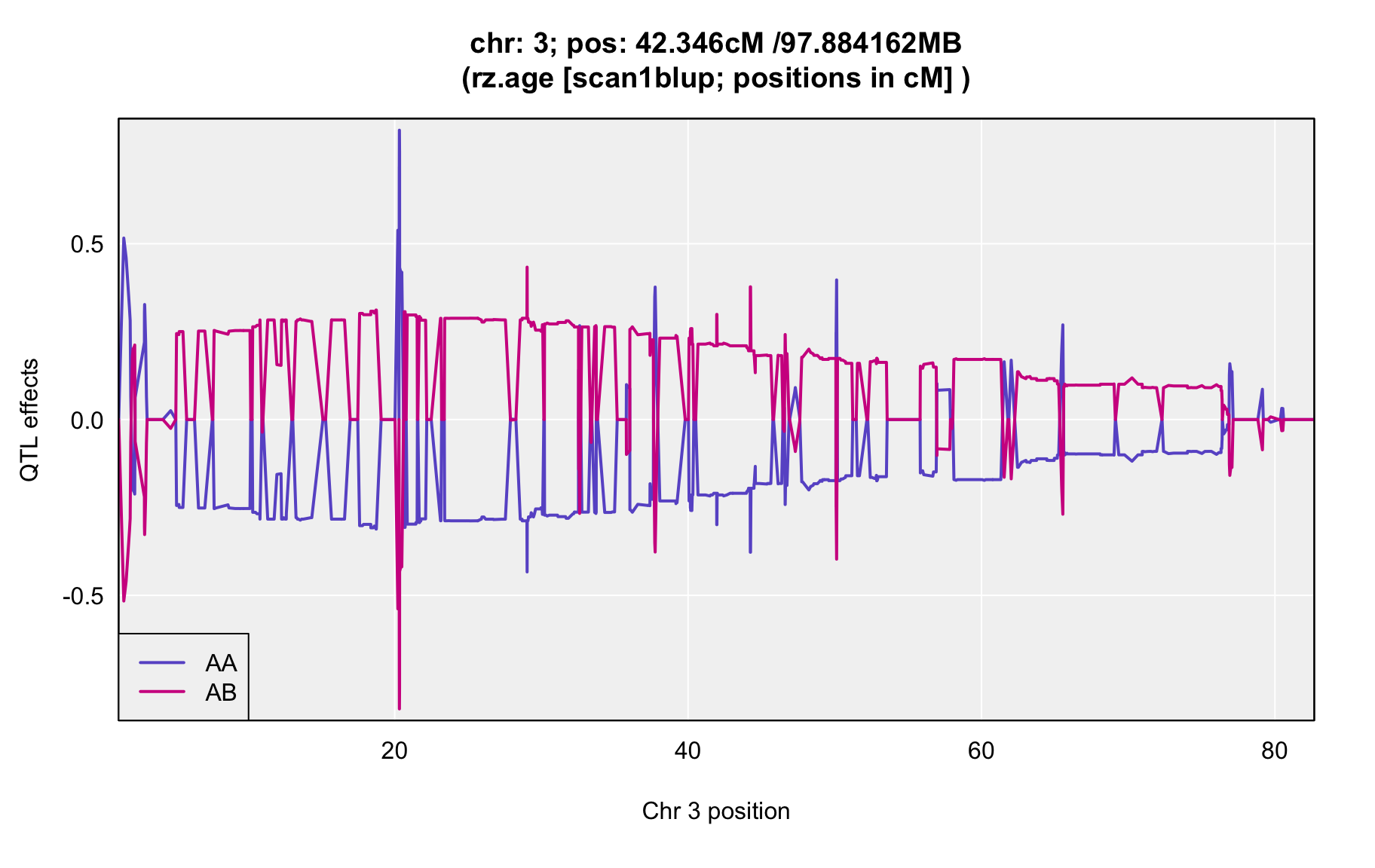
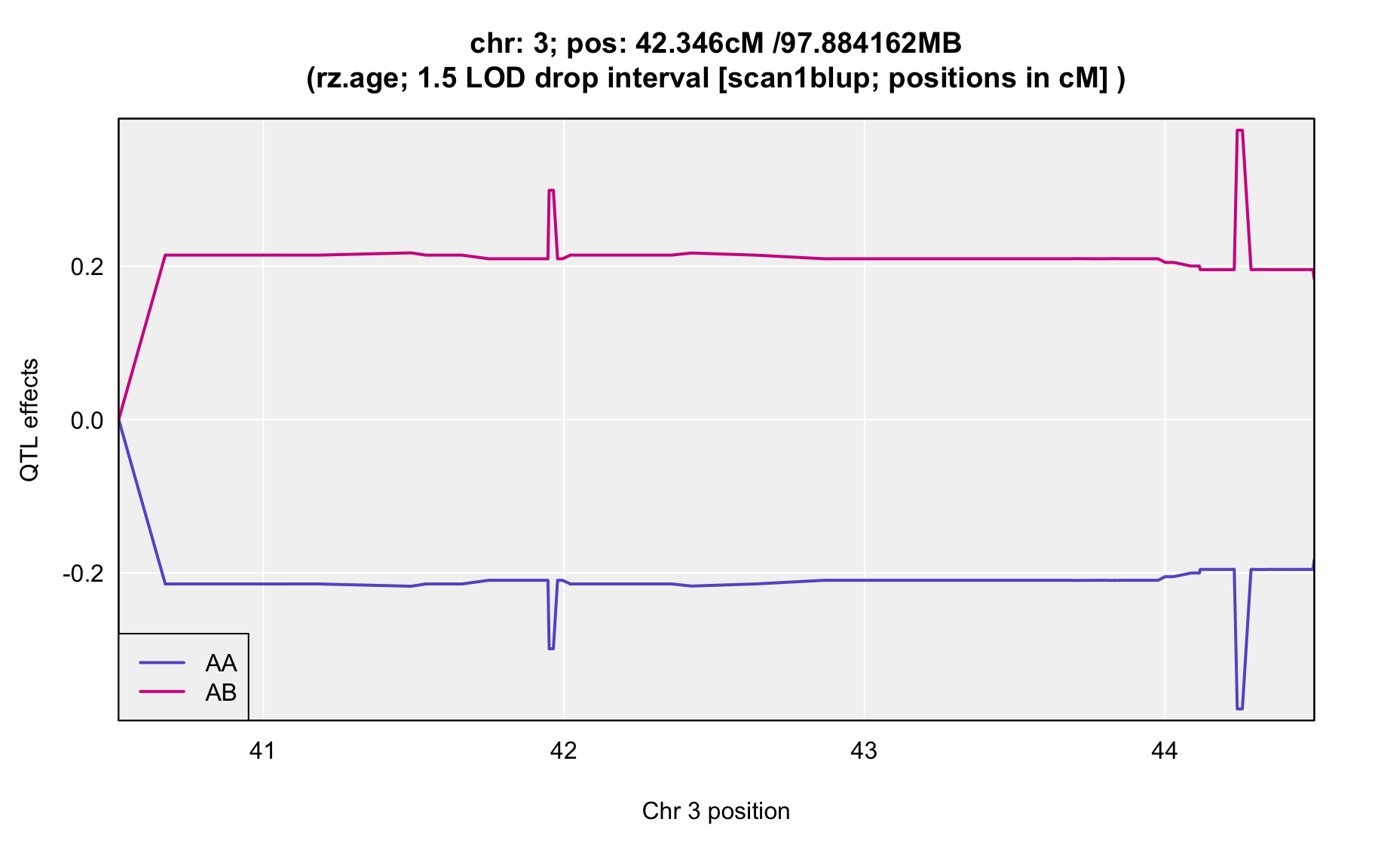
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | pseudogene | 93.90814 | 93.91191 | positive | MGI_C57BL6J_3648187 | Gm9116 | NCBI_Gene:668344,ENSEMBL:ENSMUSG00000102933 | MGI:3648187 | pseudogene | predicted gene 9116 |
| 3 | pseudogene | 93.91284 | 93.91325 | positive | MGI_C57BL6J_5610413 | Gm37185 | ENSEMBL:ENSMUSG00000102769 | MGI:5610413 | pseudogene | predicted gene, 37185 |
| 3 | gene | 93.93663 | 93.94354 | negative | MGI_C57BL6J_3704095 | Gm9117 | NCBI_Gene:668346,ENSEMBL:ENSMUSG00000095822 | MGI:3704095 | protein coding gene | predicted pseudogene 9117 |
| 3 | pseudogene | 93.95807 | 93.95827 | positive | MGI_C57BL6J_5610332 | Gm37104 | ENSEMBL:ENSMUSG00000102445 | MGI:5610332 | pseudogene | predicted gene, 37104 |
| 3 | gene | 93.96029 | 93.96131 | negative | MGI_C57BL6J_3027905 | Tdpoz5 | NCBI_Gene:399676,ENSEMBL:ENSMUSG00000094163 | MGI:3027905 | protein coding gene | TD and POZ domain containing 5 |
| 3 | pseudogene | 93.96914 | 93.96933 | positive | MGI_C57BL6J_5611035 | Gm37807 | ENSEMBL:ENSMUSG00000103841 | MGI:5611035 | pseudogene | predicted gene, 37807 |
| 3 | pseudogene | 93.97771 | 93.97811 | negative | MGI_C57BL6J_5610709 | Gm37481 | ENSEMBL:ENSMUSG00000102804 | MGI:5610709 | pseudogene | predicted gene, 37481 |
| 3 | pseudogene | 93.97904 | 93.98283 | negative | MGI_C57BL6J_3644976 | Gm9121 | NCBI_Gene:668353,ENSEMBL:ENSMUSG00000104460 | MGI:3644976 | pseudogene | predicted pseudogene 9121 |
| 3 | pseudogene | 94.00828 | 94.01139 | positive | MGI_C57BL6J_3644979 | Gm9124 | NCBI_Gene:668356,ENSEMBL:ENSMUSG00000104059 | MGI:3644979 | pseudogene | predicted pseudogene 9124 |
| 3 | pseudogene | 94.01277 | 94.01296 | negative | MGI_C57BL6J_5610313 | Gm37085 | ENSEMBL:ENSMUSG00000102857 | MGI:5610313 | pseudogene | predicted gene, 37085 |
| 3 | gene | 94.04798 | 94.05488 | negative | MGI_C57BL6J_3702970 | Tdpoz9 | NCBI_Gene:668359,ENSEMBL:ENSMUSG00000094328 | MGI:3702970 | protein coding gene | TD and POZ domain containing 9 |
| 3 | pseudogene | 94.06940 | 94.06960 | positive | MGI_C57BL6J_5610309 | Gm37081 | ENSEMBL:ENSMUSG00000102655 | MGI:5610309 | pseudogene | predicted gene, 37081 |
| 3 | gene | 94.07162 | 94.07264 | negative | MGI_C57BL6J_3710527 | Tdpoz7 | NCBI_Gene:100042761,ENSEMBL:ENSMUSG00000095251 | MGI:3710527 | protein coding gene | Td and POZ domain containing 7 |
| 3 | pseudogene | 94.07991 | 94.08010 | positive | MGI_C57BL6J_5611006 | Gm37778 | ENSEMBL:ENSMUSG00000102696 | MGI:5611006 | pseudogene | predicted gene, 37778 |
| 3 | pseudogene | 94.08848 | 94.08888 | negative | MGI_C57BL6J_5610383 | Gm37155 | ENSEMBL:ENSMUSG00000103239 | MGI:5610383 | pseudogene | predicted gene, 37155 |
| 3 | pseudogene | 94.08982 | 94.09360 | negative | MGI_C57BL6J_3779496 | Gm5542 | NCBI_Gene:433624,ENSEMBL:ENSMUSG00000102462 | MGI:3779496 | pseudogene | predicted gene 5542 |
| 3 | pseudogene | 94.11833 | 94.12144 | positive | MGI_C57BL6J_3648392 | Gm9131 | NCBI_Gene:668367,ENSEMBL:ENSMUSG00000104338 | MGI:3648392 | pseudogene | predicted gene 9131 |
| 3 | pseudogene | 94.12282 | 94.12301 | negative | MGI_C57BL6J_5611110 | Gm37882 | ENSEMBL:ENSMUSG00000102949 | MGI:5611110 | pseudogene | predicted gene, 37882 |
| 3 | pseudogene | 94.15763 | 94.15872 | negative | MGI_C57BL6J_3645007 | Gm5543 | NCBI_Gene:433625,ENSEMBL:ENSMUSG00000102311 | MGI:3645007 | pseudogene | predicted gene 5543 |
| 3 | pseudogene | 94.16655 | 94.16674 | positive | MGI_C57BL6J_5610229 | Gm37001 | ENSEMBL:ENSMUSG00000102285 | MGI:5610229 | pseudogene | predicted gene, 37001 |
| 3 | pseudogene | 94.16696 | 94.16985 | negative | MGI_C57BL6J_5434310 | Gm20955 | NCBI_Gene:100418020 | MGI:5434310 | pseudogene | predicted gene, 20955 |
| 3 | pseudogene | 94.17308 | 94.17348 | negative | MGI_C57BL6J_4414980 | Gm16560 | ENSEMBL:ENSMUSG00000089826 | MGI:4414980 | pseudogene | predicted gene 16560 |
| 3 | gene | 94.17441 | 94.17819 | negative | MGI_C57BL6J_3702972 | Spopfm2 | NCBI_Gene:100043188,ENSEMBL:ENSMUSG00000074424 | MGI:3702972 | protein coding gene | speckle-type BTB/POZ protein family member 2 |
| 3 | gene | 94.19671 | 94.19983 | positive | MGI_C57BL6J_3644284 | Spopfm3 | NCBI_Gene:383977,ENSEMBL:ENSMUSG00000090268 | MGI:3644284 | protein coding gene | speckle-type BTB/POZ protein family member 3 |
| 3 | pseudogene | 94.20121 | 94.20140 | negative | MGI_C57BL6J_4415004 | Gm16584 | ENSEMBL:ENSMUSG00000090111 | MGI:4415004 | pseudogene | predicted gene 16584 |
| 3 | pseudogene | 94.23670 | 94.23780 | negative | MGI_C57BL6J_3646925 | Gm4982 | NCBI_Gene:245290,ENSEMBL:ENSMUSG00000089689 | MGI:3646925 | pseudogene | predicted gene 4982 |
| 3 | pseudogene | 94.24244 | 94.24347 | negative | MGI_C57BL6J_3643267 | Gm9141 | NCBI_Gene:668385,ENSEMBL:ENSMUSG00000089893 | MGI:3643267 | pseudogene | predicted gene 9141 |
| 3 | gene | 94.26404 | 94.26718 | positive | MGI_C57BL6J_3643869 | Spopfm1 | NCBI_Gene:212727,ENSEMBL:ENSMUSG00000089696 | MGI:3643869 | protein coding gene | speckle-type BTB/POZ protein family member 1 |
| 3 | pseudogene | 94.27523 | 94.27542 | negative | MGI_C57BL6J_4414981 | Gm16561 | ENSEMBL:ENSMUSG00000089891 | MGI:4414981 | pseudogene | predicted gene 16561 |
| 3 | gene | 94.27797 | 94.30999 | negative | MGI_C57BL6J_5595169 | Gm36010 | NCBI_Gene:102639779 | MGI:5595169 | lncRNA gene | predicted gene, 36010 |
| 3 | gene | 94.31009 | 94.33254 | positive | MGI_C57BL6J_1923028 | Them4 | NCBI_Gene:75778,ENSEMBL:ENSMUSG00000028145 | MGI:1923028 | protein coding gene | thioesterase superfamily member 4 |
| 3 | pseudogene | 94.33439 | 94.33533 | negative | MGI_C57BL6J_5663880 | Gm43743 | ENSEMBL:ENSMUSG00000105506 | MGI:5663880 | pseudogene | predicted gene 43743 |
| 3 | gene | 94.34210 | 94.34735 | positive | MGI_C57BL6J_1913448 | Them5 | NCBI_Gene:66198,ENSEMBL:ENSMUSG00000028148 | MGI:1913448 | protein coding gene | thioesterase superfamily member 5 |
| 3 | gene | 94.35651 | 94.36084 | negative | MGI_C57BL6J_5595229 | Gm36070 | ENSEMBL:ENSMUSG00000105708 | MGI:5595229 | lncRNA gene | predicted gene, 36070 |
| 3 | gene | 94.36244 | 94.36457 | positive | MGI_C57BL6J_2685505 | C2cd4d | NCBI_Gene:271944,ENSEMBL:ENSMUSG00000091648 | MGI:2685505 | protein coding gene | C2 calcium-dependent domain containing 4D |
| 3 | gene | 94.36553 | 94.37725 | negative | MGI_C57BL6J_5621296 | Gm38411 | NCBI_Gene:329711,ENSEMBL:ENSMUSG00000104669 | MGI:5621296 | lncRNA gene | predicted gene, 38411 |
| 3 | gene | 94.37272 | 94.39828 | positive | MGI_C57BL6J_104856 | Rorc | NCBI_Gene:19885,ENSEMBL:ENSMUSG00000028150 | MGI:104856 | protein coding gene | RAR-related orphan receptor gamma |
| 3 | gene | 94.39297 | 94.40285 | negative | MGI_C57BL6J_5663538 | Gm43401 | ENSEMBL:ENSMUSG00000105391 | MGI:5663538 | lncRNA gene | predicted gene 43401 |
| 3 | gene | 94.39852 | 94.40450 | positive | MGI_C57BL6J_2444651 | Lingo4 | NCBI_Gene:320747,ENSEMBL:ENSMUSG00000044505 | MGI:2444651 | protein coding gene | leucine rich repeat and Ig domain containing 4 |
| 3 | gene | 94.40939 | 94.41294 | negative | MGI_C57BL6J_1923852 | 1700040D17Rik | NCBI_Gene:76602,ENSEMBL:ENSMUSG00000097515 | MGI:1923852 | lncRNA gene | RIKEN cDNA 1700040D17 gene |
| 3 | gene | 94.41314 | 94.43485 | positive | MGI_C57BL6J_1919884 | Tdrkh | NCBI_Gene:72634,ENSEMBL:ENSMUSG00000041912 | MGI:1919884 | protein coding gene | tudor and KH domain containing protein |
| 3 | gene | 94.41340 | 94.41829 | positive | MGI_C57BL6J_5662600 | Gm42463 | ENSEMBL:ENSMUSG00000106032 | MGI:5662600 | lncRNA gene | predicted gene 42463 |
| 3 | pseudogene | 94.42166 | 94.42243 | negative | MGI_C57BL6J_5662601 | Gm42464 | ENSEMBL:ENSMUSG00000104756 | MGI:5662601 | pseudogene | predicted gene 42464 |
| 3 | gene | 94.43241 | 94.44392 | negative | MGI_C57BL6J_1858170 | Oaz3 | NCBI_Gene:53814,ENSEMBL:ENSMUSG00000028141 | MGI:1858170 | protein coding gene | ornithine decarboxylase antizyme 3 |
| 3 | gene | 94.44315 | 94.45113 | positive | MGI_C57BL6J_2137211 | Mrpl9 | NCBI_Gene:78523,ENSEMBL:ENSMUSG00000028140 | MGI:2137211 | protein coding gene | mitochondrial ribosomal protein L9 |
| 3 | pseudogene | 94.44854 | 94.45407 | positive | MGI_C57BL6J_5589182 | Gm30023 | NCBI_Gene:102631769,ENSEMBL:ENSMUSG00000104695 | MGI:5589182 | pseudogene | predicted gene, 30023 |
| 3 | gene | 94.46498 | 94.48034 | negative | MGI_C57BL6J_1913603 | Riiad1 | NCBI_Gene:66353,ENSEMBL:ENSMUSG00000028139 | MGI:1913603 | protein coding gene | regulatory subunit of type II PKA R-subunit (RIIa) domain containing 1 |
| 3 | gene | 94.47828 | 94.49220 | positive | MGI_C57BL6J_1926034 | Celf3 | NCBI_Gene:78784,ENSEMBL:ENSMUSG00000028137 | MGI:1926034 | protein coding gene | CUGBP, Elav-like family member 3 |
| 3 | gene | 94.48334 | 94.48399 | negative | MGI_C57BL6J_5826450 | Gm46813 | NCBI_Gene:108168881 | MGI:5826450 | lncRNA gene | predicted gene, 46813 |
| 3 | gene | 94.49754 | 94.58272 | negative | MGI_C57BL6J_1923992 | Snx27 | NCBI_Gene:76742,ENSEMBL:ENSMUSG00000028136 | MGI:1923992 | protein coding gene | sorting nexin family member 27 |
| 3 | gene | 94.53381 | 94.53597 | negative | MGI_C57BL6J_5663910 | Gm43773 | ENSEMBL:ENSMUSG00000106061 | MGI:5663910 | lncRNA gene | predicted gene 43773 |
| 3 | gene | 94.53661 | 94.53749 | negative | MGI_C57BL6J_5663911 | Gm43774 | ENSEMBL:ENSMUSG00000104631 | MGI:5663911 | lncRNA gene | predicted gene 43774 |
| 3 | gene | 94.57578 | 94.57703 | negative | MGI_C57BL6J_5663905 | Gm43768 | ENSEMBL:ENSMUSG00000105412 | MGI:5663905 | lncRNA gene | predicted gene 43768 |
| 3 | pseudogene | 94.58947 | 94.58966 | positive | MGI_C57BL6J_3705756 | Gm15235 | ENSEMBL:ENSMUSG00000105522 | MGI:3705756 | pseudogene | predicted gene 15235 |
| 3 | pseudogene | 94.60438 | 94.60485 | negative | MGI_C57BL6J_3646595 | Rpl31-ps11 | NCBI_Gene:668409,ENSEMBL:ENSMUSG00000062896 | MGI:3646595 | pseudogene | ribosomal protein L31, pseudogene 11 |
| 3 | gene | 94.61275 | 94.65887 | negative | MGI_C57BL6J_109572 | Tuft1 | NCBI_Gene:22156,ENSEMBL:ENSMUSG00000005968 | MGI:109572 | protein coding gene | tuftelin 1 |
| 3 | gene | 94.64310 | 94.64360 | positive | MGI_C57BL6J_3779183 | Gm10972 | ENSEMBL:ENSMUSG00000118474 | MGI:3779183 | protein coding gene | predicted gene 10972 |
| 3 | gene | 94.65896 | 94.66012 | positive | MGI_C57BL6J_3705137 | Gm15234 | ENSEMBL:ENSMUSG00000085250 | MGI:3705137 | lncRNA gene | predicted gene 15234 |
| 3 | pseudogene | 94.66059 | 94.67070 | negative | MGI_C57BL6J_3615512 | BC021767 | NCBI_Gene:545551,ENSEMBL:ENSMUSG00000085006 | MGI:3615512 | pseudogene | cDNA sequence BC021767 |
| 3 | gene | 94.67721 | 94.68467 | positive | MGI_C57BL6J_5826451 | Gm46814 | NCBI_Gene:108168882 | MGI:5826451 | lncRNA gene | predicted gene, 46814 |
| 3 | gene | 94.69356 | 94.70441 | positive | MGI_C57BL6J_104859 | Selenbp2 | NCBI_Gene:20342,ENSEMBL:ENSMUSG00000068877 | MGI:104859 | protein coding gene | selenium binding protein 2 |
| 3 | pseudogene | 94.72002 | 94.75818 | negative | MGI_C57BL6J_3705845 | Gm15264 | NCBI_Gene:100042801,ENSEMBL:ENSMUSG00000081355 | MGI:3705845 | pseudogene | predicted gene 15264 |
| 3 | pseudogene | 94.73734 | 94.73771 | positive | MGI_C57BL6J_5663848 | Gm43711 | ENSEMBL:ENSMUSG00000104881 | MGI:5663848 | pseudogene | predicted gene 43711 |
| 3 | gene | 94.76007 | 94.78772 | negative | MGI_C57BL6J_1927237 | Cgn | NCBI_Gene:70737,ENSEMBL:ENSMUSG00000068876 | MGI:1927237 | protein coding gene | cingulin |
| 3 | gene | 94.82085 | 94.82231 | positive | MGI_C57BL6J_5622979 | Gm40094 | NCBI_Gene:105244497 | MGI:5622979 | lncRNA gene | predicted gene, 40094 |
| 3 | gene | 94.82311 | 94.88357 | positive | MGI_C57BL6J_2442117 | Pogz | NCBI_Gene:229584,ENSEMBL:ENSMUSG00000038902 | MGI:2442117 | protein coding gene | pogo transposable element with ZNF domain |
| 3 | pseudogene | 94.83298 | 94.83380 | positive | MGI_C57BL6J_3705753 | Gm15263 | NCBI_Gene:100417859,ENSEMBL:ENSMUSG00000084058 | MGI:3705753 | pseudogene | predicted gene 15263 |
| 3 | gene | 94.88409 | 94.88696 | negative | MGI_C57BL6J_1098257 | Psmb4 | NCBI_Gene:19172,ENSEMBL:ENSMUSG00000005779 | MGI:1098257 | protein coding gene | proteasome (prosome, macropain) subunit, beta type 4 |
| 3 | gene | 94.91796 | 94.92300 | positive | MGI_C57BL6J_5595396 | Gm36237 | NCBI_Gene:102640082 | MGI:5595396 | lncRNA gene | predicted gene, 36237 |
| 3 | gene | 94.92032 | 94.92042 | negative | MGI_C57BL6J_5456056 | Gm26279 | ENSEMBL:ENSMUSG00000087720 | MGI:5456056 | snRNA gene | predicted gene, 26279 |
| 3 | gene | 94.93306 | 94.94476 | positive | MGI_C57BL6J_96825 | Selenbp1 | NCBI_Gene:20341,ENSEMBL:ENSMUSG00000068874 | MGI:96825 | protein coding gene | selenium binding protein 1 |
| 3 | gene | 94.95307 | 94.96156 | positive | MGI_C57BL6J_1858421 | Rfx5 | NCBI_Gene:53970,ENSEMBL:ENSMUSG00000005774 | MGI:1858421 | protein coding gene | regulatory factor X, 5 (influences HLA class II expression) |
| 3 | gene | 94.95331 | 94.95369 | positive | MGI_C57BL6J_1922445 | 4930532J02Rik | NA | NA | unclassified gene | RIKEN cDNA 4930532J02 gene |
| 3 | gene | 94.95903 | 94.97492 | negative | MGI_C57BL6J_1925930 | B230398E01Rik | NCBI_Gene:109334,ENSEMBL:ENSMUSG00000087660 | MGI:1925930 | lncRNA gene | RIKEN cDNA B230398E01 gene |
| 3 | gene | 94.97473 | 95.00694 | positive | MGI_C57BL6J_1334433 | Pi4kb | NCBI_Gene:107650,ENSEMBL:ENSMUSG00000038861 | MGI:1334433 | protein coding gene | phosphatidylinositol 4-kinase beta |
| 3 | gene | 94.99441 | 94.99735 | negative | MGI_C57BL6J_2442973 | A730011C13Rik | ENSEMBL:ENSMUSG00000084846 | MGI:2442973 | lncRNA gene | RIKEN cDNA A730011C13 gene |
| 3 | gene | 94.99467 | 95.01545 | negative | MGI_C57BL6J_1925516 | Zfp687 | NCBI_Gene:78266,ENSEMBL:ENSMUSG00000019338 | MGI:1925516 | protein coding gene | zinc finger protein 687 |
| 3 | gene | 94.99880 | 95.00487 | negative | MGI_C57BL6J_3705193 | Gm15265 | ENSEMBL:ENSMUSG00000086288 | MGI:3705193 | lncRNA gene | predicted gene 15265 |
| 3 | gene | 95.00417 | 95.00424 | positive | MGI_C57BL6J_5530870 | Mir7013 | miRBase:MI0022862,NCBI_Gene:102465613,ENSEMBL:ENSMUSG00000099033 | MGI:5530870 | miRNA gene | microRNA 7013 |
| 3 | gene | 95.01562 | 95.02198 | positive | MGI_C57BL6J_1922187 | 4930481B07Rik | NCBI_Gene:108168883,ENSEMBL:ENSMUSG00000085956 | MGI:1922187 | lncRNA gene | RIKEN cDNA 4930481B07 gene |
| 3 | gene | 95.03269 | 95.04261 | negative | MGI_C57BL6J_1201670 | Psmd4 | NCBI_Gene:19185,ENSEMBL:ENSMUSG00000005625 | MGI:1201670 | protein coding gene | proteasome (prosome, macropain) 26S subunit, non-ATPase, 4 |
| 3 | gene | 95.04233 | 95.04293 | positive | MGI_C57BL6J_1923891 | 1700123N01Rik | NA | NA | unclassified gene | RIKEN cDNA 1700123N01 gene |
| 3 | gene | 95.05853 | 95.10693 | negative | MGI_C57BL6J_107929 | Pip5k1a | NCBI_Gene:18720,ENSEMBL:ENSMUSG00000028126 | MGI:107929 | protein coding gene | phosphatidylinositol-4-phosphate 5-kinase, type 1 alpha |
| 3 | gene | 95.09063 | 95.09071 | negative | MGI_C57BL6J_5454697 | Gm24920 | ENSEMBL:ENSMUSG00000064952 | MGI:5454697 | snoRNA gene | predicted gene, 24920 |
| 3 | pseudogene | 95.09704 | 95.09737 | negative | MGI_C57BL6J_5622980 | Gm40095 | NCBI_Gene:105244498 | MGI:5622980 | pseudogene | predicted gene, 40095 |
| 3 | gene | 95.09950 | 95.09961 | positive | MGI_C57BL6J_5452078 | Gm22301 | ENSEMBL:ENSMUSG00000064590 | MGI:5452078 | snRNA gene | predicted gene, 22301 |
| 3 | gene | 95.11102 | 95.12305 | positive | MGI_C57BL6J_1202305 | Vps72 | NCBI_Gene:21427,ENSEMBL:ENSMUSG00000008958 | MGI:1202305 | protein coding gene | vacuolar protein sorting 72 |
| 3 | gene | 95.12280 | 95.12921 | positive | MGI_C57BL6J_1355285 | Tmod4 | NCBI_Gene:50874,ENSEMBL:ENSMUSG00000005628 | MGI:1355285 | protein coding gene | tropomodulin 4 |
| 3 | gene | 95.12954 | 95.13405 | negative | MGI_C57BL6J_1341284 | Scnm1 | NCBI_Gene:69269,ENSEMBL:ENSMUSG00000092607 | MGI:1341284 | protein coding gene | sodium channel modifier 1 |
| 3 | gene | 95.13409 | 95.13953 | positive | MGI_C57BL6J_1919409 | Lysmd1 | NCBI_Gene:217779,ENSEMBL:ENSMUSG00000053769 | MGI:1919409 | protein coding gene | LysM, putative peptidoglycan-binding, domain containing 1 |
| 3 | gene | 95.13906 | 95.13919 | negative | MGI_C57BL6J_5531299 | Mir8099-1 | miRBase:MI0026027,NCBI_Gene:102465890,ENSEMBL:ENSMUSG00000098498 | MGI:5531299 | miRNA gene | microRNA 8099-1 |
| 3 | gene | 95.13952 | 95.14245 | negative | MGI_C57BL6J_1917019 | Tnfaip8l2 | NCBI_Gene:69769,ENSEMBL:ENSMUSG00000013707 | MGI:1917019 | protein coding gene | tumor necrosis factor, alpha-induced protein 8-like 2 |
| 3 | pseudogene | 95.14298 | 95.14515 | positive | MGI_C57BL6J_3648561 | Gm9169 | NCBI_Gene:668436,ENSEMBL:ENSMUSG00000098198 | MGI:3648561 | pseudogene | predicted gene 9169 |
| 3 | gene | 95.16042 | 95.17405 | positive | MGI_C57BL6J_1338032 | Sema6c | NCBI_Gene:20360,ENSEMBL:ENSMUSG00000038777 | MGI:1338032 | protein coding gene | sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6C |
| 3 | gene | 95.18177 | 95.21808 | negative | MGI_C57BL6J_95612 | Gabpb2 | NCBI_Gene:213054,ENSEMBL:ENSMUSG00000038766 | MGI:95612 | protein coding gene | GA repeat binding protein, beta 2 |
| 3 | gene | 95.21746 | 95.22666 | positive | MGI_C57BL6J_4439664 | Gm16740 | NCBI_Gene:100504029,ENSEMBL:ENSMUSG00000097167 | MGI:4439664 | lncRNA gene | predicted gene, 16740 |
| 3 | gene | 95.21854 | 95.23220 | negative | MGI_C57BL6J_1929671 | Mllt11 | NCBI_Gene:56772,ENSEMBL:ENSMUSG00000053192 | MGI:1929671 | protein coding gene | myeloid/lymphoid or mixed-lineage leukemia, translocated to, 11 |
| 3 | gene | 95.22842 | 95.23642 | positive | MGI_C57BL6J_1889510 | Cdc42se1 | NCBI_Gene:57912,ENSEMBL:ENSMUSG00000046722 | MGI:1889510 | protein coding gene | CDC42 small effector 1 |
| 3 | gene | 95.23692 | 95.24160 | negative | MGI_C57BL6J_2684974 | Gm128 | NCBI_Gene:229588,ENSEMBL:ENSMUSG00000068860 | MGI:2684974 | protein coding gene | predicted gene 128 |
| 3 | gene | 95.24127 | 95.25119 | negative | MGI_C57BL6J_2384749 | Bnipl | NCBI_Gene:171388,ENSEMBL:ENSMUSG00000028115 | MGI:2384749 | protein coding gene | BCL2/adenovirus E1B 19kD interacting protein like |
| 3 | gene | 95.25367 | 95.28208 | negative | MGI_C57BL6J_1925152 | Prune1 | NCBI_Gene:229589,ENSEMBL:ENSMUSG00000015711 | MGI:1925152 | protein coding gene | prune exopolyphosphatase |
| 3 | gene | 95.28135 | 95.29617 | positive | MGI_C57BL6J_1922257 | Mindy1 | NCBI_Gene:75007,ENSEMBL:ENSMUSG00000038712 | MGI:1922257 | protein coding gene | MINDY lysine 48 deubiquitinase 1 |
| 3 | gene | 95.29609 | 95.31489 | negative | MGI_C57BL6J_1923711 | Anxa9 | NCBI_Gene:71790,ENSEMBL:ENSMUSG00000015702 | MGI:1923711 | protein coding gene | annexin A9 |
| 3 | gene | 95.30708 | 95.31513 | negative | MGI_C57BL6J_1923443 | 6330562C20Rik | ENSEMBL:ENSMUSG00000093424 | MGI:1923443 | lncRNA gene | RIKEN cDNA 6330562C20 gene |
| 3 | gene | 95.31479 | 95.32360 | positive | MGI_C57BL6J_1924143 | Cers2 | NCBI_Gene:76893,ENSEMBL:ENSMUSG00000015714 | MGI:1924143 | protein coding gene | ceramide synthase 2 |
| 3 | gene | 95.32353 | 95.35720 | negative | MGI_C57BL6J_1934229 | Setdb1 | NCBI_Gene:84505,ENSEMBL:ENSMUSG00000015697 | MGI:1934229 | protein coding gene | SET domain, bifurcated 1 |
| 3 | pseudogene | 95.35278 | 95.35298 | positive | MGI_C57BL6J_5662715 | Gm42578 | ENSEMBL:ENSMUSG00000105860 | MGI:5662715 | pseudogene | predicted gene 42578 |
| 3 | gene | 95.35613 | 95.35757 | positive | MGI_C57BL6J_5610728 | Gm37500 | NCBI_Gene:108168884,ENSEMBL:ENSMUSG00000103900 | MGI:5610728 | protein coding gene | predicted gene, 37500 |
| 3 | gene | 95.38646 | 95.40284 | negative | MGI_C57BL6J_1914904 | 4930558C23Rik | NCBI_Gene:67654,ENSEMBL:ENSMUSG00000105734 | MGI:1914904 | protein coding gene | RIKEN cDNA 4930558C23 gene |
| 3 | pseudogene | 95.41030 | 95.41124 | negative | MGI_C57BL6J_3643219 | Gm5070 | NCBI_Gene:277589,ENSEMBL:ENSMUSG00000096177 | MGI:3643219 | pseudogene | predicted gene 5070 |
| 3 | pseudogene | 95.41498 | 95.41545 | negative | MGI_C57BL6J_3647108 | Gm9173 | NCBI_Gene:101055724,ENSEMBL:ENSMUSG00000104646 | MGI:3647108 | pseudogene | predicted gene 9173 |
| 3 | pseudogene | 95.42735 | 95.43109 | positive | MGI_C57BL6J_3782533 | Gm4349 | NCBI_Gene:100043305,ENSEMBL:ENSMUSG00000097532 | MGI:3782533 | pseudogene | predicted gene 4349 |
| 3 | gene | 95.43437 | 95.49724 | positive | MGI_C57BL6J_88071 | Arnt | NCBI_Gene:11863,ENSEMBL:ENSMUSG00000015522 | MGI:88071 | protein coding gene | aryl hydrocarbon receptor nuclear translocator |
| 3 | pseudogene | 95.46119 | 95.46154 | positive | MGI_C57BL6J_5662809 | Gm42672 | ENSEMBL:ENSMUSG00000105395 | MGI:5662809 | pseudogene | predicted gene 42672 |
| 3 | gene | 95.49921 | 95.50939 | positive | MGI_C57BL6J_107823 | Ctsk | NCBI_Gene:13038,ENSEMBL:ENSMUSG00000028111 | MGI:107823 | protein coding gene | cathepsin K |
| 3 | gene | 95.52321 | 95.52450 | negative | MGI_C57BL6J_5622981 | Gm40096 | NCBI_Gene:105244499 | MGI:5622981 | lncRNA gene | predicted gene, 40096 |
| 3 | gene | 95.52679 | 95.55640 | positive | MGI_C57BL6J_107341 | Ctss | NCBI_Gene:13040,ENSEMBL:ENSMUSG00000038642 | MGI:107341 | protein coding gene | cathepsin S |
| 3 | pseudogene | 95.53041 | 95.53104 | positive | MGI_C57BL6J_3776505 | Hmgb1-ps5 | NCBI_Gene:100043312,ENSEMBL:ENSMUSG00000083822 | MGI:3776505 | pseudogene | high mobility group box 1, pseudogene 5 |
| 3 | gene | 95.55964 | 95.58786 | positive | MGI_C57BL6J_1915231 | Hormad1 | NCBI_Gene:67981,ENSEMBL:ENSMUSG00000028109 | MGI:1915231 | protein coding gene | HORMA domain containing 1 |
| 3 | gene | 95.58588 | 95.58883 | negative | MGI_C57BL6J_5622982 | Gm40097 | NCBI_Gene:105244500 | MGI:5622982 | lncRNA gene | predicted gene, 40097 |
| 3 | gene | 95.58892 | 95.61925 | positive | MGI_C57BL6J_1917129 | Golph3l | NCBI_Gene:229593,ENSEMBL:ENSMUSG00000046519 | MGI:1917129 | protein coding gene | golgi phosphoprotein 3-like |
| 3 | gene | 95.58897 | 95.59057 | positive | MGI_C57BL6J_5662808 | Gm42671 | ENSEMBL:ENSMUSG00000105460 | MGI:5662808 | unclassified gene | predicted gene 42671 |
| 3 | pseudogene | 95.62224 | 95.62281 | positive | MGI_C57BL6J_3644167 | Rps10-ps1 | NCBI_Gene:668457,ENSEMBL:ENSMUSG00000060438 | MGI:3644167 | pseudogene | ribosomal protein S10, pseudogene 1 |
| 3 | gene | 95.62498 | 95.63212 | positive | MGI_C57BL6J_1891189 | Ensa | NCBI_Gene:56205,ENSEMBL:ENSMUSG00000038619 | MGI:1891189 | protein coding gene | endosulfine alpha |
| 3 | gene | 95.63930 | 95.64105 | positive | MGI_C57BL6J_3026962 | E330034L11Rik | ENSEMBL:ENSMUSG00000106383 | MGI:3026962 | unclassified gene | RIKEN cDNA E330034L11 gene |
| 3 | gene | 95.65841 | 95.66318 | positive | MGI_C57BL6J_101769 | Mcl1 | NCBI_Gene:17210,ENSEMBL:ENSMUSG00000038612 | MGI:101769 | protein coding gene | myeloid cell leukemia sequence 1 |
| 3 | gene | 95.67620 | 95.68813 | negative | MGI_C57BL6J_2389008 | Adamtsl4 | NCBI_Gene:229595,ENSEMBL:ENSMUSG00000015850 | MGI:2389008 | protein coding gene | ADAMTS-like 4 |
| 3 | gene | 95.73415 | 95.73987 | negative | MGI_C57BL6J_103060 | Ecm1 | NCBI_Gene:13601,ENSEMBL:ENSMUSG00000028108 | MGI:103060 | protein coding gene | extracellular matrix protein 1 |
| 3 | gene | 95.73491 | 95.73497 | negative | MGI_C57BL6J_5531219 | Mir7014 | miRBase:MI0022863,NCBI_Gene:102466787,ENSEMBL:ENSMUSG00000098737 | MGI:5531219 | miRNA gene | microRNA 7014 |
| 3 | gene | 95.73997 | 95.76021 | negative | MGI_C57BL6J_1919057 | Tars2 | NCBI_Gene:71807,ENSEMBL:ENSMUSG00000028107 | MGI:1919057 | protein coding gene | threonyl-tRNA synthetase 2, mitochondrial (putative) |
| 3 | gene | 95.75987 | 95.81979 | negative | MGI_C57BL6J_1922387 | Rprd2 | NCBI_Gene:75137,ENSEMBL:ENSMUSG00000028106 | MGI:1922387 | protein coding gene | regulation of nuclear pre-mRNA domain containing 2 |
| 3 | pseudogene | 95.78072 | 95.78087 | positive | MGI_C57BL6J_5663573 | Gm43436 | ENSEMBL:ENSMUSG00000104967 | MGI:5663573 | pseudogene | predicted gene 43436 |
| 3 | gene | 95.79670 | 95.79688 | negative | MGI_C57BL6J_5456138 | Gm26361 | ENSEMBL:ENSMUSG00000089493 | MGI:5456138 | snRNA gene | predicted gene, 26361 |
| 3 | pseudogene | 95.80482 | 95.80652 | positive | MGI_C57BL6J_5010000 | Gm17815 | NCBI_Gene:100306934,ENSEMBL:ENSMUSG00000105093 | MGI:5010000 | pseudogene | predicted gene, 17815 |
| 3 | gene | 95.81905 | 95.82190 | positive | MGI_C57BL6J_5589107 | Gm29948 | NCBI_Gene:102631665 | MGI:5589107 | lncRNA gene | predicted gene, 29948 |
| 3 | gene | 95.83012 | 95.85680 | negative | MGI_C57BL6J_1918017 | Prpf3 | NCBI_Gene:70767,ENSEMBL:ENSMUSG00000015748 | MGI:1918017 | protein coding gene | pre-mRNA processing factor 3 |
| 3 | gene | 95.85597 | 95.86149 | positive | MGI_C57BL6J_5477088 | Gm26594 | ENSEMBL:ENSMUSG00000097189 | MGI:5477088 | lncRNA gene | predicted gene, 26594 |
| 3 | gene | 95.86263 | 95.87155 | negative | MGI_C57BL6J_1913542 | Mrps21 | NCBI_Gene:66292,ENSEMBL:ENSMUSG00000054312 | MGI:1913542 | protein coding gene | mitochondrial ribosomal protein S21 |
| 3 | gene | 95.87152 | 95.88909 | positive | MGI_C57BL6J_1923991 | C920021L13Rik | NCBI_Gene:100042889,ENSEMBL:ENSMUSG00000080727 | MGI:1923991 | lncRNA gene | RIKEN cDNA C920021L13 gene |
| 3 | gene | 95.87850 | 95.88225 | negative | MGI_C57BL6J_2684975 | Ciart | NCBI_Gene:229599,ENSEMBL:ENSMUSG00000038550 | MGI:2684975 | protein coding gene | circadian associated repressor of transcription |
| 3 | gene | 95.88395 | 95.89295 | negative | MGI_C57BL6J_2385885 | BC028528 | NCBI_Gene:229600,ENSEMBL:ENSMUSG00000038543 | MGI:2385885 | protein coding gene | cDNA sequence BC028528 |
| 3 | gene | 95.89392 | 95.89859 | positive | MGI_C57BL6J_2385110 | Aph1a | NCBI_Gene:226548,ENSEMBL:ENSMUSG00000015750 | MGI:2385110 | protein coding gene | aph1 homolog A, gamma secretase subunit |
| 3 | gene | 95.89777 | 95.90512 | negative | MGI_C57BL6J_1344341 | Car14 | NCBI_Gene:23831,ENSEMBL:ENSMUSG00000038526 | MGI:1344341 | protein coding gene | carbonic anhydrase 14 |
| 3 | gene | 95.92588 | 95.92902 | negative | MGI_C57BL6J_5826452 | Gm46815 | NCBI_Gene:108168885 | MGI:5826452 | lncRNA gene | predicted gene, 46815 |
| 3 | gene | 95.92924 | 95.94739 | positive | MGI_C57BL6J_1913721 | Anp32e | NCBI_Gene:66471,ENSEMBL:ENSMUSG00000015749 | MGI:1913721 | protein coding gene | acidic (leucine-rich) nuclear phosphoprotein 32 family, member E |
| 3 | gene | 95.94009 | 95.94257 | positive | MGI_C57BL6J_5663512 | Gm43375 | ENSEMBL:ENSMUSG00000106262 | MGI:5663512 | unclassified gene | predicted gene 43375 |
| 3 | pseudogene | 95.96403 | 95.96439 | negative | MGI_C57BL6J_5434296 | Gm20940 | ENSEMBL:ENSMUSG00000106640 | MGI:5434296 | pseudogene | predicted gene, 20940 |
| 3 | pseudogene | 95.98281 | 95.98360 | positive | MGI_C57BL6J_3782273 | Gm4097 | NCBI_Gene:100042898 | MGI:3782273 | pseudogene | predicted gene 4097 |
| 3 | gene | 95.98551 | 95.98697 | positive | MGI_C57BL6J_3646918 | Gm9054 | NCBI_Gene:668224 | MGI:3646918 | lncRNA gene | predicted gene 9054 |
| 3 | gene | 95.98843 | 95.99600 | negative | MGI_C57BL6J_1914470 | Plekho1 | NCBI_Gene:67220,ENSEMBL:ENSMUSG00000015745 | MGI:1914470 | protein coding gene | pleckstrin homology domain containing, family O member 1 |
| 3 | gene | 95.99592 | 95.99646 | positive | MGI_C57BL6J_3646916 | Gm9052 | NA | NA | unclassified gene | predicted gene 9052 |
| 3 | gene | 95.99983 | 96.05847 | negative | MGI_C57BL6J_891965 | Vps45 | NCBI_Gene:22365,ENSEMBL:ENSMUSG00000015747 | MGI:891965 | protein coding gene | vacuolar protein sorting 45 |
| 3 | pseudogene | 96.08687 | 96.08706 | positive | MGI_C57BL6J_5663691 | Gm43554 | ENSEMBL:ENSMUSG00000105995 | MGI:5663691 | pseudogene | predicted gene 43554 |
| 3 | gene | 96.09817 | 96.10436 | negative | MGI_C57BL6J_5589203 | Gm30044 | NCBI_Gene:102631798 | MGI:5589203 | lncRNA gene | predicted gene, 30044 |
| 3 | gene | 96.10451 | 96.16113 | positive | MGI_C57BL6J_2654703 | Otud7b | NCBI_Gene:229603,ENSEMBL:ENSMUSG00000038495 | MGI:2654703 | protein coding gene | OTU domain containing 7B |
| 3 | gene | 96.16195 | 96.17172 | positive | MGI_C57BL6J_2652817 | Mtmr11 | NCBI_Gene:194126,ENSEMBL:ENSMUSG00000045934 | MGI:2652817 | protein coding gene | myotubularin related protein 11 |
| 3 | gene | 96.16746 | 96.17242 | negative | MGI_C57BL6J_4937324 | Gm17690 | ENSEMBL:ENSMUSG00000097204 | MGI:4937324 | lncRNA gene | predicted gene, 17690 |
| 3 | gene | 96.17233 | 96.17756 | positive | MGI_C57BL6J_109580 | Sf3b4 | NCBI_Gene:107701,ENSEMBL:ENSMUSG00000068856 | MGI:109580 | protein coding gene | splicing factor 3b, subunit 4 |
| 3 | gene | 96.17636 | 96.17649 | positive | MGI_C57BL6J_5451804 | Gm22027 | ENSEMBL:ENSMUSG00000084686 | MGI:5451804 | snoRNA gene | predicted gene, 22027 |
| 3 | gene | 96.18115 | 96.19552 | positive | MGI_C57BL6J_1927139 | Sv2a | NCBI_Gene:64051,ENSEMBL:ENSMUSG00000038486 | MGI:1927139 | protein coding gene | synaptic vesicle glycoprotein 2 a |
| 3 | gene | 96.19658 | 96.21971 | negative | MGI_C57BL6J_1916418 | Bola1 | NCBI_Gene:69168,ENSEMBL:ENSMUSG00000015943 | MGI:1916418 | protein coding gene | bolA-like 1 (E. coli) |
| 3 | gene | 96.20963 | 96.22017 | negative | MGI_C57BL6J_3705796 | Gm15444 | ENSEMBL:ENSMUSG00000086556 | MGI:3705796 | lncRNA gene | predicted gene 15444 |
| 3 | gene | 96.21986 | 96.22035 | positive | MGI_C57BL6J_2448314 | H2ac21 | NCBI_Gene:621893,ENSEMBL:ENSMUSG00000063689 | MGI:2448314 | protein coding gene | H2A clustered histone 21 |
| 3 | gene | 96.22036 | 96.22088 | negative | MGI_C57BL6J_2448316 | H2ac20 | NCBI_Gene:319176,ENSEMBL:ENSMUSG00000068855 | MGI:2448316 | protein coding gene | H2A clustered histone 20 |
| 3 | gene | 96.22050 | 96.22064 | positive | MGI_C57BL6J_5690884 | Gm44492 | ENSEMBL:ENSMUSG00000106431 | MGI:5690884 | miRNA gene | predicted gene, 44492 |
| 3 | gene | 96.22107 | 96.22374 | positive | MGI_C57BL6J_2448415 | H2bc21 | NCBI_Gene:319190,ENSEMBL:ENSMUSG00000068854 | MGI:2448415 | protein coding gene | H2B clustered histone 21 |
| 3 | gene | 96.23811 | 96.23913 | negative | MGI_C57BL6J_2448357 | H3c15 | NCBI_Gene:97114,ENSEMBL:ENSMUSG00000081058 | MGI:2448357 | protein coding gene | H3 clustered histone 15 |
| 3 | gene | 96.23939 | 96.24182 | positive | MGI_C57BL6J_5313079 | Gm20632 | ENSEMBL:ENSMUSG00000093577 | MGI:5313079 | lncRNA gene | predicted gene 20632 |
| 3 | gene | 96.23975 | 96.24037 | negative | MGI_C57BL6J_2448283 | H2ac19 | NCBI_Gene:319192,ENSEMBL:ENSMUSG00000063954 | MGI:2448283 | protein coding gene | H2A clustered histone 19 |
| 3 | gene | 96.23987 | 96.24000 | positive | MGI_C57BL6J_6096187 | Gm47311 | ENSEMBL:ENSMUSG00000106587 | MGI:6096187 | miRNA gene | predicted gene, 47311 |
| 3 | gene | 96.24355 | 96.24638 | negative | MGI_C57BL6J_5313080 | Gm20633 | ENSEMBL:ENSMUSG00000093553 | MGI:5313080 | lncRNA gene | predicted gene 20633 |
| 3 | gene | 96.24550 | 96.24608 | positive | MGI_C57BL6J_96097 | H2ac18 | NCBI_Gene:15267,ENSEMBL:ENSMUSG00000064220 | MGI:96097 | protein coding gene | H2A clustered histone 18 |
| 3 | gene | 96.24555 | 96.24851 | positive | MGI_C57BL6J_5313081 | Gm20634 | ENSEMBL:ENSMUSG00000070392 | MGI:5313081 | lncRNA gene | predicted gene 20634 |
| 3 | gene | 96.24583 | 96.24596 | negative | MGI_C57BL6J_6096191 | Gm47313 | ENSEMBL:ENSMUSG00000105610 | MGI:6096191 | miRNA gene | predicted gene, 47313 |
| 3 | gene | 96.24668 | 96.24851 | positive | MGI_C57BL6J_2448355 | H3c14 | NCBI_Gene:15077,ENSEMBL:ENSMUSG00000093769 | MGI:2448355 | protein coding gene | H3 clustered histone 14 |
| 3 | gene | 96.25462 | 96.25775 | negative | MGI_C57BL6J_5313074 | Gm20627 | NCBI_Gene:102631874,ENSEMBL:ENSMUSG00000093507 | MGI:5313074 | lncRNA gene | predicted gene 20627 |
| 3 | gene | 96.26168 | 96.26332 | negative | MGI_C57BL6J_2140113 | H4c14 | NCBI_Gene:97122,ENSEMBL:ENSMUSG00000091405 | MGI:2140113 | protein coding gene | H4 clustered histone 14 |
| 3 | gene | 96.26378 | 96.26539 | positive | MGI_C57BL6J_5611455 | Gm38227 | ENSEMBL:ENSMUSG00000103940 | MGI:5611455 | unclassified gene | predicted gene, 38227 |
| 3 | gene | 96.26708 | 96.27029 | negative | MGI_C57BL6J_5313075 | Gm20628 | ENSEMBL:ENSMUSG00000093656 | MGI:5313075 | lncRNA gene | predicted gene 20628 |
| 3 | gene | 96.26865 | 96.26915 | positive | MGI_C57BL6J_2448351 | H3c13 | NCBI_Gene:319154,ENSEMBL:ENSMUSG00000074403 | MGI:2448351 | protein coding gene | H3 clustered histone 13 |
| 3 | gene | 96.26970 | 96.27019 | positive | MGI_C57BL6J_2448413 | H2bc18 | NCBI_Gene:319189,ENSEMBL:ENSMUSG00000105827 | MGI:2448413 | protein coding gene | H2B clustered histone 18 |
| 3 | gene | 96.26972 | 96.27900 | positive | MGI_C57BL6J_5662880 | Gm42743 | ENSEMBL:ENSMUSG00000050936 | MGI:5662880 | lncRNA gene | predicted gene 42743 |
| 3 | gene | 96.28291 | 96.29397 | negative | MGI_C57BL6J_95498 | Fcgr1 | NCBI_Gene:14129,ENSEMBL:ENSMUSG00000015947 | MGI:95498 | protein coding gene | Fc receptor, IgG, high affinity I |
| 3 | gene | 96.32207 | 96.32215 | positive | MGI_C57BL6J_4413741 | n-TNgtt4 | NCBI_Gene:102467475 | MGI:4413741 | tRNA gene | nuclear encoded tRNA asparagine 4 (anticodon GTT) |
| 3 | gene | 96.32842 | 96.32859 | positive | MGI_C57BL6J_5454607 | Gm24830 | ENSEMBL:ENSMUSG00000095701 | MGI:5454607 | snRNA gene | predicted gene, 24830 |
| 3 | gene | 96.33274 | 96.33281 | positive | MGI_C57BL6J_4414101 | n-TVcac10 | NCBI_Gene:102467407 | MGI:4414101 | tRNA gene | nuclear encoded tRNA valine 10 (anticodon CAC) |
| 3 | gene | 96.33461 | 96.33469 | negative | MGI_C57BL6J_4413742 | n-TNgtt5 | NCBI_Gene:102467256 | MGI:4413742 | tRNA gene | nuclear encoded tRNA asparagine 5 (anticodon GTT) |
| 3 | gene | 96.34116 | 96.34123 | negative | MGI_C57BL6J_4413829 | n-TQctg5 | NCBI_Gene:102467292 | MGI:4413829 | tRNA gene | nuclear encoded tRNA glutamine 5 (anticodon CTG) |
| 3 | gene | 96.34529 | 96.35881 | positive | MGI_C57BL6J_4937214 | Platr30 | NCBI_Gene:100504145,ENSEMBL:ENSMUSG00000097851 | MGI:4937214 | lncRNA gene | pluripotency associated transcript 30 |
| 3 | gene | 96.34668 | 96.34675 | positive | MGI_C57BL6J_4413854 | n-TEttc3 | NCBI_Gene:102467302 | MGI:4413854 | tRNA gene | nuclear encoded tRNA glutamic acid 3 (anticodon TTC) |
| 3 | gene | 96.34863 | 96.35715 | negative | MGI_C57BL6J_5477047 | Gm26553 | NCBI_Gene:74690,ENSEMBL:ENSMUSG00000097807 | MGI:5477047 | lncRNA gene | predicted gene, 26553 |
| 3 | gene | 96.35117 | 96.35124 | negative | MGI_C57BL6J_4413866 | n-TGccc5 | NCBI_Gene:102467528 | MGI:4413866 | tRNA gene | nuclear encoded tRNA glycine 5 (anticodon CCC) |
| 3 | gene | 96.35995 | 96.36012 | negative | MGI_C57BL6J_97974 | Rnu1b1 | NCBI_Gene:19844,ENSEMBL:ENSMUSG00000095580 | MGI:97974 | snRNA gene | U1b1 small nuclear RNA |
| 3 | gene | 96.37402 | 96.37418 | positive | MGI_C57BL6J_104624 | Rnu1b2 | NCBI_Gene:19845,ENSEMBL:ENSMUSG00000093834 | MGI:104624 | snRNA gene | U1b2 small nuclear RNA |
| 3 | gene | 96.37565 | 96.38886 | negative | MGI_C57BL6J_4937209 | Gm17575 | ENSEMBL:ENSMUSG00000097420 | MGI:4937209 | lncRNA gene | predicted gene, 17575 |
| 3 | gene | 96.37698 | 96.38552 | positive | MGI_C57BL6J_5477183 | Gm26689 | NCBI_Gene:102632400,ENSEMBL:ENSMUSG00000097753 | MGI:5477183 | lncRNA gene | predicted gene, 26689 |
| 3 | gene | 96.38291 | 96.38298 | positive | MGI_C57BL6J_4413865 | n-TGccc1 | NCBI_Gene:102467307 | MGI:4413865 | tRNA gene | nuclear encoded tRNA glycine 1 (anticodon CCC) |
| 3 | gene | 96.38741 | 96.38748 | negative | MGI_C57BL6J_4413855 | n-TEttc2 | NCBI_Gene:102467523 | MGI:4413855 | tRNA gene | nuclear encoded tRNA glutamic acid 2 (anticodon TTC) |
| 3 | gene | 96.39906 | 96.39913 | negative | MGI_C57BL6J_4413828 | n-TQctg4 | NCBI_Gene:102467413 | MGI:4413828 | tRNA gene | nuclear encoded tRNA glutamine 4 (anticodon CTG) |
| 3 | gene | 96.40115 | 96.40201 | negative | MGI_C57BL6J_1922199 | 4930477E14Rik | ENSEMBL:ENSMUSG00000097539 | MGI:1922199 | lncRNA gene | RIKEN cDNA 4930477E14 gene |
| 3 | gene | 96.40222 | 96.40229 | positive | MGI_C57BL6J_4413747 | n-TNgtt10 | NCBI_Gene:102467258 | MGI:4413747 | tRNA gene | nuclear encoded tRNA asparagine 10 (anticodon GTT) |
| 3 | gene | 96.40238 | 96.40295 | positive | MGI_C57BL6J_5477148 | Gm26654 | NCBI_Gene:105244502,ENSEMBL:ENSMUSG00000097299 | MGI:5477148 | lncRNA gene | predicted gene, 26654 |
| 3 | pseudogene | 96.40738 | 96.40946 | negative | MGI_C57BL6J_103115 | Pafah1b1-ps1 | NCBI_Gene:18473,ENSEMBL:ENSMUSG00000097041 | MGI:103115 | pseudogene | platelet-activating factor acetylhydrolase, isoform 1b, beta1 subunit, pseudogene 1 |
| 3 | gene | 96.41007 | 96.41014 | positive | MGI_C57BL6J_4413896 | n-THgtg6 | NCBI_Gene:102467539 | MGI:4413896 | tRNA gene | nuclear encoded tRNA histidine 6 (anticodon GTG) |
| 3 | pseudogene | 96.41291 | 96.41336 | negative | MGI_C57BL6J_5439429 | Gm21960 | ENSEMBL:ENSMUSG00000095836 | MGI:5439429 | pseudogene | predicted gene, 21960 |
| 3 | gene | 96.41444 | 96.41486 | negative | MGI_C57BL6J_109558 | Terc | NCBI_Gene:21748,ENSEMBL:ENSMUSG00000064796,ENSEMBL:ENSMUSG00000118406 | MGI:109558 | telomerase RNA gene | telomerase RNA component |
| 3 | gene | 96.42273 | 96.42290 | positive | MGI_C57BL6J_5453913 | Gm24136 | ENSEMBL:ENSMUSG00000095513 | MGI:5453913 | snRNA gene | predicted gene, 24136 |
| 3 | gene | 96.42403 | 96.42410 | negative | MGI_C57BL6J_4413867 | n-TGccc6 | NCBI_Gene:102467308 | MGI:4413867 | tRNA gene | nuclear encoded tRNA glycine 6 (anticodon CCC) |
| 3 | gene | 96.42824 | 96.42831 | positive | MGI_C57BL6J_4413962 | n-TKctt15 | NCBI_Gene:102467347 | MGI:4413962 | tRNA gene | nuclear encoded tRNA lysine 15 (anticodon CTT) |
| 3 | gene | 96.43379 | 96.45231 | negative | MGI_C57BL6J_3618860 | BC107364 | NCBI_Gene:329716,ENSEMBL:ENSMUSG00000046317 | MGI:3618860 | protein coding gene | cDNA sequence BC107364 |
| 3 | gene | 96.43597 | 96.43604 | positive | MGI_C57BL6J_4413892 | n-THgtg2 | NCBI_Gene:22053 | MGI:4413892 | tRNA gene | nuclear encoded tRNA histidine 2 (anticodon GTG) |
| 3 | gene | 96.44994 | 96.45164 | positive | MGI_C57BL6J_5456009 | Gm26232 | NCBI_Gene:102632589,ENSEMBL:ENSMUSG00000096838 | MGI:5456009 | snRNA gene | predicted gene, 26232 |
| 3 | gene | 96.45143 | 96.45150 | positive | MGI_C57BL6J_4413748 | n-TNgtt11 | NCBI_Gene:102467478 | MGI:4413748 | tRNA gene | nuclear encoded tRNA asparagine 11 (anticodon GTT) |
| 3 | gene | 96.45249 | 96.45257 | positive | MGI_C57BL6J_4413897 | n-THgtg7 | NCBI_Gene:102467320 | MGI:4413897 | tRNA gene | nuclear encoded tRNA histidine 7 (anticodon GTG) |
| 3 | pseudogene | 96.45454 | 96.45595 | negative | MGI_C57BL6J_3779840 | Gm9207 | NCBI_Gene:668500,ENSEMBL:ENSMUSG00000104642 | MGI:3779840 | pseudogene | predicted gene 9207 |
| 3 | gene | 96.45807 | 96.45814 | negative | MGI_C57BL6J_4413899 | n-THgtg9 | NCBI_Gene:102467414 | MGI:4413899 | tRNA gene | nuclear encoded tRNA histidine 9 (anticodon GTG) |
| 3 | gene | 96.45821 | 96.47066 | positive | MGI_C57BL6J_3641752 | Gm10685 | NCBI_Gene:102632901,ENSEMBL:ENSMUSG00000106429 | MGI:3641752 | lncRNA gene | predicted gene 10685 |
| 3 | gene | 96.45900 | 96.46034 | negative | MGI_C57BL6J_5622984 | Gm40099 | NCBI_Gene:105244504 | MGI:5622984 | lncRNA gene | predicted gene, 40099 |
| 3 | gene | 96.45914 | 96.45921 | negative | MGI_C57BL6J_4413750 | n-TNgtt13 | NCBI_Gene:102467479 | MGI:4413750 | tRNA gene | nuclear encoded tRNA asparagine 13 (anticodon GTT) |
| 3 | gene | 96.46002 | 96.46019 | negative | MGI_C57BL6J_104604 | Rnu1b6 | NCBI_Gene:19847,ENSEMBL:ENSMUSG00000065773 | MGI:104604 | snRNA gene | U1b6 small nuclear RNA |
| 3 | gene | 96.46397 | 96.46404 | negative | MGI_C57BL6J_4413901 | n-THgtg1 | NCBI_Gene:102467321 | MGI:4413901 | tRNA gene | nuclear encoded tRNA histidine 1 (anticodon GTG) |
| 3 | gene | 96.46407 | 96.46692 | negative | MGI_C57BL6J_5663671 | Gm43534 | ENSEMBL:ENSMUSG00000106641 | MGI:5663671 | unclassified gene | predicted gene 43534 |
| 3 | gene | 96.47939 | 96.47946 | positive | MGI_C57BL6J_4413827 | n-TQctg3 | NCBI_Gene:102467511 | MGI:4413827 | tRNA gene | nuclear encoded tRNA glutamine 3 (anticodon CTG) |
| 3 | gene | 96.48516 | 96.48528 | positive | MGI_C57BL6J_5531383 | Gm28001 | ENSEMBL:ENSMUSG00000098392 | MGI:5531383 | rRNA gene | predicted gene, 28001 |
| 3 | gene | 96.48655 | 96.48672 | negative | MGI_C57BL6J_5455667 | Gm25890 | ENSEMBL:ENSMUSG00000095260 | MGI:5455667 | snRNA gene | predicted gene, 25890 |
| 3 | gene | 96.48873 | 96.48889 | positive | MGI_C57BL6J_5452391 | Gm22614 | ENSEMBL:ENSMUSG00000094812 | MGI:5452391 | snRNA gene | predicted gene, 22614 |
| 3 | gene | 96.49778 | 96.49785 | positive | MGI_C57BL6J_4413749 | n-TNgtt12 | NCBI_Gene:102467259 | MGI:4413749 | tRNA gene | nuclear encoded tRNA asparagine 12 (anticodon GTT) |
| 3 | gene | 96.49951 | 96.49958 | negative | MGI_C57BL6J_4413963 | n-TKctt16 | NCBI_Gene:102467567 | MGI:4413963 | tRNA gene | nuclear encoded tRNA lysine 16 (anticodon CTT) |
| 3 | gene | 96.50037 | 96.50044 | negative | MGI_C57BL6J_4413898 | n-THgtg8 | NCBI_Gene:104358 | MGI:4413898 | tRNA gene | nuclear encoded tRNA histidine 8 (anticodon GTG) |
| 3 | gene | 96.50101 | 96.50108 | negative | MGI_C57BL6J_4413890 | n-TGtcc7 | NCBI_Gene:102467318 | MGI:4413890 | tRNA gene | nuclear encoded tRNA glycine 7 (anticodon TCC) |
| 3 | gene | 96.50145 | 96.50152 | negative | MGI_C57BL6J_4413853 | n-TEctc13 | NCBI_Gene:102467428 | MGI:4413853 | tRNA gene | nuclear encoded tRNA glutamic acid 13 (anticodon CTC) |
| 3 | gene | 96.50639 | 96.50878 | negative | MGI_C57BL6J_5622985 | Gm40100 | NCBI_Gene:105244505 | MGI:5622985 | lncRNA gene | predicted gene, 40100 |
| 3 | gene | 96.52516 | 96.52922 | positive | MGI_C57BL6J_1916835 | Hjv | NCBI_Gene:69585,ENSEMBL:ENSMUSG00000038403 | MGI:1916835 | protein coding gene | hemojuvelin BMP co-receptor |
| 3 | gene | 96.55576 | 96.56680 | negative | MGI_C57BL6J_3641753 | Gm15441 | NCBI_Gene:100038464,ENSEMBL:ENSMUSG00000074398 | MGI:3641753 | lncRNA gene | predicted gene 15441 |
| 3 | gene | 96.55796 | 96.56188 | positive | MGI_C57BL6J_1889549 | Txnip | NCBI_Gene:56338,ENSEMBL:ENSMUSG00000038393 | MGI:1889549 | protein coding gene | thioredoxin interacting protein |
| 3 | gene | 96.57698 | 96.58066 | positive | MGI_C57BL6J_3826531 | Gm16253 | ENSEMBL:ENSMUSG00000087610 | MGI:3826531 | protein coding gene | predicted gene 16253 |
| 3 | gene | 96.57787 | 96.59418 | negative | MGI_C57BL6J_1917120 | Polr3gl | NCBI_Gene:69870,ENSEMBL:ENSMUSG00000028104 | MGI:1917120 | protein coding gene | polymerase (RNA) III (DNA directed) polypeptide G like |
| 3 | gene | 96.59548 | 96.60013 | positive | MGI_C57BL6J_3617846 | Ankrd34a | NCBI_Gene:545554,ENSEMBL:ENSMUSG00000049097 | MGI:3617846 | protein coding gene | ankyrin repeat domain 34A |
| 3 | gene | 96.60113 | 96.62617 | positive | MGI_C57BL6J_3036267 | Lix1l | NCBI_Gene:280411,ENSEMBL:ENSMUSG00000049288 | MGI:3036267 | protein coding gene | Lix1-like |
| 3 | pseudogene | 96.60666 | 96.62982 | negative | MGI_C57BL6J_2442586 | 6330549D23Rik | NCBI_Gene:229613,ENSEMBL:ENSMUSG00000045327 | MGI:2442586 | pseudogene | RIKEN cDNA 6330549D23 gene |
| 3 | gene | 96.62993 | 96.63379 | positive | MGI_C57BL6J_1913129 | Rbm8a | NCBI_Gene:60365,ENSEMBL:ENSMUSG00000038374 | MGI:1913129 | protein coding gene | RNA binding motif protein 8a |
| 3 | gene | 96.63536 | 96.64538 | positive | MGI_C57BL6J_1338882 | Pex11b | NCBI_Gene:18632,ENSEMBL:ENSMUSG00000028102 | MGI:1338882 | protein coding gene | peroxisomal biogenesis factor 11 beta |
| 3 | gene | 96.63547 | 96.65310 | positive | MGI_C57BL6J_5663094 | Gm42957 | ENSEMBL:ENSMUSG00000106447 | MGI:5663094 | protein coding gene | predicted gene 42957 |
| 3 | gene | 96.64558 | 96.66452 | positive | MGI_C57BL6J_2153482 | Itga10 | NCBI_Gene:213119,ENSEMBL:ENSMUSG00000090210 | MGI:2153482 | protein coding gene | integrin, alpha 10 |
| 3 | pseudogene | 96.66647 | 96.66702 | negative | MGI_C57BL6J_3648207 | Rpl21-ps11 | NCBI_Gene:433630,ENSEMBL:ENSMUSG00000105144 | MGI:3648207 | pseudogene | ribosomal protein L21, pseudogene 11 |
| 3 | gene | 96.67013 | 96.69103 | positive | MGI_C57BL6J_2442590 | Ankrd35 | NCBI_Gene:213121,ENSEMBL:ENSMUSG00000038354 | MGI:2442590 | protein coding gene | ankyrin repeat domain 35 |
| 3 | gene | 96.69386 | 96.69701 | negative | MGI_C57BL6J_2444301 | C230057H02Rik | NA | NA | unclassified gene | RIKEN cDNA C230057H02 gene |
| 3 | gene | 96.69638 | 96.70607 | positive | MGI_C57BL6J_1913126 | Pias3 | NCBI_Gene:229615,ENSEMBL:ENSMUSG00000028101 | MGI:1913126 | protein coding gene | protein inhibitor of activated STAT 3 |
| 3 | gene | 96.70589 | 96.70893 | negative | MGI_C57BL6J_1925623 | Nudt17 | NCBI_Gene:78373,ENSEMBL:ENSMUSG00000028100 | MGI:1925623 | protein coding gene | nudix (nucleoside diphosphate linked moiety X)-type motif 17 |
| 3 | gene | 96.71149 | 96.72763 | negative | MGI_C57BL6J_1921664 | Polr3c | NCBI_Gene:74414,ENSEMBL:ENSMUSG00000028099 | MGI:1921664 | protein coding gene | polymerase (RNA) III (DNA directed) polypeptide C |
| 3 | gene | 96.72427 | 96.72440 | negative | MGI_C57BL6J_5452358 | Gm22581 | ENSEMBL:ENSMUSG00000065669 | MGI:5452358 | snoRNA gene | predicted gene, 22581 |
| 3 | gene | 96.72453 | 96.72532 | negative | MGI_C57BL6J_5662920 | Gm42783 | ENSEMBL:ENSMUSG00000104795 | MGI:5662920 | unclassified gene | predicted gene 42783 |
| 3 | gene | 96.72759 | 96.79164 | positive | MGI_C57BL6J_1915095 | Rnf115 | NCBI_Gene:67845,ENSEMBL:ENSMUSG00000028098 | MGI:1915095 | protein coding gene | ring finger protein 115 |
| 3 | gene | 96.78432 | 96.78485 | positive | MGI_C57BL6J_5662919 | Gm42782 | ENSEMBL:ENSMUSG00000105976 | MGI:5662919 | unclassified gene | predicted gene 42782 |
| 3 | gene | 96.79876 | 96.82935 | negative | MGI_C57BL6J_1860383 | Cd160 | NCBI_Gene:54215,ENSEMBL:ENSMUSG00000038304 | MGI:1860383 | protein coding gene | CD160 antigen |
| 3 | gene | 96.82928 | 96.87093 | positive | MGI_C57BL6J_1928901 | Pdzk1 | NCBI_Gene:59020,ENSEMBL:ENSMUSG00000038298 | MGI:1928901 | protein coding gene | PDZ domain containing 1 |
| 3 | gene | 96.82983 | 96.83210 | negative | MGI_C57BL6J_1914833 | 4930442L01Rik | NCBI_Gene:67583,ENSEMBL:ENSMUSG00000087223 | MGI:1914833 | lncRNA gene | RIKEN cDNA 4930442L01 gene |
| 3 | gene | 96.86828 | 96.90535 | negative | MGI_C57BL6J_1914799 | Gpr89 | NCBI_Gene:67549,ENSEMBL:ENSMUSG00000028096 | MGI:1914799 | protein coding gene | G protein-coupled receptor 89 |
| 3 | gene | 96.90469 | 97.07742 | positive | MGI_C57BL6J_95716 | Gja5 | NCBI_Gene:14613,ENSEMBL:ENSMUSG00000057123 | MGI:95716 | protein coding gene | gap junction protein, alpha 5 |
| 3 | gene | 96.91357 | 96.92605 | negative | MGI_C57BL6J_99953 | Gja8 | NCBI_Gene:14616,ENSEMBL:ENSMUSG00000049908 | MGI:99953 | protein coding gene | gap junction protein, alpha 8 |
| 3 | gene | 96.99872 | 97.00077 | negative | MGI_C57BL6J_1925330 | 9230114J08Rik | ENSEMBL:ENSMUSG00000105385 | MGI:1925330 | lncRNA gene | RIKEN cDNA 9230114J08 gene |
| 3 | gene | 97.04666 | 97.09279 | negative | MGI_C57BL6J_1923138 | 4930573H18Rik | NCBI_Gene:75888,ENSEMBL:ENSMUSG00000104766 | MGI:1923138 | lncRNA gene | RIKEN cDNA 4930573H18 gene |
| 3 | gene | 97.07415 | 97.07602 | positive | MGI_C57BL6J_5590279 | Gm31120 | NCBI_Gene:102633248 | MGI:5590279 | lncRNA gene | predicted gene, 31120 |
| 3 | gene | 97.10109 | 97.10296 | positive | MGI_C57BL6J_5590342 | Gm31183 | NCBI_Gene:102633334 | MGI:5590342 | lncRNA gene | predicted gene, 31183 |
| 3 | pseudogene | 97.12551 | 97.12611 | negative | MGI_C57BL6J_5663359 | Gm43222 | ENSEMBL:ENSMUSG00000106324 | MGI:5663359 | pseudogene | predicted gene 43222 |
| 3 | gene | 97.14055 | 97.14066 | negative | MGI_C57BL6J_5454212 | Gm24435 | ENSEMBL:ENSMUSG00000088511 | MGI:5454212 | snRNA gene | predicted gene, 24435 |
| 3 | gene | 97.15808 | 97.15816 | positive | MGI_C57BL6J_4413852 | n-TEctc12 | NCBI_Gene:102467522 | MGI:4413852 | tRNA gene | nuclear encoded tRNA glutamic acid 12 (anticodon CTC) |
| 3 | gene | 97.15875 | 97.17730 | positive | MGI_C57BL6J_1931010 | Acp6 | NCBI_Gene:66659,ENSEMBL:ENSMUSG00000028093 | MGI:1931010 | protein coding gene | acid phosphatase 6, lysophosphatidic |
| 3 | gene | 97.20366 | 97.29808 | negative | MGI_C57BL6J_1924828 | Bcl9 | NCBI_Gene:77578,ENSEMBL:ENSMUSG00000038256 | MGI:1924828 | protein coding gene | B cell CLL/lymphoma 9 |
| 3 | gene | 97.26235 | 97.26565 | positive | MGI_C57BL6J_5662645 | Gm42508 | ENSEMBL:ENSMUSG00000106536 | MGI:5662645 | unclassified gene | predicted gene 42508 |
| 3 | gene | 97.27063 | 97.27162 | negative | MGI_C57BL6J_1924948 | 9230110I02Rik | NA | NA | unclassified non-coding RNA gene | RIKEN cDNA 9230110I02 gene |
| 3 | gene | 97.27388 | 97.27506 | negative | MGI_C57BL6J_5663210 | Gm43073 | ENSEMBL:ENSMUSG00000104745 | MGI:5663210 | unclassified gene | predicted gene 43073 |
| 3 | gene | 97.33042 | 97.33295 | positive | MGI_C57BL6J_5622986 | Gm40101 | NCBI_Gene:105244507 | MGI:5622986 | lncRNA gene | predicted gene, 40101 |
| 3 | gene | 97.40745 | 97.42202 | negative | MGI_C57BL6J_3031236 | Olfr1402 | NCBI_Gene:258272,ENSEMBL:ENSMUSG00000051392 | MGI:3031236 | protein coding gene | olfactory receptor 1402 |
| 3 | gene | 97.54630 | 97.57259 | positive | MGI_C57BL6J_5622987 | Gm40102 | NCBI_Gene:105244508 | MGI:5622987 | lncRNA gene | predicted gene, 40102 |
| 3 | gene | 97.56074 | 97.61020 | negative | MGI_C57BL6J_1915308 | Chd1l | NCBI_Gene:68058,ENSEMBL:ENSMUSG00000028089 | MGI:1915308 | protein coding gene | chromodomain helicase DNA binding protein 1-like |
| 3 | gene | 97.60993 | 97.61384 | positive | MGI_C57BL6J_5663212 | Gm43075 | ENSEMBL:ENSMUSG00000105004 | MGI:5663212 | unclassified gene | predicted gene 43075 |
| 3 | gene | 97.61019 | 97.65529 | positive | MGI_C57BL6J_1310004 | Fmo5 | NCBI_Gene:14263,ENSEMBL:ENSMUSG00000028088 | MGI:1310004 | protein coding gene | flavin containing monooxygenase 5 |
| 3 | gene | 97.65819 | 97.67381 | positive | MGI_C57BL6J_1336185 | Prkab2 | NCBI_Gene:108097,ENSEMBL:ENSMUSG00000038205 | MGI:1336185 | protein coding gene | protein kinase, AMP-activated, beta 2 non-catalytic subunit |
| 3 | gene | 97.67303 | 97.67469 | negative | MGI_C57BL6J_4937285 | Gm17651 | ENSEMBL:ENSMUSG00000090441 | MGI:4937285 | protein coding gene | predicted gene, 17651 |
| 3 | gene | 97.68034 | 97.68206 | positive | MGI_C57BL6J_5590401 | Gm31242 | NCBI_Gene:102633409 | MGI:5590401 | lncRNA gene | predicted gene, 31242 |
| 3 | gene | 97.68146 | 97.68153 | positive | MGI_C57BL6J_4413826 | n-TQctg2 | NCBI_Gene:102467291 | MGI:4413826 | tRNA gene | nuclear encoded tRNA glutamine 2 (anticodon CTG) |
| 3 | gene | 97.68982 | 97.88871 | negative | MGI_C57BL6J_1891434 | Pde4dip | NCBI_Gene:83679,ENSEMBL:ENSMUSG00000038170 | MGI:1891434 | protein coding gene | phosphodiesterase 4D interacting protein (myomegalin) |
| 3 | gene | 97.69051 | 97.69056 | negative | MGI_C57BL6J_5530881 | Mir7225 | miRBase:MI0023720,NCBI_Gene:102466229,ENSEMBL:ENSMUSG00000098381 | MGI:5530881 | miRNA gene | microRNA 7225 |
| 3 | gene | 97.73034 | 97.73714 | positive | MGI_C57BL6J_5590464 | Gm31305 | NCBI_Gene:102633492,ENSEMBL:ENSMUSG00000105245 | MGI:5590464 | lncRNA gene | predicted gene, 31305 |
| 3 | gene | 97.74903 | 97.75107 | positive | MGI_C57BL6J_5826453 | Gm46816 | NCBI_Gene:108168886 | MGI:5826453 | lncRNA gene | predicted gene, 46816 |
| 3 | gene | 97.76152 | 97.76277 | negative | MGI_C57BL6J_2444459 | A630078F23Rik | NA | NA | unclassified gene | RIKEN cDNA A630078F23 gene |
| 3 | pseudogene | 97.78707 | 97.78785 | negative | MGI_C57BL6J_5011008 | Gm18823 | NCBI_Gene:100417783 | MGI:5011008 | pseudogene | predicted gene, 18823 |
| 3 | gene | 97.89223 | 97.90109 | negative | MGI_C57BL6J_3802103 | Gm15999 | ENSEMBL:ENSMUSG00000086557 | MGI:3802103 | lncRNA gene | predicted gene 15999 |
| 3 | gene | 97.90119 | 97.92328 | positive | MGI_C57BL6J_1338759 | Sec22b | NCBI_Gene:20333,ENSEMBL:ENSMUSG00000027879 | MGI:1338759 | protein coding gene | SEC22 homolog B, vesicle trafficking protein |
| 3 | gene | 97.93017 | 97.96702 | positive | MGI_C57BL6J_3648209 | Gm5544 | NCBI_Gene:433632,ENSEMBL:ENSMUSG00000074388 | MGI:3648209 | lncRNA gene | predicted gene 5544 |
| 3 | gene | 98.01353 | 98.15037 | positive | MGI_C57BL6J_97364 | Notch2 | NCBI_Gene:18129,ENSEMBL:ENSMUSG00000027878 | MGI:97364 | protein coding gene | notch 2 |
| 3 | gene | 98.02231 | 98.02688 | negative | MGI_C57BL6J_5622988 | Gm40103 | NCBI_Gene:105244509 | MGI:5622988 | lncRNA gene | predicted gene, 40103 |
| 3 | gene | 98.03069 | 98.03581 | positive | MGI_C57BL6J_5662956 | Gm42819 | ENSEMBL:ENSMUSG00000105728 | MGI:5662956 | unclassified gene | predicted gene 42819 |
| 3 | gene | 98.12159 | 98.12465 | positive | MGI_C57BL6J_5662957 | Gm42820 | ENSEMBL:ENSMUSG00000106427 | MGI:5662957 | unclassified gene | predicted gene 42820 |
| 3 | gene | 98.16063 | 98.16417 | positive | MGI_C57BL6J_1918328 | Adam30 | NCBI_Gene:71078,ENSEMBL:ENSMUSG00000043468 | MGI:1918328 | protein coding gene | a disintegrin and metallopeptidase domain 30 |
| 3 | pseudogene | 98.18997 | 98.19112 | negative | MGI_C57BL6J_3642065 | Gm9771 | NCBI_Gene:622561,ENSEMBL:ENSMUSG00000106486 | MGI:3642065 | pseudogene | predicted gene 9771 |
| 3 | gene | 98.20235 | 98.20676 | positive | MGI_C57BL6J_5622989 | Gm40104 | NCBI_Gene:105244510 | MGI:5622989 | lncRNA gene | predicted gene, 40104 |
| 3 | gene | 98.22214 | 98.23675 | positive | MGI_C57BL6J_1914959 | Reg4 | NCBI_Gene:67709,ENSEMBL:ENSMUSG00000027876 | MGI:1914959 | protein coding gene | regenerating islet-derived family, member 4 |
| 3 | pseudogene | 98.24860 | 98.24909 | positive | MGI_C57BL6J_5662958 | Gm42821 | ENSEMBL:ENSMUSG00000104842 | MGI:5662958 | pseudogene | predicted gene 42821 |
| 3 | gene | 98.28043 | 98.31074 | positive | MGI_C57BL6J_101939 | Hmgcs2 | NCBI_Gene:15360,ENSEMBL:ENSMUSG00000027875 | MGI:101939 | protein coding gene | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 2 |
| 3 | gene | 98.28699 | 98.29006 | positive | MGI_C57BL6J_5663326 | Gm43189 | ENSEMBL:ENSMUSG00000105909 | MGI:5663326 | lncRNA gene | predicted gene 43189 |
| 3 | gene | 98.31317 | 98.33999 | negative | MGI_C57BL6J_1355330 | Phgdh | NCBI_Gene:236539,ENSEMBL:ENSMUSG00000053398 | MGI:1355330 | protein coding gene | 3-phosphoglycerate dehydrogenase |
| 3 | gene | 98.35151 | 98.35497 | positive | MGI_C57BL6J_5590579 | Gm31420 | NCBI_Gene:102633639 | MGI:5590579 | lncRNA gene | predicted gene, 31420 |
| 3 | gene | 98.38134 | 98.75381 | positive | MGI_C57BL6J_2139736 | Zfp697 | NCBI_Gene:242109,ENSEMBL:ENSMUSG00000050064 | MGI:2139736 | protein coding gene | zinc finger protein 697 |
| 3 | gene | 98.44567 | 98.45714 | negative | MGI_C57BL6J_3782634 | Gm4450 | NCBI_Gene:100043461,ENSEMBL:ENSMUSG00000090817 | MGI:3782634 | protein coding gene | predicted gene 4450 |
| 3 | gene | 98.44720 | 98.44898 | negative | MGI_C57BL6J_5663327 | Gm43190 | ENSEMBL:ENSMUSG00000106107 | MGI:5663327 | lncRNA gene | predicted gene 43190 |
| 3 | pseudogene | 98.46915 | 98.47322 | negative | MGI_C57BL6J_5011395 | Gm19210 | NCBI_Gene:100418432,ENSEMBL:ENSMUSG00000106129 | MGI:5011395 | pseudogene | predicted gene, 19210 |
| 3 | gene | 98.49907 | 98.51058 | negative | MGI_C57BL6J_96236 | Hsd3b4 | NCBI_Gene:15495,ENSEMBL:ENSMUSG00000095143 | MGI:96236 | protein coding gene | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 4 |
| 3 | pseudogene | 98.52274 | 98.52669 | negative | MGI_C57BL6J_5011396 | Gm19211 | NCBI_Gene:100418433,ENSEMBL:ENSMUSG00000105696 | MGI:5011396 | pseudogene | predicted gene, 19211 |
| 3 | gene | 98.55254 | 98.58881 | negative | MGI_C57BL6J_3711284 | Gm10681 | NCBI_Gene:100043456,ENSEMBL:ENSMUSG00000095388 | MGI:3711284 | protein coding gene | predicted gene 10681 |
| 3 | pseudogene | 98.57976 | 98.58057 | negative | MGI_C57BL6J_3648707 | Gm5853 | NCBI_Gene:545555,ENSEMBL:ENSMUSG00000105641 | MGI:3648707 | pseudogene | predicted gene 5853 |
| 3 | pseudogene | 98.58115 | 98.58829 | negative | MGI_C57BL6J_3780086 | Gm9678 | NCBI_Gene:676433,ENSEMBL:ENSMUSG00000106456 | MGI:3780086 | pseudogene | predicted gene 9678 |
| 3 | gene | 98.61863 | 98.64553 | negative | MGI_C57BL6J_104645 | Hsd3b5 | NCBI_Gene:15496,ENSEMBL:ENSMUSG00000038092 | MGI:104645 | protein coding gene | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 5 |
| 3 | pseudogene | 98.66985 | 98.67833 | negative | MGI_C57BL6J_3651917 | Gm12399 | NCBI_Gene:100418434,ENSEMBL:ENSMUSG00000081919 | MGI:3651917 | pseudogene | predicted gene 12399 |
| 3 | pseudogene | 98.68082 | 98.68824 | negative | MGI_C57BL6J_3651916 | Gm12400 | NCBI_Gene:668543,ENSEMBL:ENSMUSG00000083331 | MGI:3651916 | pseudogene | predicted gene 12400 |
| 3 | gene | 98.70925 | 98.72459 | negative | MGI_C57BL6J_96234 | Hsd3b2 | NCBI_Gene:15493,ENSEMBL:ENSMUSG00000063730 | MGI:96234 | protein coding gene | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 2 |
| 3 | gene | 98.74148 | 98.76515 | negative | MGI_C57BL6J_96235 | Hsd3b3 | NCBI_Gene:15494,ENSEMBL:ENSMUSG00000062410 | MGI:96235 | protein coding gene | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 3 |
| 3 | gene | 98.80550 | 98.81444 | negative | MGI_C57BL6J_109598 | Hsd3b6 | NCBI_Gene:15497,ENSEMBL:ENSMUSG00000027869 | MGI:109598 | protein coding gene | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 6 |
| 3 | pseudogene | 98.81808 | 98.82002 | negative | MGI_C57BL6J_3650847 | Gm12427 | ENSEMBL:ENSMUSG00000083222 | MGI:3650847 | pseudogene | predicted gene 12427 |
| 3 | pseudogene | 98.82085 | 98.82266 | positive | MGI_C57BL6J_3650848 | Gm12428 | NCBI_Gene:545556,ENSEMBL:ENSMUSG00000082185 | MGI:3650848 | pseudogene | predicted gene 12428 |
| 3 | gene | 98.84235 | 98.85979 | negative | MGI_C57BL6J_96233 | Hsd3b1 | NCBI_Gene:15492,ENSEMBL:ENSMUSG00000027871 | MGI:96233 | protein coding gene | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 |
| 3 | pseudogene | 98.84287 | 98.84304 | negative | MGI_C57BL6J_3649672 | Gm12431 | ENSEMBL:ENSMUSG00000082089 | MGI:3649672 | pseudogene | predicted gene 12431 |
| 3 | pseudogene | 98.84634 | 98.84652 | negative | MGI_C57BL6J_3649670 | Gm12432 | ENSEMBL:ENSMUSG00000083577 | MGI:3649670 | pseudogene | predicted gene 12432 |
| 3 | pseudogene | 98.84651 | 98.84676 | positive | MGI_C57BL6J_3649669 | Gm12433 | ENSEMBL:ENSMUSG00000082589 | MGI:3649669 | pseudogene | predicted gene 12433 |
| 3 | gene | 98.87452 | 98.89538 | negative | MGI_C57BL6J_96012 | Hao2 | NCBI_Gene:56185,ENSEMBL:ENSMUSG00000027870 | MGI:96012 | protein coding gene | hydroxyacid oxidase 2 |
| 3 | gene | 98.89783 | 98.93252 | negative | MGI_C57BL6J_5591039 | Gm31880 | NCBI_Gene:102634250 | MGI:5591039 | lncRNA gene | predicted gene, 31880 |
| 3 | gene | 98.93765 | 98.94126 | negative | MGI_C57BL6J_1917808 | 5730437C11Rik | ENSEMBL:ENSMUSG00000085121 | MGI:1917808 | lncRNA gene | RIKEN cDNA 5730437C11 gene |
| 3 | gene | 99.04982 | 99.05374 | positive | MGI_C57BL6J_3649266 | Gm12440 | NCBI_Gene:102634753,ENSEMBL:ENSMUSG00000086833 | MGI:3649266 | lncRNA gene | predicted gene 12440 |
| 3 | pseudogene | 99.10057 | 99.10115 | negative | MGI_C57BL6J_3649265 | Gm12441 | NCBI_Gene:102631982,ENSEMBL:ENSMUSG00000083888 | MGI:3649265 | pseudogene | predicted gene 12441 |
| 3 | gene | 99.11096 | 99.11570 | positive | MGI_C57BL6J_5622990 | Gm40105 | NCBI_Gene:105244511 | MGI:5622990 | lncRNA gene | predicted gene, 40105 |
| 3 | gene | 99.11316 | 99.11328 | negative | MGI_C57BL6J_5452118 | Gm22341 | ENSEMBL:ENSMUSG00000065709 | MGI:5452118 | snoRNA gene | predicted gene, 22341 |
| 3 | gene | 99.12419 | 99.12430 | negative | MGI_C57BL6J_5453993 | Gm24216 | ENSEMBL:ENSMUSG00000088884 | MGI:5453993 | snoRNA gene | predicted gene, 24216 |
| 3 | gene | 99.14062 | 99.23919 | positive | MGI_C57BL6J_1917810 | Wars2 | NCBI_Gene:70560,ENSEMBL:ENSMUSG00000004233 | MGI:1917810 | protein coding gene | tryptophanyl tRNA synthetase 2 (mitochondrial) |
| 3 | gene | 99.14460 | 99.14895 | positive | MGI_C57BL6J_5663161 | Gm43024 | ENSEMBL:ENSMUSG00000105472 | MGI:5663161 | lncRNA gene | predicted gene 43024 |
| 3 | pseudogene | 99.18075 | 99.18139 | negative | MGI_C57BL6J_5011167 | Gm18982 | NCBI_Gene:100418070,ENSEMBL:ENSMUSG00000105798 | MGI:5011167 | pseudogene | predicted gene, 18982 |
| 3 | gene | 99.19934 | 99.19945 | negative | MGI_C57BL6J_5531126 | Mir6481 | miRBase:MI0022051,NCBI_Gene:102465230,ENSEMBL:ENSMUSG00000098918 | MGI:5531126 | miRNA gene | microRNA 6481 |
| 3 | gene | 99.20642 | 99.20988 | positive | MGI_C57BL6J_5662854 | Gm42717 | ENSEMBL:ENSMUSG00000106197 | MGI:5662854 | lncRNA gene | predicted gene 42717 |
| 3 | gene | 99.23691 | 99.23912 | positive | MGI_C57BL6J_5621353 | Gm38468 | NCBI_Gene:102634827 | MGI:5621353 | lncRNA gene | predicted gene, 38468 |
| 3 | gene | 99.24035 | 99.35426 | positive | MGI_C57BL6J_1277234 | Tbx15 | NCBI_Gene:21384,ENSEMBL:ENSMUSG00000027868 | MGI:1277234 | protein coding gene | T-box 15 |
| 3 | gene | 99.24066 | 99.24189 | negative | MGI_C57BL6J_5826454 | Gm46817 | NCBI_Gene:108168887 | MGI:5826454 | lncRNA gene | predicted gene, 46817 |
| 3 | gene | 99.25470 | 99.26129 | negative | MGI_C57BL6J_5663257 | Gm43120 | ENSEMBL:ENSMUSG00000105689 | MGI:5663257 | lncRNA gene | predicted gene 43120 |
| 3 | pseudogene | 99.46359 | 99.46382 | negative | MGI_C57BL6J_3650542 | Gm12449 | ENSEMBL:ENSMUSG00000081183 | MGI:3650542 | pseudogene | predicted gene 12449 |
| 3 | pseudogene | 99.50037 | 99.50261 | positive | MGI_C57BL6J_96905 | M6pr-ps | NCBI_Gene:17114,ENSEMBL:ENSMUSG00000080832 | MGI:96905 | pseudogene | mannose-6-phosphate receptor, pseudogene |
| 3 | pseudogene | 99.54441 | 99.54506 | negative | MGI_C57BL6J_3651230 | Gm12448 | ENSEMBL:ENSMUSG00000083938 | MGI:3651230 | pseudogene | predicted gene 12448 |
| 3 | gene | 99.70774 | 99.70785 | negative | MGI_C57BL6J_5452282 | Gm22505 | ENSEMBL:ENSMUSG00000064890 | MGI:5452282 | snRNA gene | predicted gene, 22505 |
| 3 | gene | 99.88541 | 100.14332 | positive | MGI_C57BL6J_1921612 | Spag17 | NCBI_Gene:74362,ENSEMBL:ENSMUSG00000027867 | MGI:1921612 | protein coding gene | sperm associated antigen 17 |
| 3 | pseudogene | 100.00832 | 100.00873 | positive | MGI_C57BL6J_3649699 | Rpl36-ps7 | NCBI_Gene:623045,ENSEMBL:ENSMUSG00000080754 | MGI:3649699 | pseudogene | ribosomal protein L36, pseudogene 7 |
| 3 | pseudogene | 100.07848 | 100.07864 | negative | MGI_C57BL6J_3650850 | Tra2b-ps | ENSEMBL:ENSMUSG00000082763 | MGI:3650850 | pseudogene | transformer 2 beta, pseudogene |
| 3 | gene | 100.10068 | 100.10689 | negative | MGI_C57BL6J_2444368 | Spag17os | ENSEMBL:ENSMUSG00000085244 | MGI:2444368 | antisense lncRNA gene | sperm associated antigen 17, opposite strand |
| 3 | gene | 100.13818 | 100.16241 | negative | MGI_C57BL6J_2443143 | Wdr3 | NCBI_Gene:269470,ENSEMBL:ENSMUSG00000033285 | MGI:2443143 | protein coding gene | WD repeat domain 3 |
| 3 | gene | 100.16238 | 100.20699 | positive | MGI_C57BL6J_1338001 | Gdap2 | NCBI_Gene:14547,ENSEMBL:ENSMUSG00000027865 | MGI:1338001 | protein coding gene | ganglioside-induced differentiation-associated-protein 2 |
| 3 | gene | 100.30329 | 100.30336 | positive | MGI_C57BL6J_5455579 | Gm25802 | ENSEMBL:ENSMUSG00000064573 | MGI:5455579 | snoRNA gene | predicted gene, 25802 |
| 3 | pseudogene | 100.41355 | 100.41478 | positive | MGI_C57BL6J_3650645 | Gm12464 | NCBI_Gene:195514,ENSEMBL:ENSMUSG00000081440 | MGI:3650645 | pseudogene | predicted gene 12464 |
| 3 | gene | 100.41808 | 100.42242 | positive | MGI_C57BL6J_5591630 | Gm32471 | NCBI_Gene:102635032 | MGI:5591630 | lncRNA gene | predicted gene, 32471 |
| 3 | gene | 100.42641 | 100.43775 | positive | MGI_C57BL6J_1921056 | 4930406D18Rik | NCBI_Gene:73806,ENSEMBL:ENSMUSG00000086923 | MGI:1921056 | lncRNA gene | RIKEN cDNA 4930406D18 gene |
| 3 | gene | 100.43783 | 100.43914 | positive | MGI_C57BL6J_5663258 | Gm43121 | ENSEMBL:ENSMUSG00000106639 | MGI:5663258 | lncRNA gene | predicted gene 43121 |
| 3 | gene | 100.45163 | 100.48932 | negative | MGI_C57BL6J_1921895 | Tent5c | NCBI_Gene:74645,ENSEMBL:ENSMUSG00000044468 | MGI:1921895 | protein coding gene | terminal nucleotidyltransferase 5C |
| 3 | gene | 100.45181 | 100.45958 | negative | MGI_C57BL6J_5621415 | Gm38530 | NCBI_Gene:102641070 | MGI:5621415 | lncRNA gene | predicted gene, 38530 |
| 3 | gene | 100.48932 | 100.53121 | positive | MGI_C57BL6J_3649378 | Gm12474 | NCBI_Gene:545557,ENSEMBL:ENSMUSG00000053957 | MGI:3649378 | lncRNA gene | predicted gene 12474 |
| 3 | gene | 100.49536 | 100.54135 | negative | MGI_C57BL6J_5622991 | Gm40106 | NCBI_Gene:105244512 | MGI:5622991 | lncRNA gene | predicted gene, 40106 |
| 3 | gene | 100.49743 | 100.50012 | positive | MGI_C57BL6J_5663005 | Gm42868 | ENSEMBL:ENSMUSG00000104943 | MGI:5663005 | lncRNA gene | predicted gene 42868 |
| 3 | gene | 100.56220 | 100.68550 | negative | MGI_C57BL6J_104676 | Man1a2 | NCBI_Gene:17156,ENSEMBL:ENSMUSG00000008763 | MGI:104676 | protein coding gene | mannosidase, alpha, class 1A, member 2 |
| 3 | gene | 100.58231 | 100.58753 | negative | MGI_C57BL6J_5663603 | Gm43466 | ENSEMBL:ENSMUSG00000105824 | MGI:5663603 | lncRNA gene | predicted gene 43466 |
| 3 | gene | 100.62490 | 100.62702 | negative | MGI_C57BL6J_5663006 | Gm42869 | ENSEMBL:ENSMUSG00000105137 | MGI:5663006 | lncRNA gene | predicted gene 42869 |
| 3 | gene | 100.62730 | 100.63094 | negative | MGI_C57BL6J_5663599 | Gm43462 | ENSEMBL:ENSMUSG00000105561 | MGI:5663599 | lncRNA gene | predicted gene 43462 |
| 3 | gene | 100.67829 | 100.68009 | negative | MGI_C57BL6J_5663600 | Gm43463 | ENSEMBL:ENSMUSG00000106184 | MGI:5663600 | lncRNA gene | predicted gene 43463 |
| 3 | pseudogene | 100.68027 | 100.68055 | positive | MGI_C57BL6J_3651876 | Gm12485 | ENSEMBL:ENSMUSG00000083842 | MGI:3651876 | pseudogene | predicted gene 12485 |
| 3 | gene | 100.69271 | 100.70008 | positive | MGI_C57BL6J_5663601 | Gm43464 | ENSEMBL:ENSMUSG00000106227 | MGI:5663601 | lncRNA gene | predicted gene 43464 |
| 3 | gene | 100.73845 | 100.73871 | negative | MGI_C57BL6J_5453242 | Gm23465 | ENSEMBL:ENSMUSG00000087759 | MGI:5453242 | unclassified non-coding RNA gene | predicted gene, 23465 |
| 3 | gene | 100.81154 | 100.81775 | negative | MGI_C57BL6J_5622992 | Gm40107 | NCBI_Gene:105244513 | MGI:5622992 | lncRNA gene | predicted gene, 40107 |
| 3 | gene | 100.82040 | 100.82702 | negative | MGI_C57BL6J_5591879 | Gm32720 | NCBI_Gene:102635361 | MGI:5591879 | lncRNA gene | predicted gene, 32720 |
| 3 | gene | 100.82546 | 100.89692 | positive | MGI_C57BL6J_3039619 | Vtcn1 | NCBI_Gene:242122,ENSEMBL:ENSMUSG00000051076 | MGI:3039619 | protein coding gene | V-set domain containing T cell activation inhibitor 1 |
| 3 | gene | 100.90894 | 100.91522 | positive | MGI_C57BL6J_5622993 | Gm40108 | NCBI_Gene:105244514 | MGI:5622993 | lncRNA gene | predicted gene, 40108 |
| 3 | gene | 100.92220 | 100.93693 | positive | MGI_C57BL6J_1918187 | Trim45 | NCBI_Gene:229644,ENSEMBL:ENSMUSG00000033233 | MGI:1918187 | protein coding gene | tripartite motif-containing 45 |
| 3 | gene | 100.93886 | 100.96972 | negative | MGI_C57BL6J_1921294 | Ttf2 | NCBI_Gene:74044,ENSEMBL:ENSMUSG00000033222 | MGI:1921294 | protein coding gene | transcription termination factor, RNA polymerase II |
| 3 | pseudogene | 100.95544 | 100.95594 | positive | MGI_C57BL6J_3650947 | Gm12488 | ENSEMBL:ENSMUSG00000082000 | MGI:3650947 | pseudogene | predicted gene 12488 |
| 3 | gene | 100.99353 | 101.02956 | negative | MGI_C57BL6J_2685862 | Cd101 | NCBI_Gene:630146,ENSEMBL:ENSMUSG00000086564 | MGI:2685862 | protein coding gene | CD101 antigen |
| 3 | gene | 101.04023 | 101.11075 | negative | MGI_C57BL6J_1277114 | Ptgfrn | NCBI_Gene:19221,ENSEMBL:ENSMUSG00000027864 | MGI:1277114 | protein coding gene | prostaglandin F2 receptor negative regulator |
| 3 | gene | 101.08459 | 101.08739 | negative | MGI_C57BL6J_5663602 | Gm43465 | ENSEMBL:ENSMUSG00000105717 | MGI:5663602 | lncRNA gene | predicted gene 43465 |
| 3 | pseudogene | 101.20666 | 101.20748 | negative | MGI_C57BL6J_3650949 | Gm12486 | NCBI_Gene:668578,ENSEMBL:ENSMUSG00000082415 | MGI:3650949 | pseudogene | predicted gene 12486 |
| 3 | pseudogene | 101.23346 | 101.23412 | positive | MGI_C57BL6J_107797 | Tpt1-ps1 | NCBI_Gene:668583,ENSEMBL:ENSMUSG00000083546 | MGI:107797 | pseudogene | tumor protein, translationally-controlled, pseudogene 1 |
| 3 | pseudogene | 101.26092 | 101.26148 | positive | MGI_C57BL6J_3651827 | Gm12490 | NCBI_Gene:669017,ENSEMBL:ENSMUSG00000081569 | MGI:3651827 | pseudogene | predicted gene 12490 |
| 3 | gene | 101.27590 | 101.28794 | negative | MGI_C57BL6J_88320 | Cd2 | NCBI_Gene:12481,ENSEMBL:ENSMUSG00000027863 | MGI:88320 | protein coding gene | CD2 antigen |
| 3 | gene | 101.29100 | 101.29661 | negative | MGI_C57BL6J_5591962 | Gm32803 | NCBI_Gene:102635474 | MGI:5591962 | lncRNA gene | predicted gene, 32803 |
| 3 | pseudogene | 101.30643 | 101.30719 | negative | MGI_C57BL6J_3641762 | Gm10355 | NCBI_Gene:100043490,ENSEMBL:ENSMUSG00000072381 | MGI:3641762 | pseudogene | predicted gene 10355 |
| 3 | pseudogene | 101.31085 | 101.31128 | positive | MGI_C57BL6J_5663604 | Gm43467 | ENSEMBL:ENSMUSG00000106316 | MGI:5663604 | pseudogene | predicted gene 43467 |
| 3 | gene | 101.31591 | 101.32129 | positive | MGI_C57BL6J_5592007 | Gm32848 | NCBI_Gene:102635540 | MGI:5592007 | lncRNA gene | predicted gene, 32848 |
| 3 | gene | 101.33963 | 101.35461 | negative | MGI_C57BL6J_5592059 | Gm32900 | NCBI_Gene:102635610 | MGI:5592059 | lncRNA gene | predicted gene, 32900 |
| 3 | gene | 101.37708 | 101.46306 | positive | MGI_C57BL6J_1926158 | Igsf3 | NCBI_Gene:78908,ENSEMBL:ENSMUSG00000042035 | MGI:1926158 | protein coding gene | immunoglobulin superfamily, member 3 |
| 3 | gene | 101.40779 | 101.41000 | positive | MGI_C57BL6J_5663076 | Gm42939 | ENSEMBL:ENSMUSG00000104639 | MGI:5663076 | lncRNA gene | predicted gene 42939 |
| 3 | gene | 101.41154 | 101.41261 | positive | MGI_C57BL6J_5663075 | Gm42938 | ENSEMBL:ENSMUSG00000105611 | MGI:5663075 | lncRNA gene | predicted gene 42938 |
| 3 | gene | 101.42462 | 101.42694 | negative | MGI_C57BL6J_5662675 | Gm42538 | ENSEMBL:ENSMUSG00000106137 | MGI:5662675 | lncRNA gene | predicted gene 42538 |
| 3 | gene | 101.50193 | 101.50785 | negative | MGI_C57BL6J_5622994 | Gm40109 | NCBI_Gene:105244515 | MGI:5622994 | lncRNA gene | predicted gene, 40109 |
| 3 | gene | 101.51156 | 101.53625 | negative | MGI_C57BL6J_5592321 | Gm33162 | NCBI_Gene:102635957 | MGI:5592321 | lncRNA gene | predicted gene, 33162 |
| 3 | gene | 101.57622 | 101.60471 | negative | MGI_C57BL6J_88105 | Atp1a1 | NCBI_Gene:11928,ENSEMBL:ENSMUSG00000033161 | MGI:88105 | protein coding gene | ATPase, Na+/K+ transporting, alpha 1 polypeptide |
| 3 | gene | 101.58030 | 101.58106 | negative | MGI_C57BL6J_5663074 | Gm42937 | ENSEMBL:ENSMUSG00000104737 | MGI:5663074 | lncRNA gene | predicted gene 42937 |
| 3 | gene | 101.59581 | 101.59713 | negative | MGI_C57BL6J_5663078 | Gm42941 | ENSEMBL:ENSMUSG00000104693 | MGI:5663078 | lncRNA gene | predicted gene 42941 |
| 3 | gene | 101.63405 | 101.65080 | negative | MGI_C57BL6J_5592374 | Gm33215 | NCBI_Gene:102636027 | MGI:5592374 | lncRNA gene | predicted gene, 33215 |
| 3 | gene | 101.65909 | 101.66730 | negative | MGI_C57BL6J_5592424 | Gm33265 | NCBI_Gene:102636098 | MGI:5592424 | lncRNA gene | predicted gene, 33265 |
| 3 | gene | 101.69179 | 101.69963 | positive | MGI_C57BL6J_5663272 | Gm43135 | ENSEMBL:ENSMUSG00000106070 | MGI:5663272 | lncRNA gene | predicted gene 43135 |
| 3 | gene | 101.72714 | 101.73754 | negative | MGI_C57BL6J_5592468 | Gm33309 | NCBI_Gene:102636166 | MGI:5592468 | lncRNA gene | predicted gene, 33309 |
| 3 | gene | 101.74812 | 101.75001 | positive | MGI_C57BL6J_5663271 | Gm43134 | ENSEMBL:ENSMUSG00000105642 | MGI:5663271 | lncRNA gene | predicted gene 43134 |
| 3 | gene | 101.81308 | 101.84896 | negative | MGI_C57BL6J_2446273 | Mab21l3 | NCBI_Gene:242125,ENSEMBL:ENSMUSG00000044313 | MGI:2446273 | protein coding gene | mab-21-like 3 |
| 3 | gene | 101.82688 | 101.83499 | positive | MGI_C57BL6J_5592540 | Gm33381 | NCBI_Gene:102636267 | MGI:5592540 | lncRNA gene | predicted gene, 33381 |
| 3 | gene | 101.85444 | 101.92454 | negative | MGI_C57BL6J_3607704 | Slc22a15 | NCBI_Gene:242126,ENSEMBL:ENSMUSG00000033147 | MGI:3607704 | protein coding gene | solute carrier family 22 (organic anion/cation transporter), member 15 |
| 3 | gene | 102.01008 | 102.01549 | positive | MGI_C57BL6J_97324 | Nhlh2 | NCBI_Gene:18072,ENSEMBL:ENSMUSG00000048540 | MGI:97324 | protein coding gene | nescient helix loop helix 2 |
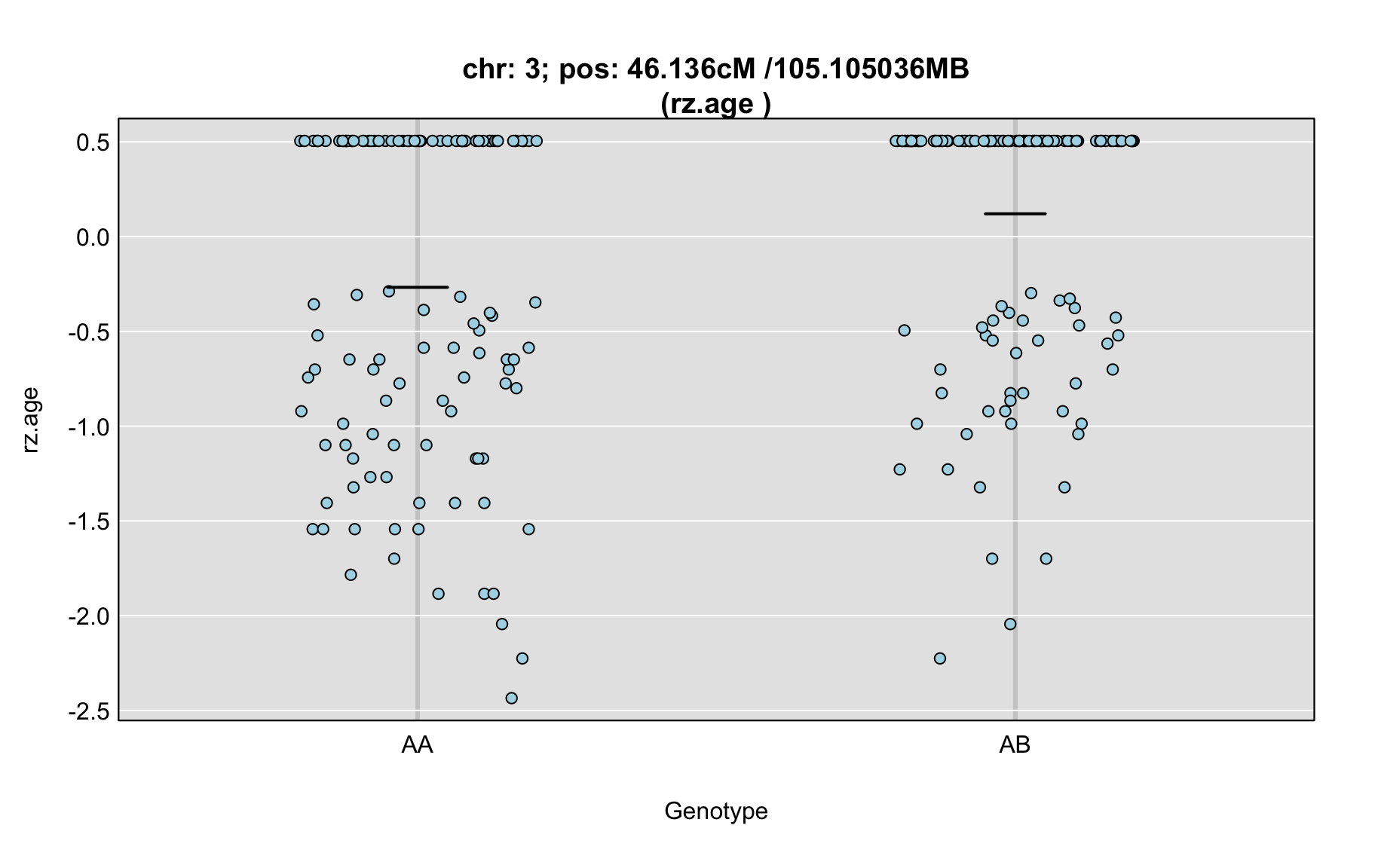
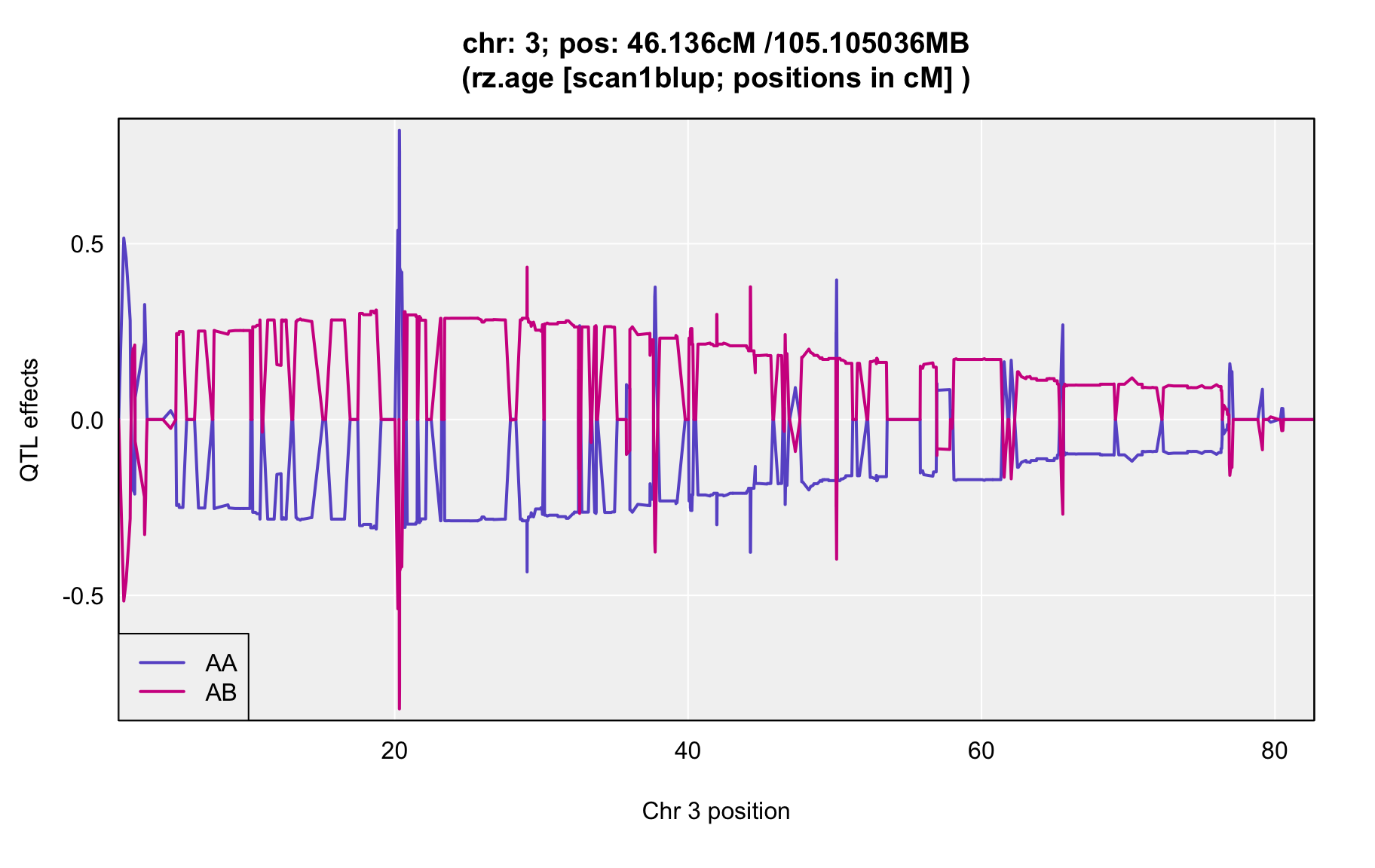
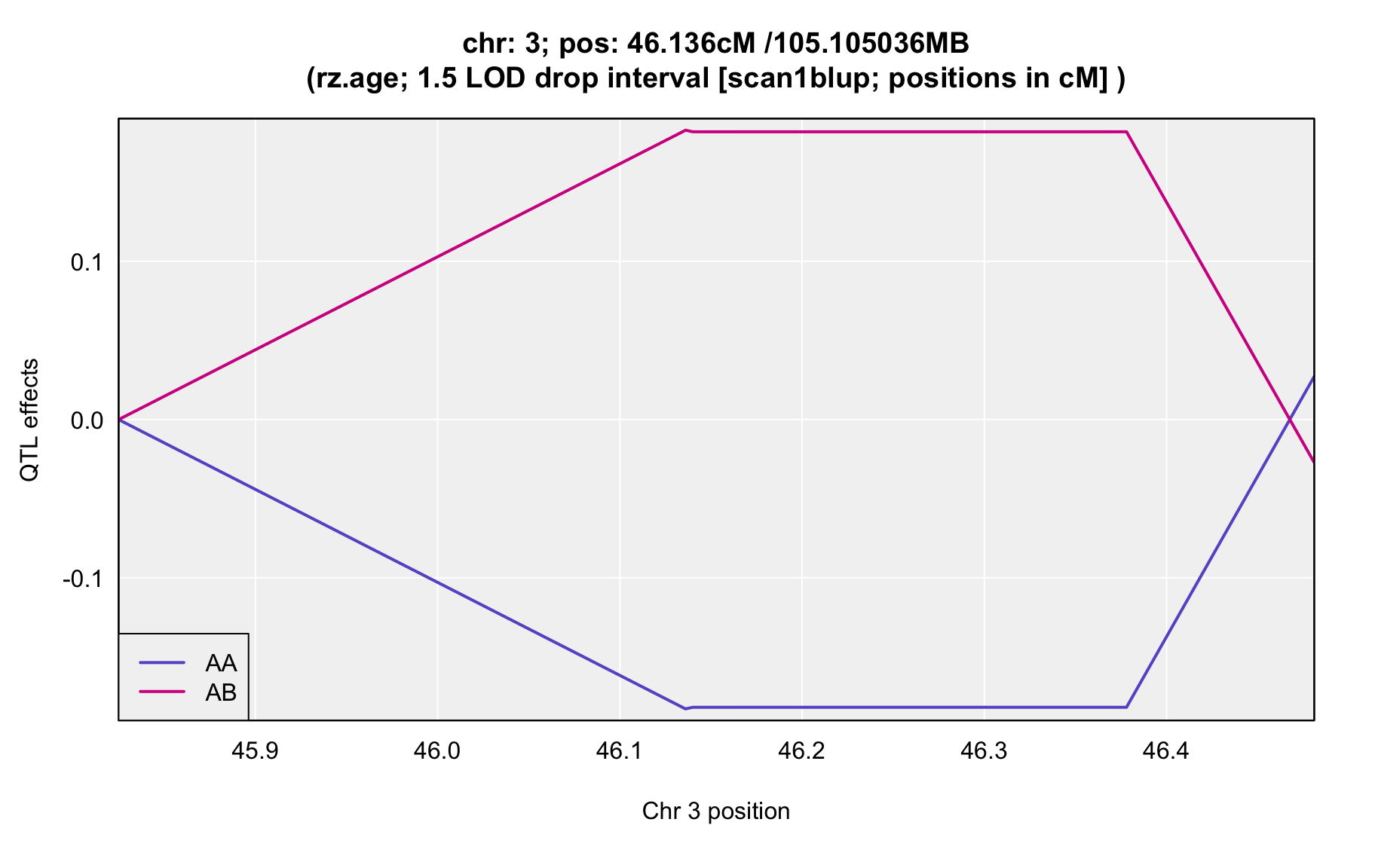
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 104.8228 | 104.8645 | negative | MGI_C57BL6J_106227 | Capza1 | NCBI_Gene:12340,ENSEMBL:ENSMUSG00000070372 | MGI:106227 | protein coding gene | capping protein (actin filament) muscle Z-line, alpha 1 |
| 3 | gene | 104.8263 | 104.8279 | negative | MGI_C57BL6J_5662795 | Gm42658 | ENSEMBL:ENSMUSG00000106251 | MGI:5662795 | lncRNA gene | predicted gene 42658 |
| 3 | gene | 104.8364 | 104.8382 | negative | MGI_C57BL6J_5662796 | Gm42659 | ENSEMBL:ENSMUSG00000105655 | MGI:5662796 | lncRNA gene | predicted gene 42659 |
| 3 | gene | 104.8640 | 104.9301 | positive | MGI_C57BL6J_2386964 | St7l | NCBI_Gene:229681,ENSEMBL:ENSMUSG00000045576 | MGI:2386964 | protein coding gene | suppression of tumorigenicity 7-like |
| 3 | pseudogene | 104.9159 | 104.9162 | negative | MGI_C57BL6J_3782710 | Gm4525 | NCBI_Gene:100043572,ENSEMBL:ENSMUSG00000106105 | MGI:3782710 | pseudogene | predicted gene 4525 |
| 3 | gene | 104.9448 | 104.9619 | negative | MGI_C57BL6J_1261834 | Wnt2b | NCBI_Gene:22414,ENSEMBL:ENSMUSG00000027840 | MGI:1261834 | protein coding gene | wingless-type MMTV integration site family, member 2B |
| 3 | gene | 104.9574 | 104.9584 | negative | MGI_C57BL6J_5663983 | Gm43846 | ENSEMBL:ENSMUSG00000105925 | MGI:5663983 | lncRNA gene | predicted gene 43846 |
| 3 | gene | 104.9816 | 104.9862 | positive | MGI_C57BL6J_5826456 | Gm46819 | NCBI_Gene:108168890 | MGI:5826456 | lncRNA gene | predicted gene, 46819 |
| 3 | gene | 104.9922 | 105.0040 | positive | MGI_C57BL6J_5623002 | Gm40117 | NCBI_Gene:105244523,ENSEMBL:ENSMUSG00000105355 | MGI:5623002 | lncRNA gene | predicted gene, 40117 |
| 3 | gene | 105.0019 | 105.0531 | negative | MGI_C57BL6J_1933137 | Cttnbp2nl | NCBI_Gene:80281,ENSEMBL:ENSMUSG00000062127 | MGI:1933137 | protein coding gene | CTTNBP2 N-terminal like |
| 3 | gene | 105.0081 | 105.0178 | positive | MGI_C57BL6J_5623001 | Gm40116 | NCBI_Gene:105244522 | MGI:5623001 | lncRNA gene | predicted gene, 40116 |
| 3 | gene | 105.0668 | 105.0788 | positive | MGI_C57BL6J_1922519 | 4930564D02Rik | NCBI_Gene:75269,ENSEMBL:ENSMUSG00000051788 | MGI:1922519 | protein coding gene | RIKEN cDNA 4930564D02 gene |
| 3 | gene | 105.0976 | 105.0983 | positive | MGI_C57BL6J_5593940 | Gm34781 | NA | NA | protein coding gene | predicted gene%2c 34781 |
| 3 | gene | 105.1074 | 105.1350 | positive | MGI_C57BL6J_5623003 | Gm40118 | NCBI_Gene:105244524 | MGI:5623003 | lncRNA gene | predicted gene, 40118 |
| 3 | gene | 105.3551 | 105.3683 | positive | MGI_C57BL6J_5593996 | Gm34837 | NCBI_Gene:102638225 | MGI:5593996 | lncRNA gene | predicted gene, 34837 |
| 3 | gene | 105.4416 | 105.4530 | negative | MGI_C57BL6J_1925885 | Kcnd3os | NCBI_Gene:78635,ENSEMBL:ENSMUSG00000074346 | MGI:1925885 | antisense lncRNA gene | potassium voltage-gated channel, Shal-related family, member 3, opposite strand |
| 3 | gene | 105.4522 | 105.6740 | positive | MGI_C57BL6J_1928743 | Kcnd3 | NCBI_Gene:56543,ENSEMBL:ENSMUSG00000040896 | MGI:1928743 | protein coding gene | potassium voltage-gated channel, Shal-related family, member 3 |
| 3 | gene | 105.5029 | 105.5048 | positive | MGI_C57BL6J_5663984 | Gm43847 | ENSEMBL:ENSMUSG00000105753 | MGI:5663984 | lncRNA gene | predicted gene 43847 |
| 3 | gene | 105.6783 | 105.6881 | negative | MGI_C57BL6J_1858415 | Ddx20 | NCBI_Gene:53975,ENSEMBL:ENSMUSG00000027905 | MGI:1858415 | protein coding gene | DEAD (Asp-Glu-Ala-Asp) box polypeptide 20 |
| 3 | gene | 105.7046 | 105.7208 | positive | MGI_C57BL6J_1923497 | Inka2 | NCBI_Gene:109050,ENSEMBL:ENSMUSG00000048458 | MGI:1923497 | protein coding gene | inka box actin regulator 2 |
| 3 | gene | 105.7259 | 105.7271 | negative | MGI_C57BL6J_5663985 | Gm43848 | ENSEMBL:ENSMUSG00000106317 | MGI:5663985 | lncRNA gene | predicted gene 43848 |
| 3 | gene | 105.7273 | 105.8014 | negative | MGI_C57BL6J_97852 | Rap1a | NCBI_Gene:109905,ENSEMBL:ENSMUSG00000068798 | MGI:97852 | protein coding gene | RAS-related protein 1a |
| 3 | gene | 105.7324 | 105.7339 | negative | MGI_C57BL6J_5663465 | Gm43328 | ENSEMBL:ENSMUSG00000105677 | MGI:5663465 | lncRNA gene | predicted gene 43328 |
| 3 | gene | 105.7670 | 105.7702 | negative | MGI_C57BL6J_5663466 | Gm43329 | ENSEMBL:ENSMUSG00000105842 | MGI:5663466 | lncRNA gene | predicted gene 43329 |
| 3 | gene | 105.7829 | 105.7850 | negative | MGI_C57BL6J_5663467 | Gm43330 | ENSEMBL:ENSMUSG00000106341 | MGI:5663467 | lncRNA gene | predicted gene 43330 |
| 3 | gene | 105.7876 | 105.7893 | negative | MGI_C57BL6J_5663468 | Gm43331 | ENSEMBL:ENSMUSG00000104910 | MGI:5663468 | lncRNA gene | predicted gene 43331 |
| 3 | gene | 105.8155 | 105.8174 | positive | MGI_C57BL6J_3648206 | Gm5547 | NCBI_Gene:433637,ENSEMBL:ENSMUSG00000104835 | MGI:3648206 | lncRNA gene | predicted gene 5547 |
| 3 | gene | 105.8218 | 105.8259 | negative | MGI_C57BL6J_5826475 | Gm46838 | NCBI_Gene:108168921 | MGI:5826475 | lncRNA gene | predicted gene, 46838 |
| 3 | gene | 105.8709 | 105.9089 | positive | MGI_C57BL6J_104847 | Adora3 | NCBI_Gene:11542,ENSEMBL:ENSMUSG00000000562 | MGI:104847 | protein coding gene | adenosine A3 receptor |
| 3 | gene | 105.8709 | 105.9240 | positive | MGI_C57BL6J_5604098 | Tmigd3 | NCBI_Gene:69296,ENSEMBL:ENSMUSG00000074344 | MGI:5604098 | protein coding gene | transmembrane and immunoglobulin domain containing 3 |
| 3 | gene | 105.9244 | 105.9329 | negative | MGI_C57BL6J_3588284 | I830077J02Rik | NCBI_Gene:433638,ENSEMBL:ENSMUSG00000074342 | MGI:3588284 | protein coding gene | RIKEN cDNA I830077J02 gene |
| 3 | gene | 105.9427 | 105.9602 | negative | MGI_C57BL6J_1100495 | Atp5pb | NCBI_Gene:11950,ENSEMBL:ENSMUSG00000000563 | MGI:1100495 | protein coding gene | ATP synthase peripheral stalk-membrane subunit b |
| 3 | gene | 105.9594 | 105.9700 | positive | MGI_C57BL6J_1917715 | Wdr77 | NCBI_Gene:70465,ENSEMBL:ENSMUSG00000000561 | MGI:1917715 | protein coding gene | WD repeat domain 77 |
| 3 | gene | 105.9595 | 105.9846 | positive | MGI_C57BL6J_5663027 | Gm42890 | ENSEMBL:ENSMUSG00000105852 | MGI:5663027 | lncRNA gene | predicted gene 42890 |
| 3 | gene | 105.9737 | 105.9874 | positive | MGI_C57BL6J_106661 | Ovgp1 | NCBI_Gene:12659,ENSEMBL:ENSMUSG00000074340 | MGI:106661 | protein coding gene | oviductal glycoprotein 1 |
| 3 | gene | 105.9970 | 106.0146 | negative | MGI_C57BL6J_1923670 | Pifo | NCBI_Gene:100503311,ENSEMBL:ENSMUSG00000010136 | MGI:1923670 | protein coding gene | primary cilia formation |
| 3 | gene | 106.0016 | 106.0175 | positive | MGI_C57BL6J_5662859 | Gm42722 | ENSEMBL:ENSMUSG00000106093 | MGI:5662859 | lncRNA gene | predicted gene 42722 |
| 3 | gene | 106.0079 | 106.0094 | negative | MGI_C57BL6J_5662858 | Gm42721 | ENSEMBL:ENSMUSG00000104592 | MGI:5662858 | lncRNA gene | predicted gene 42721 |
| 3 | gene | 106.0169 | 106.0331 | negative | MGI_C57BL6J_2676649 | Chil5 | NCBI_Gene:229687,ENSEMBL:ENSMUSG00000043873 | MGI:2676649 | protein coding gene | chitinase-like 5 |
| 3 | pseudogene | 106.0347 | 106.0351 | positive | MGI_C57BL6J_3782724 | Gm4540 | NCBI_Gene:100043594,ENSEMBL:ENSMUSG00000092072 | MGI:3782724 | pseudogene | predicted gene 4540 |
| 3 | pseudogene | 106.0417 | 106.0448 | negative | MGI_C57BL6J_3779914 | Gm9504 | NCBI_Gene:102638412,ENSEMBL:ENSMUSG00000105277 | MGI:3779914 | pseudogene | predicted gene 9504 |
| 3 | pseudogene | 106.0909 | 106.0915 | positive | MGI_C57BL6J_3782425 | Gm4248 | NCBI_Gene:100043129,ENSEMBL:ENSMUSG00000105051 | MGI:3782425 | pseudogene | predicted pseudogene 4248 |
| 3 | gene | 106.1132 | 106.1321 | positive | MGI_C57BL6J_1932052 | Chia1 | NCBI_Gene:81600,ENSEMBL:ENSMUSG00000062778 | MGI:1932052 | protein coding gene | chitinase, acidic 1 |
| 3 | gene | 106.1476 | 106.1676 | negative | MGI_C57BL6J_1330860 | Chil3 | NCBI_Gene:12655,ENSEMBL:ENSMUSG00000040809 | MGI:1330860 | protein coding gene | chitinase-like 3 |
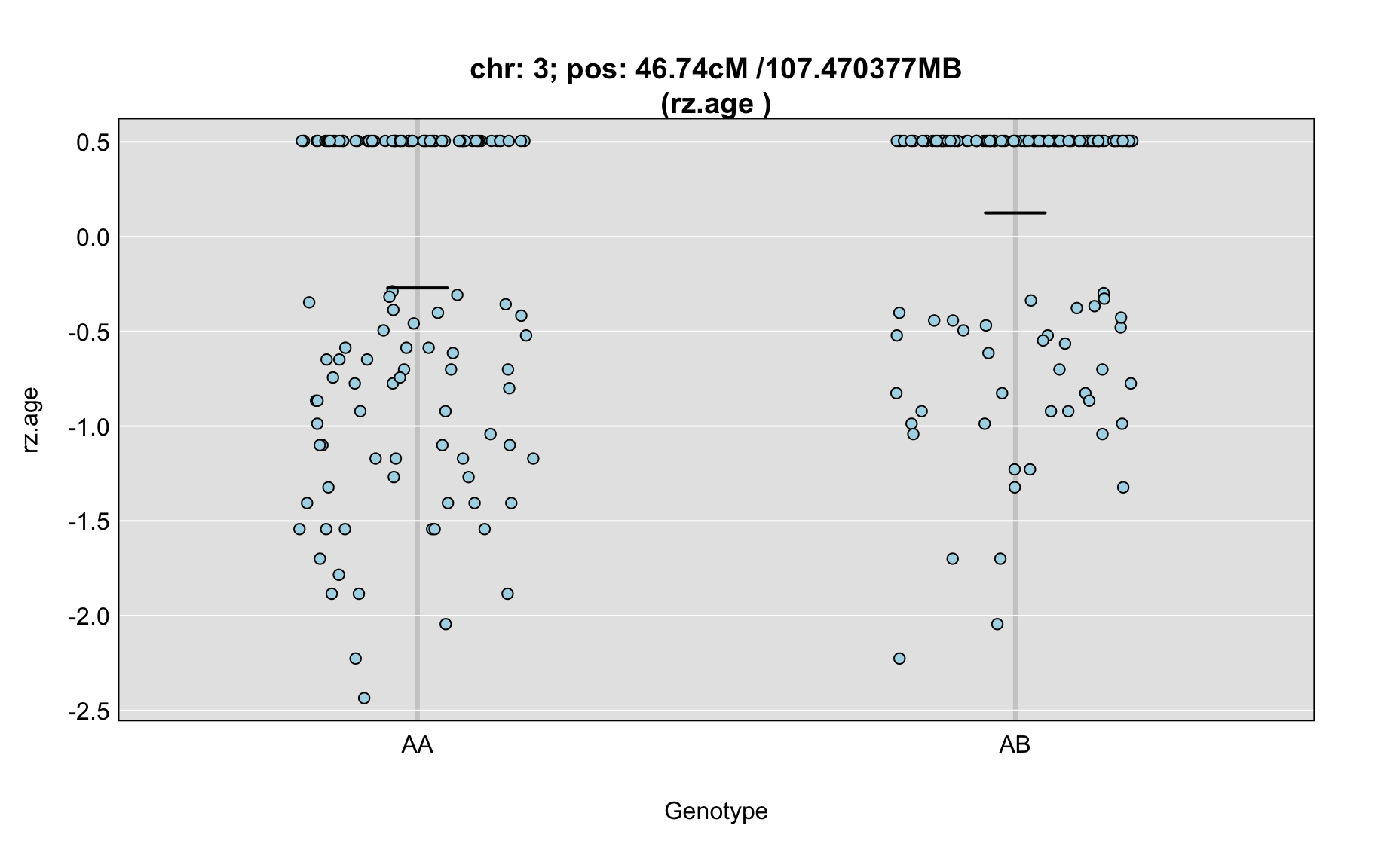
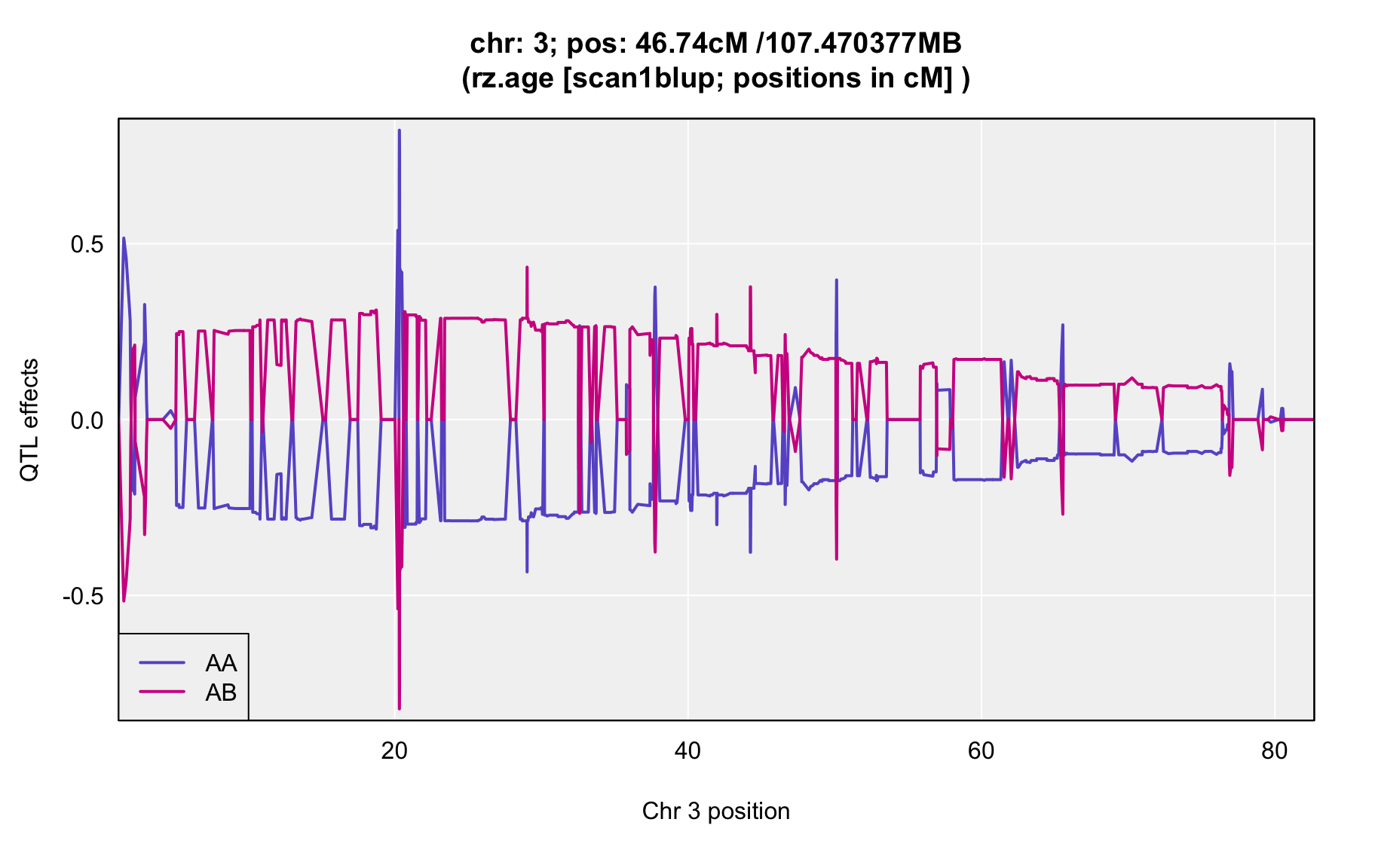
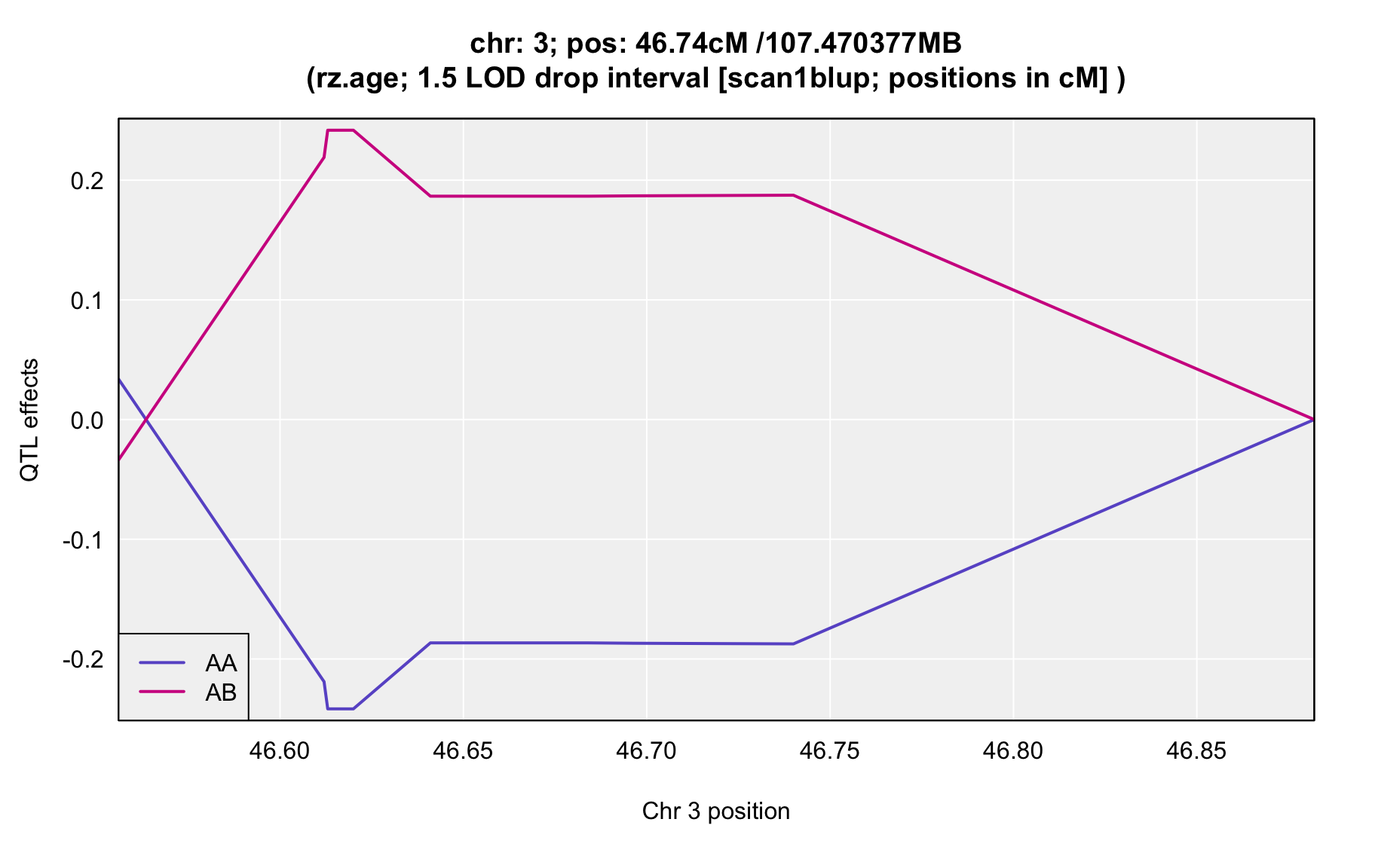
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 106.9032 | 107.0939 | negative | MGI_C57BL6J_5504123 | Gm27008 | NCBI_Gene:102639188,ENSEMBL:ENSMUSG00000098108 | MGI:5504123 | lncRNA gene | predicted gene, 27008 |
| 3 | gene | 107.0362 | 107.0381 | positive | MGI_C57BL6J_96660 | Kcna3 | NCBI_Gene:16491,ENSEMBL:ENSMUSG00000047959 | MGI:96660 | protein coding gene | potassium voltage-gated channel, shaker-related subfamily, member 3 |
| 3 | gene | 107.0395 | 107.0543 | positive | MGI_C57BL6J_2139742 | AI504432 | NCBI_Gene:229694,ENSEMBL:ENSMUSG00000056145 | MGI:2139742 | lncRNA gene | expressed sequence AI504432 |
| 3 | gene | 107.0549 | 107.0667 | negative | MGI_C57BL6J_5621597 | Gm38712 | NCBI_Gene:105242447 | MGI:5621597 | lncRNA gene | predicted gene, 38712 |
| 3 | gene | 107.0909 | 107.1150 | positive | MGI_C57BL6J_96659 | Kcna2 | NCBI_Gene:16490,ENSEMBL:ENSMUSG00000040724 | MGI:96659 | protein coding gene | potassium voltage-gated channel, shaker-related subfamily, member 2 |
| 3 | gene | 107.0909 | 107.0984 | positive | MGI_C57BL6J_1924480 | A930002I21Rik | ENSEMBL:ENSMUSG00000050179 | MGI:1924480 | lncRNA gene | RIKEN cDNA A930002I21 gene |
| 3 | gene | 107.1831 | 107.1957 | positive | MGI_C57BL6J_3037820 | Kcna10 | NCBI_Gene:242151,ENSEMBL:ENSMUSG00000042861 | MGI:3037820 | protein coding gene | potassium voltage-gated channel, shaker-related subfamily, member 10 |
| 3 | gene | 107.2113 | 107.2217 | negative | MGI_C57BL6J_2684977 | Cym | NCBI_Gene:229697,ENSEMBL:ENSMUSG00000046213 | MGI:2684977 | protein coding gene | chymosin |
| 3 | gene | 107.2306 | 107.2347 | positive | MGI_C57BL6J_2442999 | A630076J17Rik | NCBI_Gene:319929,ENSEMBL:ENSMUSG00000094613 | MGI:2442999 | protein coding gene | RIKEN cDNA A630076J17 gene |
| 3 | gene | 107.2319 | 107.2397 | negative | MGI_C57BL6J_2180370 | Prok1 | NCBI_Gene:246691,ENSEMBL:ENSMUSG00000070368 | MGI:2180370 | protein coding gene | prokineticin 1 |
| 3 | gene | 107.2789 | 107.2841 | positive | MGI_C57BL6J_1915826 | Lamtor5 | NCBI_Gene:68576,ENSEMBL:ENSMUSG00000087260 | MGI:1915826 | protein coding gene | late endosomal/lysosomal adaptor, MAPK and MTOR activator 5 |
| 3 | gene | 107.2912 | 107.3121 | positive | MGI_C57BL6J_2385183 | Slc16a4 | NCBI_Gene:229699,ENSEMBL:ENSMUSG00000027896 | MGI:2385183 | protein coding gene | solute carrier family 16 (monocarboxylic acid transporters), member 4 |
| 3 | gene | 107.3055 | 107.3350 | negative | MGI_C57BL6J_2443205 | Rbm15 | NCBI_Gene:229700,ENSEMBL:ENSMUSG00000048109 | MGI:2443205 | protein coding gene | RNA binding motif protein 15 |
| 3 | pseudogene | 107.3430 | 107.3437 | positive | MGI_C57BL6J_3647225 | Gm5279 | NCBI_Gene:383901,ENSEMBL:ENSMUSG00000105532 | MGI:3647225 | pseudogene | predicted gene 5279 |
| 3 | gene | 107.3648 | 107.3731 | positive | MGI_C57BL6J_5594859 | Gm35700 | NCBI_Gene:102639370 | MGI:5594859 | lncRNA gene | predicted gene, 35700 |
| 3 | gene | 107.3808 | 107.3826 | positive | MGI_C57BL6J_5623005 | Gm40120 | NCBI_Gene:105244526 | MGI:5623005 | lncRNA gene | predicted gene, 40120 |
| 3 | gene | 107.4159 | 107.4162 | negative | MGI_C57BL6J_5454160 | Gm24383 | ENSEMBL:ENSMUSG00000084640 | MGI:5454160 | unclassified non-coding RNA gene | predicted gene, 24383 |
| 3 | gene | 107.4383 | 107.4596 | negative | MGI_C57BL6J_96670 | Kcnc4 | NCBI_Gene:99738,ENSEMBL:ENSMUSG00000027895 | MGI:96670 | protein coding gene | potassium voltage gated channel, Shaw-related subfamily, member 4 |
| 3 | gene | 107.4403 | 107.4430 | negative | MGI_C57BL6J_2443689 | A330043J11Rik | NA | NA | unclassified gene | RIKEN cDNA A330043J11 gene |
| 3 | gene | 107.4675 | 107.5180 | negative | MGI_C57BL6J_2442535 | Slc6a17 | NCBI_Gene:229706,ENSEMBL:ENSMUSG00000027894 | MGI:2442535 | protein coding gene | solute carrier family 6 (neurotransmitter transporter), member 17 |
| 3 | gene | 107.5479 | 107.5542 | positive | MGI_C57BL6J_5595256 | Gm36097 | NCBI_Gene:102639893 | MGI:5595256 | lncRNA gene | predicted gene, 36097 |
| 3 | gene | 107.5537 | 107.5551 | negative | MGI_C57BL6J_1914841 | Ubl4b | NCBI_Gene:67591,ENSEMBL:ENSMUSG00000055891 | MGI:1914841 | protein coding gene | ubiquitin-like 4B |
| 3 | gene | 107.5950 | 107.6059 | positive | MGI_C57BL6J_1277097 | Alx3 | NCBI_Gene:11694,ENSEMBL:ENSMUSG00000014603 | MGI:1277097 | protein coding gene | aristaless-like homeobox 3 |
| 3 | gene | 107.6125 | 107.6317 | negative | MGI_C57BL6J_2443884 | Strip1 | NCBI_Gene:229707,ENSEMBL:ENSMUSG00000014601 | MGI:2443884 | protein coding gene | striatin interacting protein 1 |
| 3 | gene | 107.6313 | 107.6348 | positive | MGI_C57BL6J_3779172 | Gm10961 | NCBI_Gene:102639972,ENSEMBL:ENSMUSG00000078604 | MGI:3779172 | lncRNA gene | predicted gene 10961 |
| 3 | gene | 107.6631 | 107.6966 | negative | MGI_C57BL6J_2385184 | Ahcyl1 | NCBI_Gene:229709,ENSEMBL:ENSMUSG00000027893 | MGI:2385184 | protein coding gene | S-adenosylhomocysteine hydrolase-like 1 |
| 3 | gene | 107.6967 | 107.8130 | positive | MGI_C57BL6J_5623006 | Gm40121 | NCBI_Gene:105244527 | MGI:5623006 | lncRNA gene | predicted gene, 40121 |
| 3 | gene | 107.7410 | 107.7605 | negative | MGI_C57BL6J_1339753 | Csf1 | NCBI_Gene:12977,ENSEMBL:ENSMUSG00000014599 | MGI:1339753 | protein coding gene | colony stimulating factor 1 (macrophage) |
| 3 | pseudogene | 107.7819 | 107.7825 | positive | MGI_C57BL6J_5663370 | Gm43233 | ENSEMBL:ENSMUSG00000106531 | MGI:5663370 | pseudogene | predicted gene 43233 |
| 3 | gene | 107.8202 | 107.8335 | negative | MGI_C57BL6J_5595370 | Gm36211 | ENSEMBL:ENSMUSG00000106426 | MGI:5595370 | lncRNA gene | predicted gene, 36211 |
| 3 | pseudogene | 107.8680 | 107.8687 | negative | MGI_C57BL6J_3645483 | Gm5075 | NCBI_Gene:280047,ENSEMBL:ENSMUSG00000105944 | MGI:3645483 | pseudogene | predicted gene 5075 |
| 3 | gene | 107.8739 | 107.8929 | positive | MGI_C57BL6J_2139743 | Eps8l3 | NCBI_Gene:99662,ENSEMBL:ENSMUSG00000040600 | MGI:2139743 | protein coding gene | EPS8-like 3 |
| 3 | gene | 107.8748 | 107.8755 | positive | MGI_C57BL6J_1922590 | 4930554G22Rik | ENSEMBL:ENSMUSG00000106359 | MGI:1922590 | unclassified gene | RIKEN cDNA 4930554G22 gene |
| 3 | gene | 107.8860 | 107.8878 | positive | MGI_C57BL6J_5663882 | Gm43745 | ENSEMBL:ENSMUSG00000105456 | MGI:5663882 | unclassified gene | predicted gene 43745 |
| 3 | gene | 107.8860 | 107.8878 | negative | MGI_C57BL6J_3642497 | Gm10667 | NA | NA | unclassified gene | predicted gene 10667 |
| 3 | gene | 107.8889 | 107.8962 | negative | MGI_C57BL6J_3584041 | 4933431E20Rik | NCBI_Gene:329735,ENSEMBL:ENSMUSG00000086968 | MGI:3584041 | lncRNA gene | RIKEN cDNA 4933431E20 gene |
| 3 | gene | 107.8958 | 107.8987 | positive | MGI_C57BL6J_1309466 | Gstm5 | NCBI_Gene:14866,ENSEMBL:ENSMUSG00000004032 | MGI:1309466 | protein coding gene | glutathione S-transferase, mu 5 |
| 3 | pseudogene | 107.9068 | 107.9113 | negative | MGI_C57BL6J_3779925 | Gm9515 | NCBI_Gene:671031,ENSEMBL:ENSMUSG00000105872 | MGI:3779925 | pseudogene | predicted gene 9515 |
| 3 | pseudogene | 107.9197 | 107.9198 | positive | MGI_C57BL6J_3650952 | Gm12489 | ENSEMBL:ENSMUSG00000082720 | MGI:3650952 | pseudogene | predicted gene 12489 |
| 3 | gene | 107.9263 | 107.9319 | negative | MGI_C57BL6J_1915562 | Gstm7 | NCBI_Gene:68312,ENSEMBL:ENSMUSG00000004035 | MGI:1915562 | protein coding gene | glutathione S-transferase, mu 7 |
| 3 | gene | 107.9388 | 107.9535 | negative | MGI_C57BL6J_1309467 | Gstm6 | NCBI_Gene:14867,ENSEMBL:ENSMUSG00000068762 | MGI:1309467 | protein coding gene | glutathione S-transferase, mu 6 |
| 3 | pseudogene | 107.9508 | 107.9531 | negative | MGI_C57BL6J_3702525 | Gm12494 | ENSEMBL:ENSMUSG00000089664 | MGI:3702525 | pseudogene | predicted gene 12494 |
| 3 | gene | 107.9637 | 107.9693 | negative | MGI_C57BL6J_106026 | Gstm3 | NCBI_Gene:14864,ENSEMBL:ENSMUSG00000004038 | MGI:106026 | protein coding gene | glutathione S-transferase, mu 3 |
| 3 | pseudogene | 107.9796 | 107.9800 | positive | MGI_C57BL6J_3650111 | Gm12497 | NCBI_Gene:624701,ENSEMBL:ENSMUSG00000049231 | MGI:3650111 | pseudogene | predicted pseudogene 12497 |
| 3 | gene | 107.9817 | 107.9865 | negative | MGI_C57BL6J_95861 | Gstm2 | NCBI_Gene:14863,ENSEMBL:ENSMUSG00000040562 | MGI:95861 | protein coding gene | glutathione S-transferase, mu 2 |
| 3 | pseudogene | 107.9994 | 107.9998 | negative | MGI_C57BL6J_5663883 | Gm43746 | ENSEMBL:ENSMUSG00000106614 | MGI:5663883 | pseudogene | predicted gene 43746 |
| 3 | gene | 108.0122 | 108.0180 | negative | MGI_C57BL6J_95860 | Gstm1 | NCBI_Gene:14862,ENSEMBL:ENSMUSG00000058135 | MGI:95860 | protein coding gene | glutathione S-transferase, mu 1 |
| 3 | gene | 108.0183 | 108.0259 | negative | MGI_C57BL6J_3650533 | Gm12498 | NCBI_Gene:108168891,ENSEMBL:ENSMUSG00000087357 | MGI:3650533 | lncRNA gene | predicted gene 12498 |
| 3 | pseudogene | 108.0197 | 108.0204 | positive | MGI_C57BL6J_3650307 | Gm12502 | NCBI_Gene:102640327,ENSEMBL:ENSMUSG00000083627 | MGI:3650307 | pseudogene | predicted gene 12502 |
| 3 | pseudogene | 108.0296 | 108.0314 | negative | MGI_C57BL6J_3650532 | Gm12499 | ENSEMBL:ENSMUSG00000081688 | MGI:3650532 | pseudogene | predicted gene 12499 |
| 3 | gene | 108.0306 | 108.0451 | negative | MGI_C57BL6J_95862 | Gstm4 | NCBI_Gene:14865,ENSEMBL:ENSMUSG00000027890 | MGI:95862 | protein coding gene | glutathione S-transferase, mu 4 |
| 3 | gene | 108.0308 | 108.0370 | negative | MGI_C57BL6J_5595657 | Gm36498 | NCBI_Gene:102640435 | MGI:5595657 | lncRNA gene | predicted gene, 36498 |
| 3 | gene | 108.0340 | 108.0342 | positive | MGI_C57BL6J_5455369 | Gm25592 | ENSEMBL:ENSMUSG00000064569 | MGI:5455369 | snoRNA gene | predicted gene, 25592 |
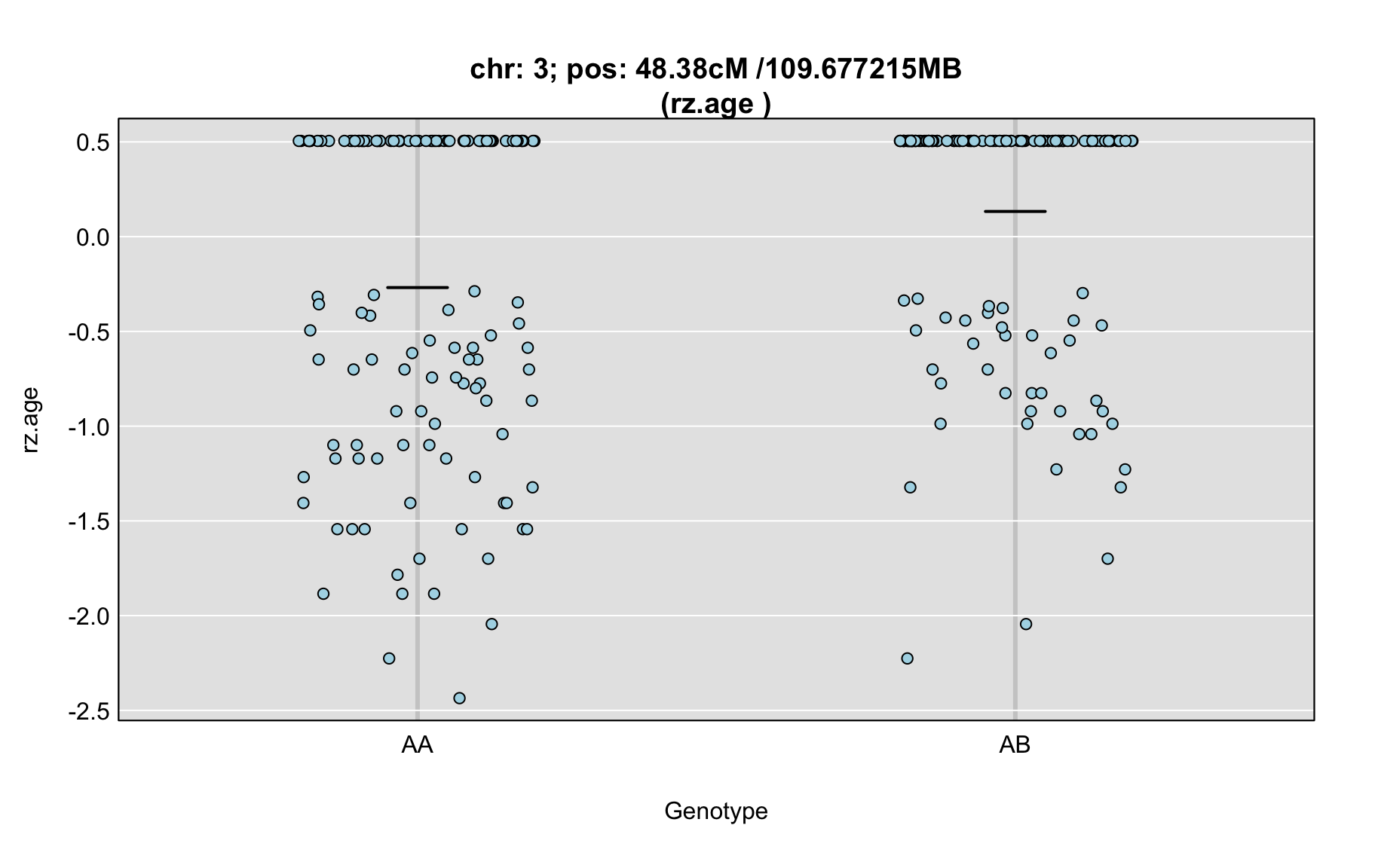
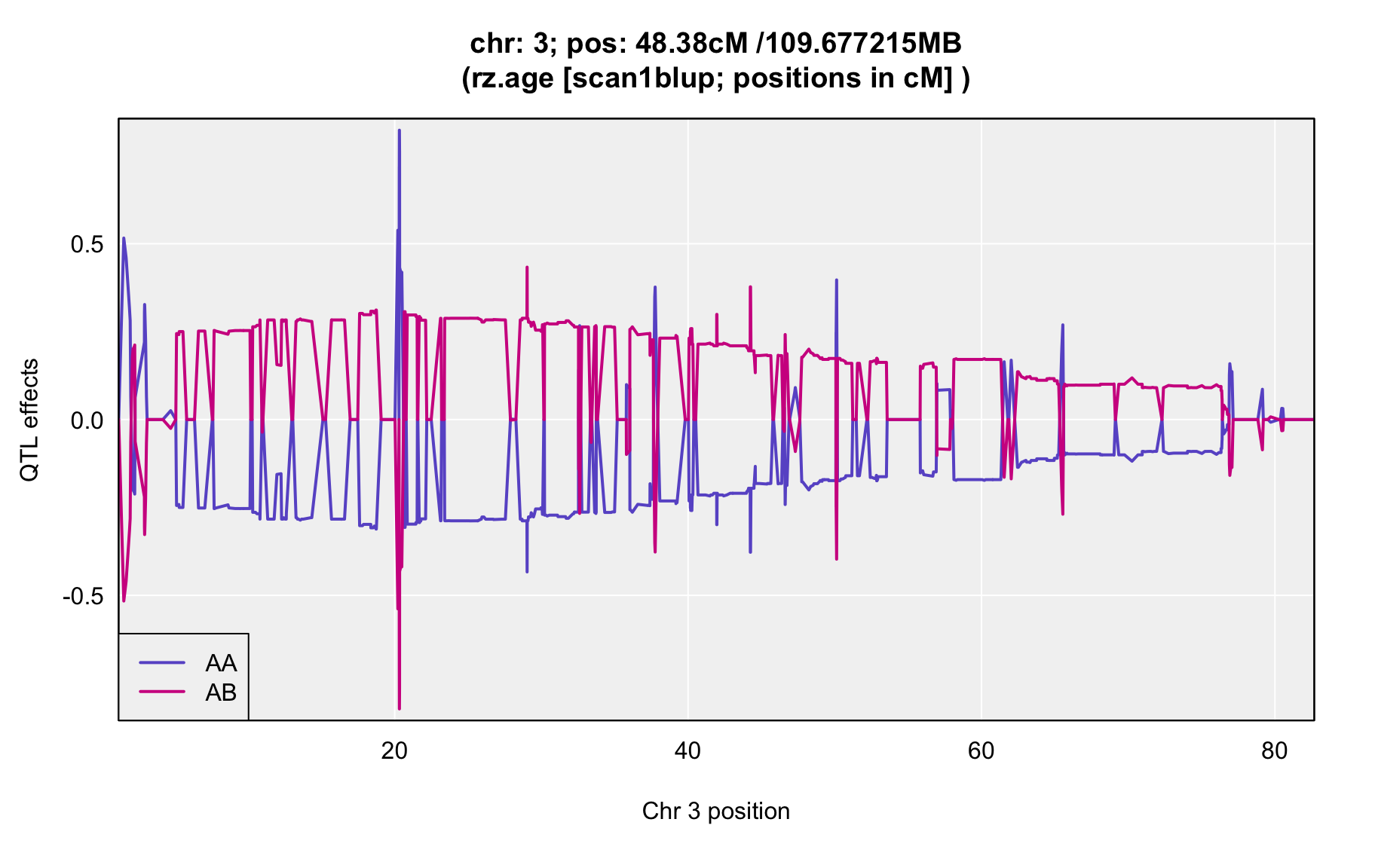
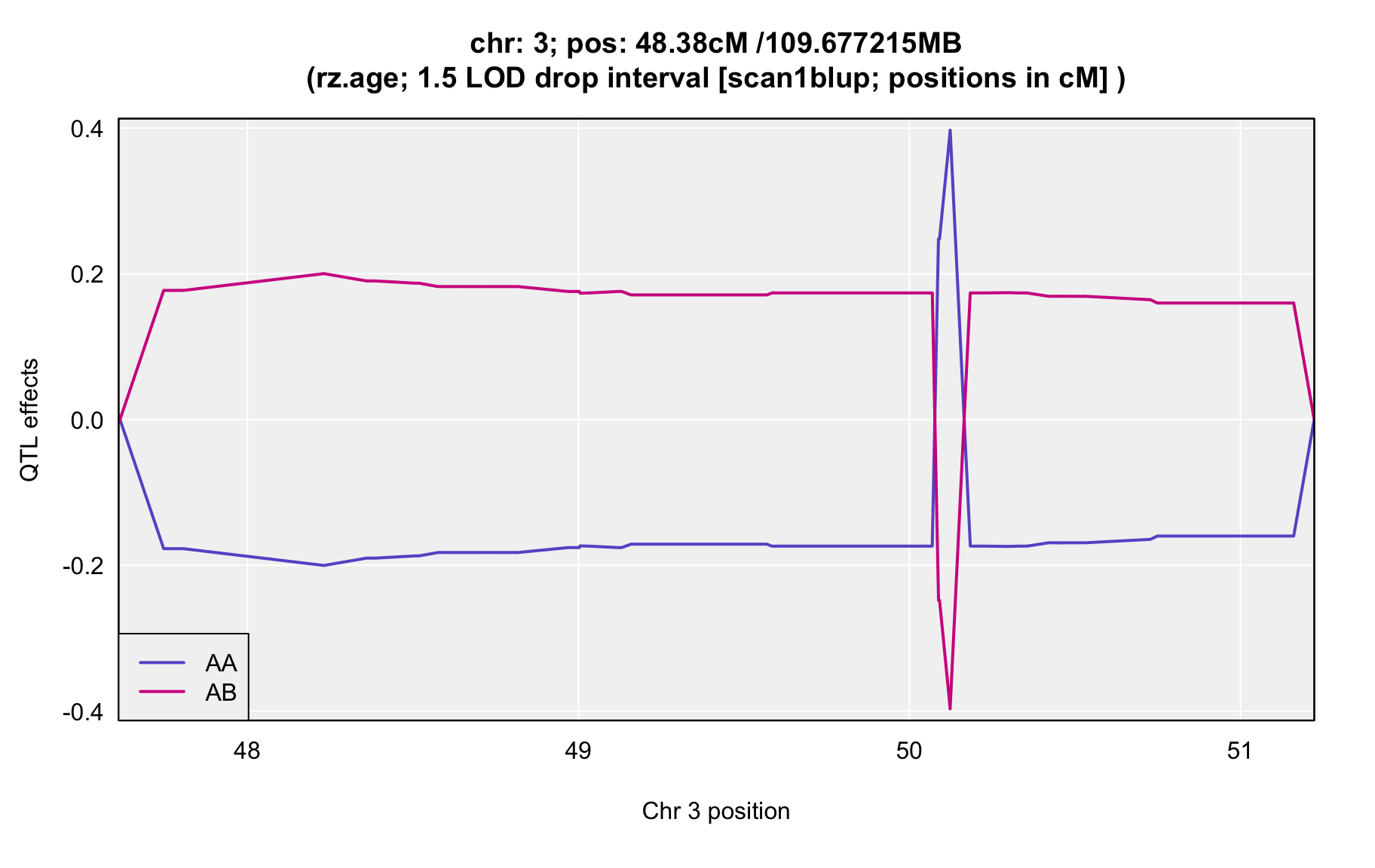
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 108.8537 | 108.8900 | negative | MGI_C57BL6J_2443535 | Fndc7 | NCBI_Gene:320181,ENSEMBL:ENSMUSG00000045326 | MGI:2443535 | protein coding gene | fibronectin type III domain containing 7 |
| 3 | gene | 108.9028 | 108.9118 | negative | MGI_C57BL6J_1914171 | Prpf38b | NCBI_Gene:66921,ENSEMBL:ENSMUSG00000027881 | MGI:1914171 | protein coding gene | PRP38 pre-mRNA processing factor 38 (yeast) domain containing B |
| 3 | gene | 108.9385 | 108.9608 | positive | MGI_C57BL6J_1913965 | Henmt1 | NCBI_Gene:66715,ENSEMBL:ENSMUSG00000045662 | MGI:1913965 | protein coding gene | HEN1 methyltransferase homolog 1 (Arabidopsis) |
| 3 | gene | 108.9399 | 108.9404 | negative | MGI_C57BL6J_3704224 | Gm9857 | ENSEMBL:ENSMUSG00000051638 | MGI:3704224 | protein coding gene | predicted gene 9857 |
| 3 | gene | 108.9710 | 109.0276 | negative | MGI_C57BL6J_3036259 | Fam102b | NCBI_Gene:329739,ENSEMBL:ENSMUSG00000040339 | MGI:3036259 | protein coding gene | family with sequence similarity 102, member B |
| 3 | gene | 109.0000 | 109.0021 | positive | MGI_C57BL6J_5663358 | Gm43221 | ENSEMBL:ENSMUSG00000106108 | MGI:5663358 | lncRNA gene | predicted gene 43221 |
| 3 | gene | 109.0556 | 109.0557 | positive | MGI_C57BL6J_5690776 | Gm44384 | ENSEMBL:ENSMUSG00000105371 | MGI:5690776 | miRNA gene | predicted gene, 44384 |
| 3 | gene | 109.0557 | 109.0654 | positive | MGI_C57BL6J_5595937 | Gm36778 | NCBI_Gene:102640795 | MGI:5595937 | lncRNA gene | predicted gene, 36778 |
| 3 | gene | 109.0786 | 109.0804 | negative | MGI_C57BL6J_1924115 | 4930408K08Rik | ENSEMBL:ENSMUSG00000105638 | MGI:1924115 | lncRNA gene | RIKEN cDNA 4930408K08 gene |
| 3 | gene | 109.0805 | 109.1167 | positive | MGI_C57BL6J_1921936 | Slc25a54 | NCBI_Gene:74686,ENSEMBL:ENSMUSG00000027880 | MGI:1921936 | protein coding gene | solute carrier family 25, member 54 |
| 3 | gene | 109.1205 | 109.1231 | negative | MGI_C57BL6J_5011576 | Gm19391 | NCBI_Gene:105244530,ENSEMBL:ENSMUSG00000105139 | MGI:5011576 | lncRNA gene | predicted gene, 19391 |
| 3 | gene | 109.1231 | 109.1685 | positive | MGI_C57BL6J_1917160 | Slc25a24 | NCBI_Gene:229731,ENSEMBL:ENSMUSG00000040322 | MGI:1917160 | protein coding gene | solute carrier family 25 (mitochondrial carrier, phosphate carrier), member 24 |
| 3 | pseudogene | 109.1920 | 109.1929 | negative | MGI_C57BL6J_3651348 | Gm13865 | NCBI_Gene:100417395,ENSEMBL:ENSMUSG00000083717 | MGI:3651348 | pseudogene | predicted gene 13865 |
| 3 | gene | 109.3403 | 109.6857 | positive | MGI_C57BL6J_1888518 | Vav3 | NCBI_Gene:57257,ENSEMBL:ENSMUSG00000033721 | MGI:1888518 | protein coding gene | vav 3 oncogene |
| 3 | gene | 109.4509 | 109.4547 | positive | MGI_C57BL6J_2444726 | C130083B21Rik | NA | NA | unclassified gene | RIKEN cDNA C130083B21 gene |
| 3 | gene | 109.7800 | 110.1440 | negative | MGI_C57BL6J_1934028 | Ntng1 | NCBI_Gene:80883,ENSEMBL:ENSMUSG00000059857 | MGI:1934028 | protein coding gene | netrin G1 |
| 3 | gene | 109.8638 | 109.8639 | positive | MGI_C57BL6J_5455458 | Gm25681 | ENSEMBL:ENSMUSG00000065872 | MGI:5455458 | snRNA gene | predicted gene, 25681 |
| 3 | pseudogene | 109.9074 | 109.9079 | positive | MGI_C57BL6J_3650178 | Gm12535 | ENSEMBL:ENSMUSG00000082624 | MGI:3650178 | pseudogene | predicted gene 12535 |
| 3 | gene | 110.2439 | 110.2446 | negative | MGI_C57BL6J_5662653 | Gm42516 | ENSEMBL:ENSMUSG00000105639 | MGI:5662653 | unclassified gene | predicted gene 42516 |
| 3 | gene | 110.2461 | 110.2510 | negative | MGI_C57BL6J_2139971 | Prmt6 | NCBI_Gene:99890,ENSEMBL:ENSMUSG00000049300 | MGI:2139971 | protein coding gene | protein arginine N-methyltransferase 6 |
| 3 | gene | 110.2500 | 110.2537 | positive | MGI_C57BL6J_3697343 | C130013H08Rik | ENSEMBL:ENSMUSG00000105509 | MGI:3697343 | lncRNA gene | RIKEN cDNA C130013H08 gene |
| 3 | pseudogene | 110.2771 | 110.2775 | negative | MGI_C57BL6J_5663542 | Gm43405 | ENSEMBL:ENSMUSG00000104944 | MGI:5663542 | pseudogene | predicted gene 43405 |
| 3 | pseudogene | 110.3991 | 110.3998 | positive | MGI_C57BL6J_5011370 | Gm19185 | NCBI_Gene:100418401,ENSEMBL:ENSMUSG00000105859 | MGI:5011370 | pseudogene | predicted gene, 19185 |
| 3 | pseudogene | 110.5641 | 110.5644 | negative | MGI_C57BL6J_5663543 | Gm43406 | ENSEMBL:ENSMUSG00000105121 | MGI:5663543 | pseudogene | predicted gene 43406 |
| 3 | gene | 110.5644 | 110.5645 | negative | MGI_C57BL6J_5455853 | Gm26076 | ENSEMBL:ENSMUSG00000077253 | MGI:5455853 | snRNA gene | predicted gene, 26076 |
| 3 | gene | 110.6343 | 110.6369 | positive | MGI_C57BL6J_5663541 | Gm43404 | ENSEMBL:ENSMUSG00000106650 | MGI:5663541 | unclassified gene | predicted gene 43404 |
| 3 | pseudogene | 111.1085 | 111.1088 | positive | MGI_C57BL6J_5663544 | Gm43407 | ENSEMBL:ENSMUSG00000106487 | MGI:5663544 | pseudogene | predicted gene 43407 |
| 3 | pseudogene | 111.3211 | 111.3230 | negative | MGI_C57BL6J_3647777 | Gm9314 | NCBI_Gene:668712,ENSEMBL:ENSMUSG00000106100 | MGI:3647777 | pseudogene | predicted gene 9314 |
| 3 | gene | 111.7895 | 111.7911 | positive | MGI_C57BL6J_5663042 | Gm42905 | ENSEMBL:ENSMUSG00000105882 | MGI:5663042 | unclassified gene | predicted gene 42905 |
| 3 | pseudogene | 111.8094 | 111.8098 | negative | MGI_C57BL6J_5663991 | Gm43854 | ENSEMBL:ENSMUSG00000104711 | MGI:5663991 | pseudogene | predicted gene 43854 |
| 3 | gene | 111.8819 | 112.0820 | positive | MGI_C57BL6J_3779614 | Gm6602 | NCBI_Gene:625585,ENSEMBL:ENSMUSG00000104543 | MGI:3779614 | lncRNA gene | predicted gene 6602 |
| 3 | gene | 112.1989 | 112.1990 | negative | MGI_C57BL6J_5455296 | Gm25519 | ENSEMBL:ENSMUSG00000089017 | MGI:5455296 | snoRNA gene | predicted gene, 25519 |
| 3 | pseudogene | 112.2027 | 112.2030 | negative | MGI_C57BL6J_5663990 | Gm43853 | ENSEMBL:ENSMUSG00000105507 | MGI:5663990 | pseudogene | predicted gene 43853 |
| 3 | pseudogene | 112.3023 | 112.3033 | positive | MGI_C57BL6J_3646906 | Gm4859 | NCBI_Gene:229746,ENSEMBL:ENSMUSG00000105077 | MGI:3646906 | pseudogene | predicted gene 4859 |
| 3 | pseudogene | 112.3309 | 112.3318 | negative | MGI_C57BL6J_3647781 | Gm9317 | NCBI_Gene:668717,ENSEMBL:ENSMUSG00000105073 | MGI:3647781 | pseudogene | predicted gene 9317 |
| 3 | pseudogene | 112.5916 | 112.5920 | positive | MGI_C57BL6J_5662663 | Gm42526 | ENSEMBL:ENSMUSG00000105785 | MGI:5662663 | pseudogene | predicted gene 42526 |
| 3 | pseudogene | 112.7668 | 112.7670 | negative | MGI_C57BL6J_5663724 | Gm43587 | ENSEMBL:ENSMUSG00000106666 | MGI:5663724 | pseudogene | predicted gene 43587 |
| 3 | pseudogene | 113.0195 | 113.0230 | positive | MGI_C57BL6J_3051682 | Utp14b-ps1 | NCBI_Gene:383905,ENSEMBL:ENSMUSG00000105588 | MGI:3051682 | pseudogene | UTP14B small subunit processome component, pseudogene 1 |
| 3 | pseudogene | 113.0525 | 113.0543 | negative | MGI_C57BL6J_3644344 | Gm5548 | NCBI_Gene:102632213,ENSEMBL:ENSMUSG00000045952 | MGI:3644344 | pseudogene | predicted pseudogene 5548 |
| 3 | pseudogene | 113.0558 | 113.0566 | negative | MGI_C57BL6J_5618687 | Gm38395 | NCBI_Gene:100416050,ENSEMBL:ENSMUSG00000105665 | MGI:5618687 | pseudogene | predicted gene, 38395 |
| 3 | pseudogene | 113.1477 | 113.1567 | negative | MGI_C57BL6J_104547 | Amy2b | NCBI_Gene:545562,ENSEMBL:ENSMUSG00000083079 | MGI:104547 | pseudogene | amylase 2b |
| 3 | pseudogene | 113.1822 | 113.2067 | negative | MGI_C57BL6J_104546 | Amy2-ps1 | NCBI_Gene:668723,ENSEMBL:ENSMUSG00000081506 | MGI:104546 | pseudogene | amylase 2, pseudogene 1 |
| 3 | gene | 113.2472 | 113.2477 | positive | MGI_C57BL6J_1916346 | 1810008N23Rik | NA | NA | unclassified non-coding RNA gene | RIKEN cDNA 1810008N23 gene |
| 3 | gene | 113.2472 | 113.2588 | negative | MGI_C57BL6J_88020 | Amy2a5 | NCBI_Gene:109959,ENSEMBL:ENSMUSG00000074268 | MGI:88020 | protein coding gene | amylase 2a5 |
| 3 | gene | 113.2798 | 113.2914 | negative | MGI_C57BL6J_3711258 | Amy2a4 | NCBI_Gene:100043684,ENSEMBL:ENSMUSG00000096770 | MGI:3711258 | protein coding gene | amylase 2a4 |
| 3 | gene | 113.3124 | 113.3241 | negative | MGI_C57BL6J_3714985 | Amy2a3 | NCBI_Gene:100043686,ENSEMBL:ENSMUSG00000093931 | MGI:3714985 | protein coding gene | amylase 2a3 |
| 3 | gene | 113.3450 | 113.3567 | negative | MGI_C57BL6J_3711220 | Amy2a2 | NCBI_Gene:100043688,ENSEMBL:ENSMUSG00000096569 | MGI:3711220 | protein coding gene | amylase 2a2 |
| 3 | gene | 113.5281 | 113.5324 | negative | MGI_C57BL6J_104548 | Amy2a1 | NCBI_Gene:100043207,ENSEMBL:ENSMUSG00000070360 | MGI:104548 | protein coding gene | amylase 2a1 |
| 3 | gene | 113.5434 | 113.5458 | negative | MGI_C57BL6J_5826457 | Gm46820 | NCBI_Gene:108168892 | MGI:5826457 | lncRNA gene | predicted gene, 46820 |
| 3 | gene | 113.5557 | 113.6067 | negative | MGI_C57BL6J_88019 | Amy1 | NCBI_Gene:11722,ENSEMBL:ENSMUSG00000074264 | MGI:88019 | protein coding gene | amylase 1, salivary |
| 3 | gene | 113.6051 | 113.6301 | negative | MGI_C57BL6J_1914475 | Rnpc3 | NCBI_Gene:67225,ENSEMBL:ENSMUSG00000027981 | MGI:1914475 | protein coding gene | RNA-binding region (RNP1, RRM) containing 3 |
| 3 | gene | 113.6299 | 113.6313 | positive | MGI_C57BL6J_3698878 | C530020B09Rik | NA | NA | unclassified non-coding RNA gene | RIKEN cDNA C530020B09 gene |
| 3 | pseudogene | 113.6316 | 113.6322 | negative | MGI_C57BL6J_5010139 | Gm17954 | NCBI_Gene:100416168,ENSEMBL:ENSMUSG00000105836 | MGI:5010139 | pseudogene | predicted gene, 17954 |
| 3 | pseudogene | 113.7286 | 113.7295 | positive | MGI_C57BL6J_5009939 | Gm17775 | NCBI_Gene:100046591,ENSEMBL:ENSMUSG00000106454 | MGI:5009939 | pseudogene | predicted gene, 17775 |
| 3 | pseudogene | 113.8547 | 113.8564 | positive | MGI_C57BL6J_5010348 | Gm18163 | NCBI_Gene:100416594,ENSEMBL:ENSMUSG00000106084 | MGI:5010348 | pseudogene | predicted gene, 18163 |
| 3 | gene | 114.0305 | 114.2208 | positive | MGI_C57BL6J_88446 | Col11a1 | NCBI_Gene:12814,ENSEMBL:ENSMUSG00000027966 | MGI:88446 | protein coding gene | collagen, type XI, alpha 1 |
| 3 | pseudogene | 114.5444 | 114.5448 | negative | MGI_C57BL6J_5663886 | Gm43749 | ENSEMBL:ENSMUSG00000106452 | MGI:5663886 | pseudogene | predicted gene 43749 |
| 3 | gene | 114.8242 | 114.8243 | negative | MGI_C57BL6J_5453527 | Gm23750 | ENSEMBL:ENSMUSG00000077410 | MGI:5453527 | snoRNA gene | predicted gene, 23750 |
| 3 | pseudogene | 114.8280 | 114.8287 | negative | MGI_C57BL6J_5011332 | Gm19147 | NCBI_Gene:100418337,ENSEMBL:ENSMUSG00000105225 | MGI:5011332 | pseudogene | predicted gene, 19147 |
| 3 | gene | 114.9041 | 115.1258 | positive | MGI_C57BL6J_2387329 | Olfm3 | NCBI_Gene:229759,ENSEMBL:ENSMUSG00000027965 | MGI:2387329 | protein coding gene | olfactomedin 3 |
| 3 | pseudogene | 115.1716 | 115.1721 | negative | MGI_C57BL6J_5010614 | Gm18429 | NCBI_Gene:100417157,ENSEMBL:ENSMUSG00000106510 | MGI:5010614 | pseudogene | predicted gene, 18429 |
| 3 | gene | 115.3916 | 115.3917 | positive | MGI_C57BL6J_5452114 | Gm22337 | ENSEMBL:ENSMUSG00000088346 | MGI:5452114 | snoRNA gene | predicted gene, 22337 |
| 3 | gene | 115.4840 | 115.4841 | negative | MGI_C57BL6J_5455124 | Gm25347 | ENSEMBL:ENSMUSG00000080676 | MGI:5455124 | miRNA gene | predicted gene, 25347 |
| 3 | pseudogene | 115.5186 | 115.5190 | positive | MGI_C57BL6J_5663806 | Gm43669 | ENSEMBL:ENSMUSG00000105829 | MGI:5663806 | pseudogene | predicted gene 43669 |
| 3 | gene | 115.6367 | 115.6417 | negative | MGI_C57BL6J_5623009 | Gm40124 | NCBI_Gene:105244531 | MGI:5623009 | lncRNA gene | predicted gene, 40124 |
| 3 | gene | 115.7104 | 115.7151 | negative | MGI_C57BL6J_1096355 | S1pr1 | NCBI_Gene:13609,ENSEMBL:ENSMUSG00000045092 | MGI:1096355 | protein coding gene | sphingosine-1-phosphate receptor 1 |
| 3 | gene | 115.7151 | 115.7167 | positive | MGI_C57BL6J_3642568 | Gm9889 | ENSEMBL:ENSMUSG00000052769 | MGI:3642568 | lncRNA gene | predicted gene 9889 |
| 3 | gene | 115.8816 | 115.8881 | negative | MGI_C57BL6J_1915411 | A930005H10Rik | NCBI_Gene:68161,ENSEMBL:ENSMUSG00000054426 | MGI:1915411 | lncRNA gene | RIKEN cDNA A930005H10 gene |
| 3 | gene | 115.8878 | 115.9344 | positive | MGI_C57BL6J_1916990 | Dph5 | NCBI_Gene:69740,ENSEMBL:ENSMUSG00000033554 | MGI:1916990 | protein coding gene | diphthamide biosynthesis 5 |
| 3 | gene | 115.8893 | 115.8923 | positive | MGI_C57BL6J_2442378 | A230060L02Rik | NA | NA | unclassified gene | RIKEN cDNA A230060L02 gene |
| 3 | gene | 115.9104 | 115.9105 | positive | MGI_C57BL6J_5662824 | Gm42687 | ENSEMBL:ENSMUSG00000105108 | MGI:5662824 | unclassified gene | predicted gene 42687 |
| 3 | gene | 115.9138 | 115.9141 | negative | MGI_C57BL6J_5663244 | Gm43107 | ENSEMBL:ENSMUSG00000105017 | MGI:5663244 | unclassified gene | predicted gene 43107 |
| 3 | gene | 115.9139 | 115.9142 | positive | MGI_C57BL6J_5663245 | Gm43108 | ENSEMBL:ENSMUSG00000104803 | MGI:5663245 | unclassified gene | predicted gene 43108 |
| 3 | gene | 115.9390 | 116.0074 | negative | MGI_C57BL6J_1913750 | Slc30a7 | NCBI_Gene:66500,ENSEMBL:ENSMUSG00000054414 | MGI:1913750 | protein coding gene | solute carrier family 30 (zinc transporter), member 7 |
| 3 | pseudogene | 115.9425 | 115.9431 | positive | MGI_C57BL6J_5663243 | Gm43106 | ENSEMBL:ENSMUSG00000104723 | MGI:5663243 | pseudogene | predicted gene 43106 |
| 3 | pseudogene | 115.9431 | 115.9443 | positive | MGI_C57BL6J_3779621 | Gm6649 | NCBI_Gene:626061,ENSEMBL:ENSMUSG00000105645 | MGI:3779621 | pseudogene | predicted gene 6649 |
| 3 | gene | 115.9799 | 115.9800 | positive | MGI_C57BL6J_3837039 | Mir669n | miRBase:MI0009951,NCBI_Gene:100316705,ENSEMBL:ENSMUSG00000089261 | MGI:3837039 | miRNA gene | microRNA 669n |
| 3 | gene | 116.0074 | 116.0290 | positive | MGI_C57BL6J_1889574 | Extl2 | NCBI_Gene:58193,ENSEMBL:ENSMUSG00000027963 | MGI:1889574 | protein coding gene | exostoses (multiple)-like 2 |
| 3 | gene | 116.0658 | 116.0659 | negative | MGI_C57BL6J_5456121 | Gm26344 | ENSEMBL:ENSMUSG00000077741 | MGI:5456121 | snoRNA gene | predicted gene, 26344 |
| 3 | gene | 116.0936 | 116.1024 | positive | MGI_C57BL6J_5590543 | Gm31384 | NCBI_Gene:102633598 | MGI:5590543 | lncRNA gene | predicted gene, 31384 |
| 3 | gene | 116.1099 | 116.1297 | negative | MGI_C57BL6J_98926 | Vcam1 | NCBI_Gene:22329,ENSEMBL:ENSMUSG00000027962 | MGI:98926 | protein coding gene | vascular cell adhesion molecule 1 |
| 3 | gene | 116.1285 | 116.1410 | positive | MGI_C57BL6J_5663246 | Gm43109 | ENSEMBL:ENSMUSG00000104653 | MGI:5663246 | lncRNA gene | predicted gene 43109 |
| 3 | gene | 116.1349 | 116.1491 | positive | MGI_C57BL6J_1924313 | 5330425B07Rik | NCBI_Gene:102633677,ENSEMBL:ENSMUSG00000105729 | MGI:1924313 | lncRNA gene | RIKEN cDNA 5330425B07 gene |
| 3 | gene | 116.1505 | 116.1658 | positive | MGI_C57BL6J_5477038 | Gm26544 | NCBI_Gene:102633872,ENSEMBL:ENSMUSG00000097362 | MGI:5477038 | lncRNA gene | predicted gene, 26544 |
| 3 | gene | 116.1982 | 116.2036 | positive | MGI_C57BL6J_5590810 | Gm31651 | NCBI_Gene:102633953,ENSEMBL:ENSMUSG00000104607 | MGI:5590810 | lncRNA gene | predicted gene, 31651 |
| 3 | gene | 116.2497 | 116.2535 | negative | MGI_C57BL6J_1927653 | Gpr88 | NCBI_Gene:64378,ENSEMBL:ENSMUSG00000068696 | MGI:1927653 | protein coding gene | G-protein coupled receptor 88 |
| 3 | gene | 116.2579 | 116.2700 | positive | MGI_C57BL6J_5826476 | Gm46839 | NCBI_Gene:108168922 | MGI:5826476 | lncRNA gene | predicted gene, 46839 |
| 3 | gene | 116.2726 | 116.4287 | negative | MGI_C57BL6J_2442676 | Cdc14a | NCBI_Gene:229776,ENSEMBL:ENSMUSG00000033502 | MGI:2442676 | protein coding gene | CDC14 cell division cycle 14A |
| 3 | gene | 116.3501 | 116.3529 | positive | MGI_C57BL6J_5579857 | Gm29151 | ENSEMBL:ENSMUSG00000106005 | MGI:5579857 | lncRNA gene | predicted gene 29151 |
| 3 | gene | 116.4242 | 116.4354 | positive | MGI_C57BL6J_5623010 | Gm40125 | NCBI_Gene:105244532 | MGI:5623010 | lncRNA gene | predicted gene, 40125 |
| 3 | gene | 116.4484 | 116.4486 | negative | MGI_C57BL6J_5452138 | Gm22361 | ENSEMBL:ENSMUSG00000089060 | MGI:5452138 | snRNA gene | predicted gene, 22361 |
| 3 | gene | 116.4666 | 116.4667 | positive | MGI_C57BL6J_5456135 | Gm26358 | ENSEMBL:ENSMUSG00000088840 | MGI:5456135 | snoRNA gene | predicted gene, 26358 |
| 3 | gene | 116.4890 | 116.5082 | negative | MGI_C57BL6J_1913618 | Rtca | NCBI_Gene:66368,ENSEMBL:ENSMUSG00000000339 | MGI:1913618 | protein coding gene | RNA 3’-terminal phosphate cyclase |
| 3 | gene | 116.4890 | 116.4934 | positive | MGI_C57BL6J_5477024 | Gm26530 | ENSEMBL:ENSMUSG00000097737 | MGI:5477024 | lncRNA gene | predicted gene, 26530 |
| 3 | gene | 116.5131 | 116.5500 | positive | MGI_C57BL6J_105386 | Dbt | NCBI_Gene:13171,ENSEMBL:ENSMUSG00000000340 | MGI:105386 | protein coding gene | dihydrolipoamide branched chain transacylase E2 |
| 3 | pseudogene | 116.5273 | 116.5287 | negative | MGI_C57BL6J_3782779 | Gm4596 | NCBI_Gene:100043702,ENSEMBL:ENSMUSG00000082394 | MGI:3782779 | pseudogene | predicted gene 4596 |
| 3 | gene | 116.5321 | 116.5326 | positive | MGI_C57BL6J_5663788 | Gm43651 | ENSEMBL:ENSMUSG00000105992 | MGI:5663788 | unclassified gene | predicted gene 43651 |
| 3 | pseudogene | 116.5606 | 116.5615 | negative | MGI_C57BL6J_3708640 | Gm9761 | NCBI_Gene:100043705,ENSEMBL:ENSMUSG00000034437 | MGI:3708640 | pseudogene | predicted gene 9761 |
| 3 | gene | 116.5630 | 116.5831 | positive | MGI_C57BL6J_1924557 | Lrrc39 | NCBI_Gene:109245,ENSEMBL:ENSMUSG00000027961 | MGI:1924557 | protein coding gene | leucine rich repeat containing 39 |
| 3 | gene | 116.5721 | 116.5744 | negative | MGI_C57BL6J_5663077 | Gm42940 | ENSEMBL:ENSMUSG00000106025 | MGI:5663077 | unclassified gene | predicted gene 42940 |
| 3 | gene | 116.5766 | 116.6146 | negative | MGI_C57BL6J_1925219 | Trmt13 | NCBI_Gene:229780,ENSEMBL:ENSMUSG00000033439 | MGI:1925219 | protein coding gene | tRNA methyltransferase 13 |
| 3 | gene | 116.5950 | 116.6310 | positive | MGI_C57BL6J_1920026 | Sass6 | NCBI_Gene:72776,ENSEMBL:ENSMUSG00000027959 | MGI:1920026 | protein coding gene | SAS-6 centriolar assembly protein |
| 3 | gene | 116.6292 | 116.6310 | negative | MGI_C57BL6J_5663030 | Gm42893 | ENSEMBL:ENSMUSG00000106415 | MGI:5663030 | lncRNA gene | predicted gene 42893 |
| 3 | gene | 116.6312 | 116.6627 | negative | MGI_C57BL6J_1201609 | Mfsd14a | NCBI_Gene:15247,ENSEMBL:ENSMUSG00000089911 | MGI:1201609 | protein coding gene | major facilitator superfamily domain containing 14A |
| 3 | gene | 116.6317 | 116.6322 | positive | MGI_C57BL6J_5662594 | Gm42457 | ENSEMBL:ENSMUSG00000105339 | MGI:5662594 | lncRNA gene | predicted gene 42457 |
| 3 | gene | 116.6419 | 116.6813 | negative | MGI_C57BL6J_5663328 | Gm43191 | ENSEMBL:ENSMUSG00000105103 | MGI:5663328 | protein coding gene | predicted gene 43191 |
| 3 | gene | 116.6637 | 116.6642 | positive | MGI_C57BL6J_1925434 | 4930512P04Rik | ENSEMBL:ENSMUSG00000106125 | MGI:1925434 | lncRNA gene | RIKEN cDNA 4930512P04 gene |
| 3 | gene | 116.6695 | 116.7137 | negative | MGI_C57BL6J_1917648 | Slc35a3 | NCBI_Gene:229782,ENSEMBL:ENSMUSG00000027957 | MGI:1917648 | protein coding gene | solute carrier family 35 (UDP-N-acetylglucosamine (UDP-GlcNAc) transporter), member 3 |
| 3 | gene | 116.6904 | 116.6934 | negative | MGI_C57BL6J_5623011 | Gm40126 | NCBI_Gene:105244533 | MGI:5623011 | lncRNA gene | predicted gene, 40126 |
| 3 | gene | 116.7400 | 116.8082 | negative | MGI_C57BL6J_1924809 | Agl | NCBI_Gene:77559,ENSEMBL:ENSMUSG00000033400 | MGI:1924809 | protein coding gene | amylo-1,6-glucosidase, 4-alpha-glucanotransferase |
| 3 | gene | 116.8188 | 116.8190 | positive | MGI_C57BL6J_5663657 | Gm43520 | ENSEMBL:ENSMUSG00000106128 | MGI:5663657 | unclassified gene | predicted gene 43520 |
| 3 | gene | 116.8223 | 116.8225 | negative | MGI_C57BL6J_5663658 | Gm43521 | ENSEMBL:ENSMUSG00000106511 | MGI:5663658 | unclassified gene | predicted gene 43521 |
| 3 | gene | 116.8230 | 116.8298 | positive | MGI_C57BL6J_5623012 | Gm40127 | NCBI_Gene:105244534 | MGI:5623012 | lncRNA gene | predicted gene, 40127 |
| 3 | gene | 116.8354 | 116.8415 | negative | MGI_C57BL6J_5623013 | Gm40128 | NCBI_Gene:105244535 | MGI:5623013 | lncRNA gene | predicted gene, 40128 |
| 3 | pseudogene | 116.8446 | 116.8450 | positive | MGI_C57BL6J_5588895 | Gm29736 | NCBI_Gene:101055867,ENSEMBL:ENSMUSG00000105250 | MGI:5588895 | pseudogene | predicted gene, 29736 |
| 3 | pseudogene | 116.8469 | 116.8473 | positive | MGI_C57BL6J_5663679 | Gm43542 | ENSEMBL:ENSMUSG00000105241 | MGI:5663679 | pseudogene | predicted gene 43542 |
| 3 | gene | 116.8595 | 116.9082 | positive | MGI_C57BL6J_108076 | Frrs1 | NCBI_Gene:20321,ENSEMBL:ENSMUSG00000033386 | MGI:108076 | protein coding gene | ferric-chelate reductase 1 |
| 3 | gene | 116.8695 | 116.8708 | positive | MGI_C57BL6J_1924370 | 9030201C23Rik | NA | NA | unclassified gene | RIKEN cDNA 9030201C23 gene |
| 3 | gene | 116.9183 | 116.9690 | negative | MGI_C57BL6J_2148896 | Palmd | NCBI_Gene:114301,ENSEMBL:ENSMUSG00000033377 | MGI:2148896 | protein coding gene | palmdelphin |
| 3 | gene | 116.9404 | 116.9443 | negative | MGI_C57BL6J_5663029 | Gm42892 | ENSEMBL:ENSMUSG00000106073 | MGI:5663029 | unclassified gene | predicted gene 42892 |
| 3 | gene | 116.9531 | 116.9601 | positive | MGI_C57BL6J_5591307 | Gm32148 | NCBI_Gene:102634607 | MGI:5591307 | lncRNA gene | predicted gene, 32148 |
| 3 | gene | 116.9683 | 116.9844 | positive | MGI_C57BL6J_1921904 | 4930455H04Rik | NCBI_Gene:74654,ENSEMBL:ENSMUSG00000080907 | MGI:1921904 | protein coding gene | RIKEN cDNA 4930455H04 gene |
| 3 | pseudogene | 116.9988 | 116.9996 | negative | MGI_C57BL6J_3643926 | Gm9332 | NCBI_Gene:668749,ENSEMBL:ENSMUSG00000105221 | MGI:3643926 | pseudogene | predicted gene 9332 |
| 3 | gene | 117.0605 | 117.0778 | negative | MGI_C57BL6J_1920612 | 1700061I17Rik | NCBI_Gene:433646,ENSEMBL:ENSMUSG00000028009 | MGI:1920612 | lncRNA gene | RIKEN cDNA 1700061I17 gene |
| 3 | pseudogene | 117.0741 | 117.0748 | negative | MGI_C57BL6J_5010081 | Gm17896 | NCBI_Gene:100416059 | MGI:5010081 | pseudogene | predicted gene, 17896 |
| 3 | pseudogene | 117.1008 | 117.1015 | negative | MGI_C57BL6J_5010615 | Gm18430 | NCBI_Gene:100417158,ENSEMBL:ENSMUSG00000106343 | MGI:5010615 | pseudogene | predicted gene, 18430 |
| 3 | pseudogene | 117.1759 | 117.1768 | positive | MGI_C57BL6J_3780039 | Gm9632 | NCBI_Gene:674908,ENSEMBL:ENSMUSG00000106314 | MGI:3780039 | pseudogene | predicted gene 9632 |
| 3 | pseudogene | 117.2229 | 117.2236 | negative | MGI_C57BL6J_5011428 | Gm19243 | NCBI_Gene:100462736,ENSEMBL:ENSMUSG00000105914 | MGI:5011428 | pseudogene | predicted gene, 19243 |
| 3 | gene | 117.2346 | 117.2562 | positive | MGI_C57BL6J_5591375 | Gm32216 | NCBI_Gene:102634688 | MGI:5591375 | lncRNA gene | predicted gene, 32216 |
| 3 | gene | 117.2510 | 117.2631 | negative | MGI_C57BL6J_5623015 | Gm40130 | NCBI_Gene:105244537 | MGI:5623015 | lncRNA gene | predicted gene, 40130 |
| 3 | gene | 117.2551 | 117.2558 | positive | MGI_C57BL6J_5663028 | Gm42891 | ENSEMBL:ENSMUSG00000106342 | MGI:5663028 | lncRNA gene | predicted gene 42891 |
| 3 | gene | 117.3191 | 117.3609 | negative | MGI_C57BL6J_106530 | Plppr4 | NCBI_Gene:229791,ENSEMBL:ENSMUSG00000044667 | MGI:106530 | protein coding gene | phospholipid phosphatase related 4 |
| 3 | gene | 117.3229 | 117.3230 | positive | MGI_C57BL6J_5452185 | Gm22408 | ENSEMBL:ENSMUSG00000092666 | MGI:5452185 | miRNA gene | predicted gene, 22408 |
| 3 | gene | 117.5710 | 117.6895 | positive | MGI_C57BL6J_1923019 | Plppr5 | NCBI_Gene:75769,ENSEMBL:ENSMUSG00000033342 | MGI:1923019 | protein coding gene | phospholipid phosphatase related 5 |
| 3 | gene | 117.7815 | 117.8689 | negative | MGI_C57BL6J_1923811 | Snx7 | NCBI_Gene:76561,ENSEMBL:ENSMUSG00000028007 | MGI:1923811 | protein coding gene | sorting nexin 7 |
| 3 | gene | 117.8486 | 117.8487 | negative | MGI_C57BL6J_5454588 | Gm24811 | ENSEMBL:ENSMUSG00000093054 | MGI:5454588 | miRNA gene | predicted gene, 24811 |
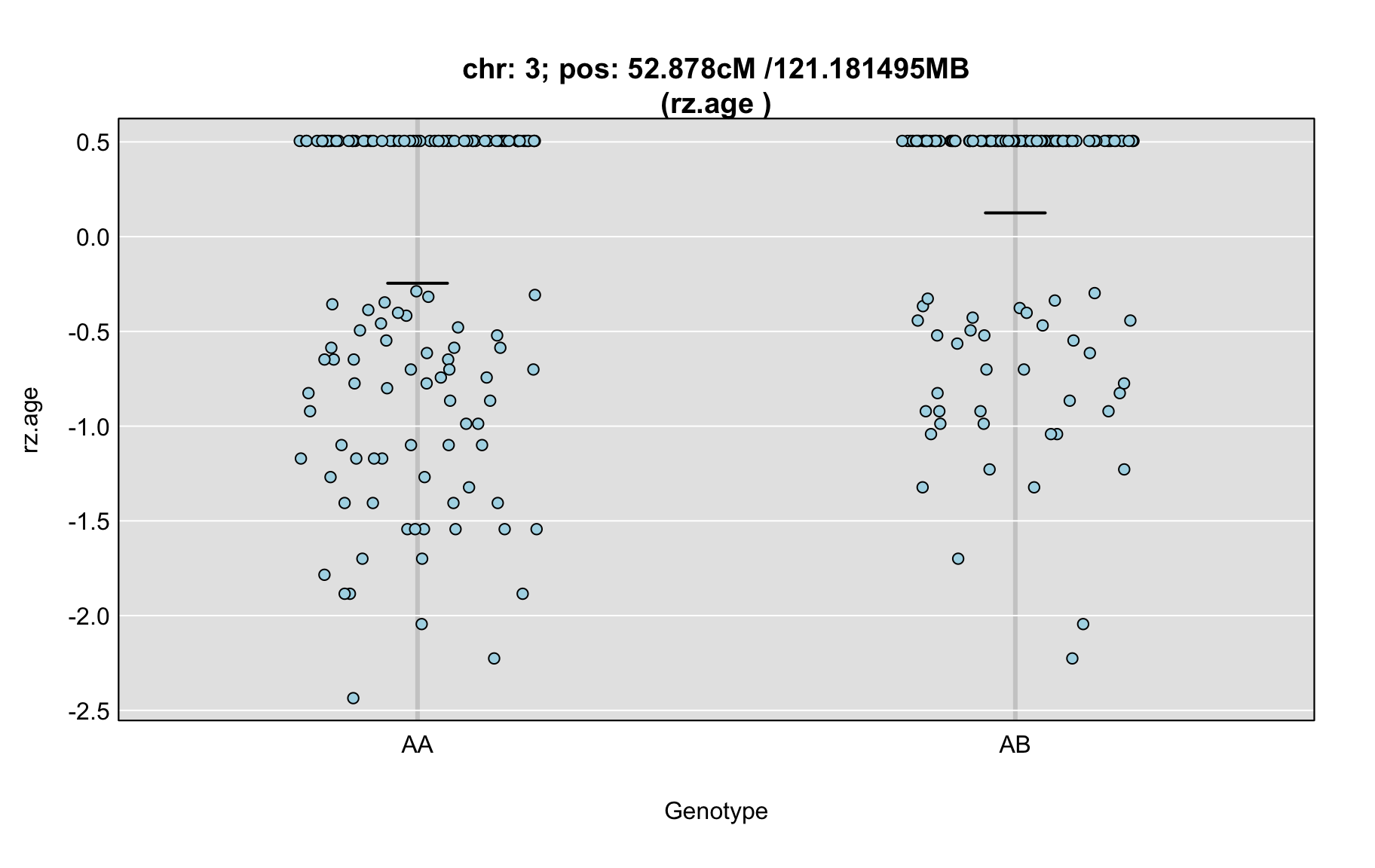
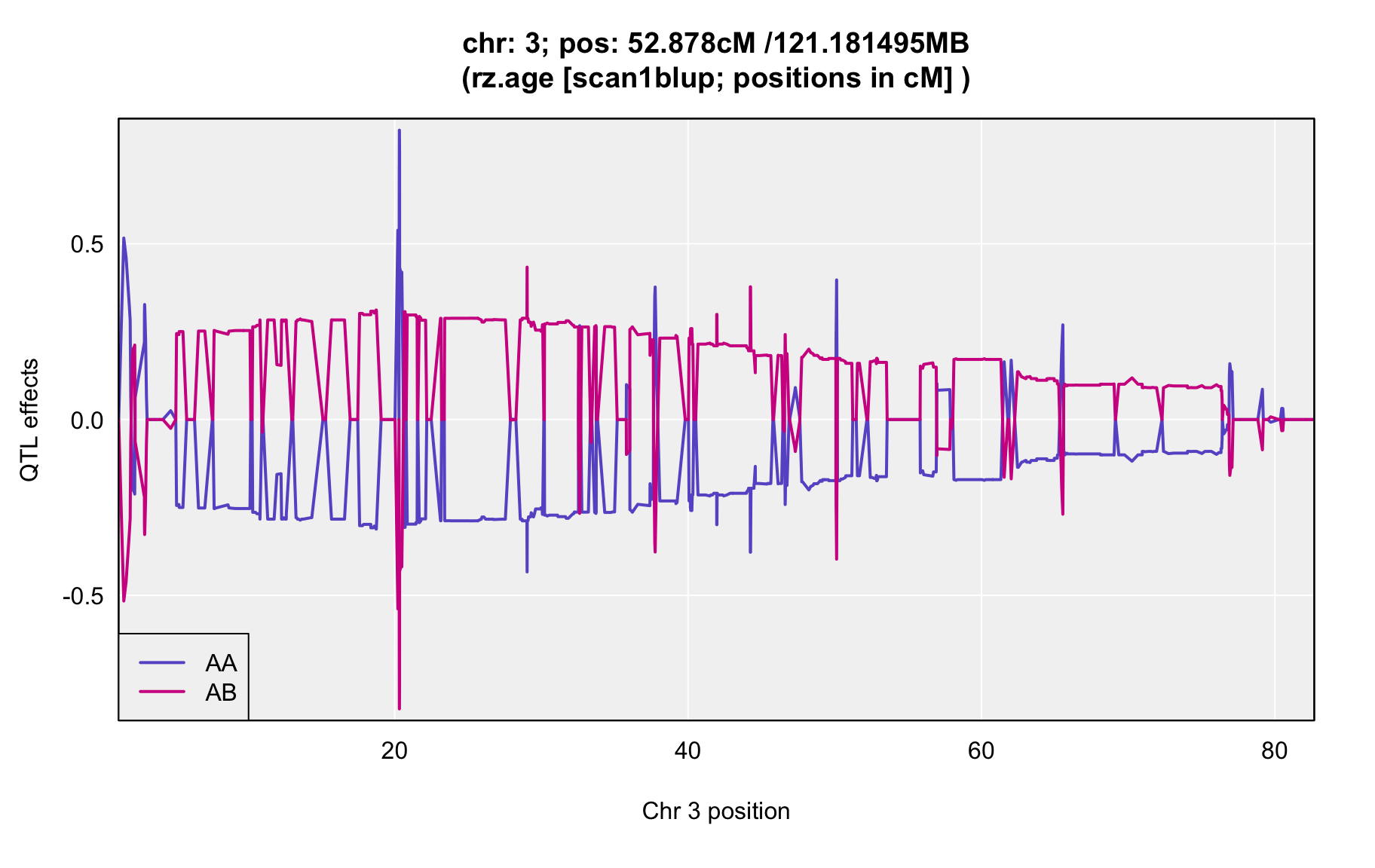
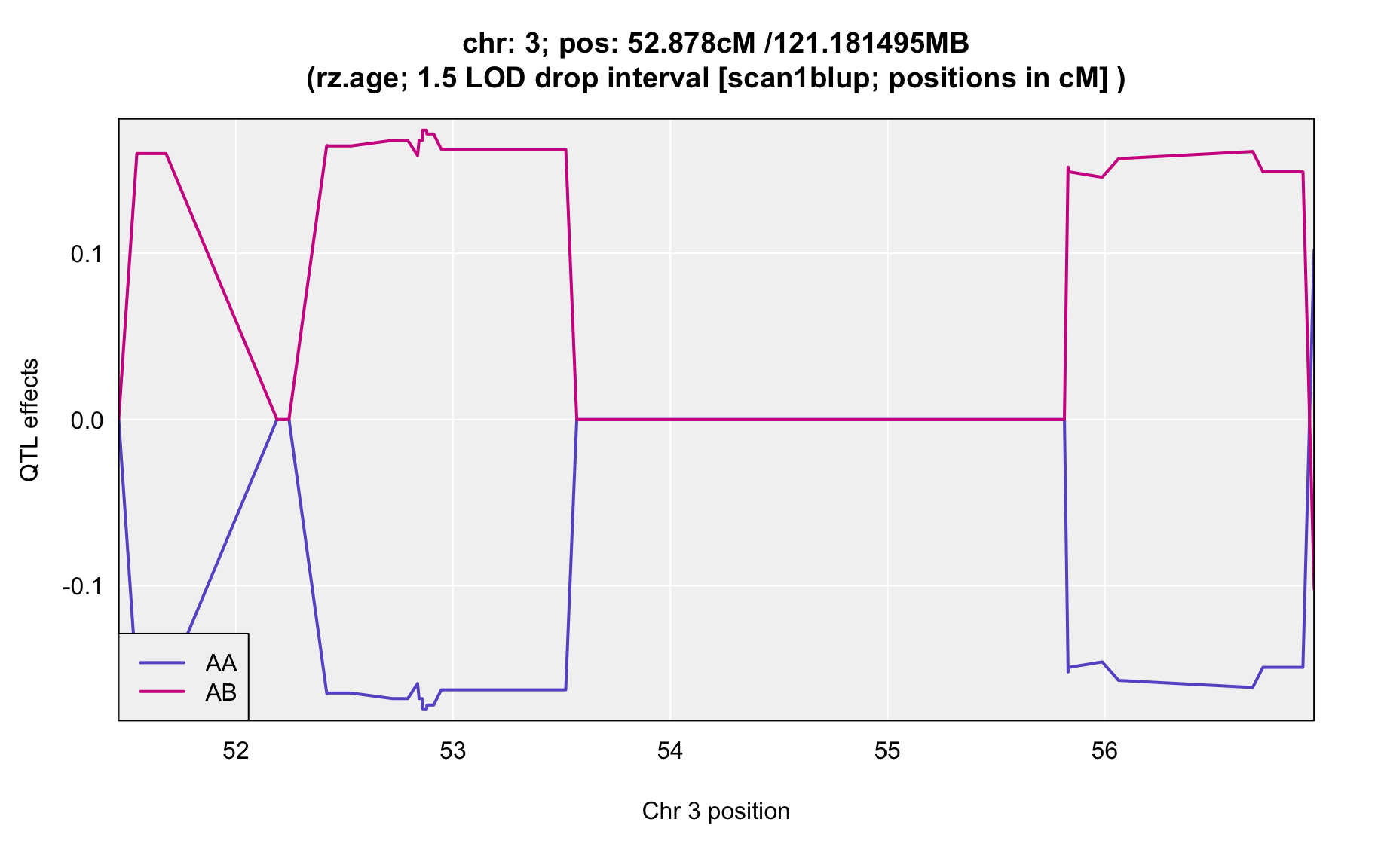
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 118.4303 | 118.5396 | positive | MGI_C57BL6J_5477365 | Gm26871 | NCBI_Gene:102634933,ENSEMBL:ENSMUSG00000097311 | MGI:5477365 | lncRNA gene | predicted gene, 26871 |
| 3 | gene | 118.4339 | 118.4339 | positive | MGI_C57BL6J_2676822 | Mir137 | miRBase:MI0000163,NCBI_Gene:387155,ENSEMBL:ENSMUSG00000065569 | MGI:2676822 | miRNA gene | microRNA 137 |
| 3 | gene | 118.4347 | 118.4348 | positive | MGI_C57BL6J_5453932 | Gm24155 | ENSEMBL:ENSMUSG00000093000 | MGI:5453932 | miRNA gene | predicted gene, 24155 |
| 3 | gene | 118.4348 | 118.4360 | positive | MGI_C57BL6J_3642633 | Gm9916 | ENSEMBL:ENSMUSG00000104178 | MGI:3642633 | lncRNA gene | predicted gene 9916 |
| 3 | gene | 118.5262 | 118.5283 | positive | MGI_C57BL6J_5611001 | Gm37773 | ENSEMBL:ENSMUSG00000104033 | MGI:5611001 | unclassified gene | predicted gene, 37773 |
| 3 | gene | 118.5309 | 118.5350 | positive | MGI_C57BL6J_5611156 | Gm37928 | ENSEMBL:ENSMUSG00000104362 | MGI:5611156 | unclassified gene | predicted gene, 37928 |
| 3 | gene | 118.5370 | 118.5405 | positive | MGI_C57BL6J_5591603 | Gm32444 | ENSEMBL:ENSMUSG00000102729 | MGI:5591603 | unclassified non-coding RNA gene | predicted gene, 32444 |
| 3 | gene | 118.5621 | 119.4329 | positive | MGI_C57BL6J_2139667 | Dpyd | NCBI_Gene:99586,ENSEMBL:ENSMUSG00000033308 | MGI:2139667 | protein coding gene | dihydropyrimidine dehydrogenase |
| 3 | gene | 118.8682 | 118.8705 | positive | MGI_C57BL6J_5663547 | Gm43410 | ENSEMBL:ENSMUSG00000106542 | MGI:5663547 | unclassified gene | predicted gene 43410 |
| 3 | gene | 119.4372 | 119.4404 | negative | MGI_C57BL6J_5591736 | Gm32577 | NCBI_Gene:102635174 | MGI:5591736 | lncRNA gene | predicted gene, 32577 |
| 3 | gene | 119.4568 | 119.4751 | negative | MGI_C57BL6J_5591791 | Gm32632 | NCBI_Gene:102635242 | MGI:5591791 | lncRNA gene | predicted gene, 32632 |
| 3 | gene | 119.4838 | 119.4900 | negative | MGI_C57BL6J_5623016 | Gm40131 | NCBI_Gene:105244538 | MGI:5623016 | lncRNA gene | predicted gene, 40131 |
| 3 | gene | 119.6387 | 119.6389 | negative | MGI_C57BL6J_5453209 | Gm23432 | ENSEMBL:ENSMUSG00000087806 | MGI:5453209 | snoRNA gene | predicted gene, 23432 |
| 3 | pseudogene | 119.7095 | 119.7099 | positive | MGI_C57BL6J_3782515 | Gm4332 | NCBI_Gene:100043278,ENSEMBL:ENSMUSG00000106037 | MGI:3782515 | pseudogene | predicted gene 4332 |
| 3 | gene | 119.7187 | 119.7845 | negative | MGI_C57BL6J_1860489 | Ptbp2 | NCBI_Gene:56195,ENSEMBL:ENSMUSG00000028134 | MGI:1860489 | protein coding gene | polypyrimidine tract binding protein 2 |
| 3 | gene | 119.7277 | 119.7298 | negative | MGI_C57BL6J_5663437 | Gm43300 | ENSEMBL:ENSMUSG00000105572 | MGI:5663437 | unclassified gene | predicted gene 43300 |
| 3 | gene | 119.7432 | 119.7464 | negative | MGI_C57BL6J_5662801 | Gm42664 | ENSEMBL:ENSMUSG00000106099 | MGI:5662801 | lncRNA gene | predicted gene 42664 |
| 3 | pseudogene | 119.7573 | 119.7575 | positive | MGI_C57BL6J_5662941 | Gm42804 | ENSEMBL:ENSMUSG00000106075 | MGI:5662941 | pseudogene | predicted gene 42804 |
| 3 | gene | 119.7838 | 119.7849 | positive | MGI_C57BL6J_5621433 | Gm38548 | NCBI_Gene:102641335 | MGI:5621433 | lncRNA gene | predicted gene, 38548 |
| 3 | gene | 119.8319 | 119.8321 | positive | MGI_C57BL6J_5454521 | Gm24744 | ENSEMBL:ENSMUSG00000094593 | MGI:5454521 | miRNA gene | predicted gene, 24744 |
| 3 | gene | 119.8379 | 119.8380 | positive | MGI_C57BL6J_5455914 | Gm26137 | ENSEMBL:ENSMUSG00000096196 | MGI:5455914 | miRNA gene | predicted gene, 26137 |
| 3 | pseudogene | 119.9687 | 119.9695 | positive | MGI_C57BL6J_5010569 | Gm18384 | NCBI_Gene:100417046,ENSEMBL:ENSMUSG00000104726 | MGI:5010569 | pseudogene | predicted gene, 18384 |
| 3 | gene | 120.3179 | 120.3180 | negative | MGI_C57BL6J_5453510 | Gm23733 | ENSEMBL:ENSMUSG00000065684 | MGI:5453510 | snRNA gene | predicted gene, 23733 |
| 3 | gene | 120.4543 | 120.8867 | negative | MGI_C57BL6J_2444638 | 6530403H02Rik | NCBI_Gene:320739,ENSEMBL:ENSMUSG00000098097 | MGI:2444638 | lncRNA gene | RIKEN cDNA 6530403H02 gene |
| 3 | gene | 120.6100 | 120.6101 | negative | MGI_C57BL6J_5454790 | Gm25013 | ENSEMBL:ENSMUSG00000089027 | MGI:5454790 | snoRNA gene | predicted gene, 25013 |
| 3 | pseudogene | 120.7357 | 120.7396 | negative | MGI_C57BL6J_5663064 | Gm42927 | ENSEMBL:ENSMUSG00000105207 | MGI:5663064 | pseudogene | predicted gene 42927 |
| 3 | gene | 120.9321 | 120.9347 | negative | MGI_C57BL6J_5623018 | Gm40133 | NCBI_Gene:105244540 | MGI:5623018 | lncRNA gene | predicted gene, 40133 |
| 3 | gene | 121.0205 | 121.0208 | negative | MGI_C57BL6J_5663581 | Gm43444 | ENSEMBL:ENSMUSG00000105243 | MGI:5663581 | unclassified gene | predicted gene 43444 |
| 3 | gene | 121.0291 | 121.0298 | negative | MGI_C57BL6J_5663063 | Gm42926 | ENSEMBL:ENSMUSG00000104749 | MGI:5663063 | unclassified gene | predicted gene 42926 |
| 3 | gene | 121.1554 | 121.1718 | negative | MGI_C57BL6J_1920420 | Rwdd3 | NCBI_Gene:66568,ENSEMBL:ENSMUSG00000028133 | MGI:1920420 | protein coding gene | RWD domain containing 3 |
| 3 | gene | 121.1687 | 121.1713 | positive | MGI_C57BL6J_3642648 | Gm10652 | ENSEMBL:ENSMUSG00000106329 | MGI:3642648 | unclassified gene | predicted gene 10652 |
| 3 | pseudogene | 121.1799 | 121.1811 | negative | MGI_C57BL6J_3643595 | Gm5710 | NCBI_Gene:435752,ENSEMBL:ENSMUSG00000098042 | MGI:3643595 | pseudogene | predicted gene 5710 |
| 3 | gene | 121.2018 | 121.2831 | negative | MGI_C57BL6J_1923195 | Tmem56 | NCBI_Gene:99887,ENSEMBL:ENSMUSG00000028132 | MGI:1923195 | protein coding gene | transmembrane protein 56 |
| 3 | pseudogene | 121.2500 | 121.2508 | negative | MGI_C57BL6J_3644601 | Gm9353 | NCBI_Gene:668778,ENSEMBL:ENSMUSG00000106619 | MGI:3644601 | pseudogene | predicted gene 9353 |
| 3 | pseudogene | 121.2775 | 121.2783 | positive | MGI_C57BL6J_5589676 | Gm30517 | ENSEMBL:ENSMUSG00000104902 | MGI:5589676 | pseudogene | predicted gene, 30517 |
| 3 | gene | 121.2917 | 121.3631 | positive | MGI_C57BL6J_1914039 | Alg14 | NCBI_Gene:66789,ENSEMBL:ENSMUSG00000039887 | MGI:1914039 | protein coding gene | asparagine-linked glycosylation 14 |
| 3 | gene | 121.3377 | 121.3380 | positive | MGI_C57BL6J_5455804 | Gm26027 | ENSEMBL:ENSMUSG00000084674 | MGI:5455804 | unclassified non-coding RNA gene | predicted gene, 26027 |
| 3 | gene | 121.3536 | 121.3613 | positive | MGI_C57BL6J_5592469 | Gm33310 | NCBI_Gene:102636167 | MGI:5592469 | lncRNA gene | predicted gene, 33310 |
| 3 | gene | 121.3690 | 121.3887 | positive | MGI_C57BL6J_5592522 | Gm33363 | NCBI_Gene:102636241 | MGI:5592522 | lncRNA gene | predicted gene, 33363 |
| 3 | pseudogene | 121.3801 | 121.3808 | negative | MGI_C57BL6J_3643596 | Gm5711 | NCBI_Gene:435753,ENSEMBL:ENSMUSG00000106455 | MGI:3643596 | pseudogene | predicted gene 5711 |
| 3 | gene | 121.3893 | 121.3927 | positive | MGI_C57BL6J_5623019 | Gm40134 | NCBI_Gene:105244541 | MGI:5623019 | lncRNA gene | predicted gene, 40134 |
| 3 | gene | 121.4242 | 121.4276 | negative | MGI_C57BL6J_5663065 | Gm42928 | ENSEMBL:ENSMUSG00000105813 | MGI:5663065 | unclassified gene | predicted gene 42928 |
| 3 | gene | 121.4265 | 121.4582 | positive | MGI_C57BL6J_1919244 | Cnn3 | NCBI_Gene:71994,ENSEMBL:ENSMUSG00000053931 | MGI:1919244 | protein coding gene | calponin 3, acidic |
| 3 | gene | 121.4280 | 121.4293 | positive | MGI_C57BL6J_5663066 | Gm42929 | ENSEMBL:ENSMUSG00000106022 | MGI:5663066 | unclassified gene | predicted gene 42929 |
| 3 | gene | 121.4595 | 121.5324 | negative | MGI_C57BL6J_2384860 | Slc44a3 | NCBI_Gene:213603,ENSEMBL:ENSMUSG00000039865 | MGI:2384860 | protein coding gene | solute carrier family 44, member 3 |
| 3 | gene | 121.5310 | 121.5325 | positive | MGI_C57BL6J_5621569 | Gm38684 | NCBI_Gene:105242407 | MGI:5621569 | lncRNA gene | predicted gene, 38684 |
| 3 | gene | 121.5316 | 121.5362 | positive | MGI_C57BL6J_2442825 | A530020G20Rik | NCBI_Gene:319839,ENSEMBL:ENSMUSG00000097124 | MGI:2442825 | lncRNA gene | RIKEN cDNA A530020G20 gene |
| 3 | gene | 121.5452 | 121.5627 | positive | MGI_C57BL6J_5623020 | Gm40135 | NCBI_Gene:105244542 | MGI:5623020 | lncRNA gene | predicted gene, 40135 |
| 3 | gene | 121.5642 | 121.5762 | negative | MGI_C57BL6J_5592706 | Gm33547 | NCBI_Gene:102636496 | MGI:5592706 | lncRNA gene | predicted gene, 33547 |
| 3 | gene | 121.5805 | 121.6066 | negative | MGI_C57BL6J_5592644 | Gm33485 | NCBI_Gene:102636411 | MGI:5592644 | lncRNA gene | predicted gene, 33485 |
| 3 | gene | 121.5929 | 121.5951 | positive | MGI_C57BL6J_5592867 | Gm33708 | NCBI_Gene:102636712 | MGI:5592867 | lncRNA gene | predicted gene, 33708 |
| 3 | gene | 121.6349 | 121.6465 | negative | MGI_C57BL6J_2442643 | A730020M07Rik | NCBI_Gene:100503044,ENSEMBL:ENSMUSG00000044522 | MGI:2442643 | lncRNA gene | RIKEN cDNA A730020M07 gene |
| 3 | gene | 121.6708 | 121.6830 | positive | MGI_C57BL6J_3588188 | 4930432M17Rik | NCBI_Gene:619318,ENSEMBL:ENSMUSG00000074248 | MGI:3588188 | protein coding gene | RIKEN cDNA 4930432M17 gene |
| 3 | gene | 121.7015 | 121.7102 | negative | MGI_C57BL6J_5663745 | Gm43608 | ENSEMBL:ENSMUSG00000106036 | MGI:5663745 | lncRNA gene | predicted gene 43608 |
| 3 | gene | 121.7116 | 121.7124 | negative | MGI_C57BL6J_5663744 | Gm43607 | ENSEMBL:ENSMUSG00000105767 | MGI:5663744 | lncRNA gene | predicted gene 43607 |
| 3 | gene | 121.7166 | 121.7234 | negative | MGI_C57BL6J_5623021 | Gm40136 | NCBI_Gene:105244543 | MGI:5623021 | lncRNA gene | predicted gene, 40136 |
| 3 | gene | 121.7216 | 121.7234 | negative | MGI_C57BL6J_5663960 | Gm43823 | ENSEMBL:ENSMUSG00000106224 | MGI:5663960 | unclassified gene | predicted gene 43823 |
| 3 | gene | 121.7235 | 121.7351 | positive | MGI_C57BL6J_88381 | F3 | NCBI_Gene:14066,ENSEMBL:ENSMUSG00000028128 | MGI:88381 | protein coding gene | coagulation factor III |
| 3 | gene | 121.7550 | 121.7586 | negative | MGI_C57BL6J_5662914 | Gm42777 | ENSEMBL:ENSMUSG00000104676 | MGI:5662914 | unclassified gene | predicted gene 42777 |
| 3 | gene | 121.7588 | 121.8153 | negative | MGI_C57BL6J_1349216 | Abcd3 | NCBI_Gene:19299,ENSEMBL:ENSMUSG00000028127 | MGI:1349216 | protein coding gene | ATP-binding cassette, sub-family D (ALD), member 3 |
| 3 | gene | 121.7622 | 121.7649 | negative | MGI_C57BL6J_5663746 | Gm43609 | ENSEMBL:ENSMUSG00000106087 | MGI:5663746 | unclassified gene | predicted gene 43609 |
| 3 | gene | 121.7875 | 121.7886 | negative | MGI_C57BL6J_1918078 | 4633401B06Rik | ENSEMBL:ENSMUSG00000105597 | MGI:1918078 | unclassified gene | RIKEN cDNA 4633401B06 gene |
| 3 | pseudogene | 121.8234 | 121.8238 | negative | MGI_C57BL6J_5662913 | Gm42776 | ENSEMBL:ENSMUSG00000105889 | MGI:5662913 | pseudogene | predicted gene 42776 |
| 3 | gene | 121.8461 | 121.8472 | negative | MGI_C57BL6J_5593083 | Gm33924 | NCBI_Gene:102637009 | MGI:5593083 | lncRNA gene | predicted gene, 33924 |
| 3 | gene | 121.8511 | 121.8552 | positive | MGI_C57BL6J_5593146 | Gm33987 | NCBI_Gene:102637085 | MGI:5593146 | lncRNA gene | predicted gene, 33987 |
| 3 | pseudogene | 121.8628 | 121.8632 | positive | MGI_C57BL6J_5662730 | Gm42593 | ENSEMBL:ENSMUSG00000104595 | MGI:5662730 | pseudogene | predicted gene 42593 |
| 3 | gene | 121.8961 | 121.9034 | negative | MGI_C57BL6J_5593209 | Gm34050 | NCBI_Gene:102637167 | MGI:5593209 | lncRNA gene | predicted gene, 34050 |
| 3 | gene | 121.9036 | 121.9358 | positive | MGI_C57BL6J_5593278 | Gm34119 | NCBI_Gene:102637254 | MGI:5593278 | lncRNA gene | predicted gene, 34119 |
| 3 | gene | 121.9269 | 121.9366 | negative | MGI_C57BL6J_5826458 | Gm46821 | NCBI_Gene:108168893 | MGI:5826458 | lncRNA gene | predicted gene, 46821 |
| 3 | gene | 121.9525 | 122.0168 | positive | MGI_C57BL6J_2443818 | Arhgap29 | NCBI_Gene:214137,ENSEMBL:ENSMUSG00000039831 | MGI:2443818 | protein coding gene | Rho GTPase activating protein 29 |
| 3 | gene | 122.0238 | 122.0271 | positive | MGI_C57BL6J_5662681 | Gm42544 | ENSEMBL:ENSMUSG00000104869 | MGI:5662681 | unclassified gene | predicted gene 42544 |
| 3 | gene | 122.0439 | 122.1801 | positive | MGI_C57BL6J_109424 | Abca4 | NCBI_Gene:11304,ENSEMBL:ENSMUSG00000028125 | MGI:109424 | protein coding gene | ATP-binding cassette, sub-family A (ABC1), member 4 |
| 3 | gene | 122.1055 | 122.1318 | negative | MGI_C57BL6J_5623023 | Gm40138 | NCBI_Gene:105244545 | MGI:5623023 | lncRNA gene | predicted gene, 40138 |
| 3 | pseudogene | 122.1251 | 122.2274 | negative | MGI_C57BL6J_5663295 | Gm43158 | ENSEMBL:ENSMUSG00000104887 | MGI:5663295 | pseudogene | predicted gene 43158 |
| 3 | pseudogene | 122.1522 | 122.1526 | negative | MGI_C57BL6J_5663294 | Gm43157 | ENSEMBL:ENSMUSG00000105351 | MGI:5663294 | pseudogene | predicted gene 43157 |
| 3 | pseudogene | 122.1835 | 122.1850 | negative | MGI_C57BL6J_3643912 | Gm9361 | NCBI_Gene:668788,ENSEMBL:ENSMUSG00000104799 | MGI:3643912 | pseudogene | predicted gene 9361 |
| 3 | gene | 122.1974 | 122.2454 | negative | MGI_C57BL6J_5826459 | Gm46822 | NCBI_Gene:108168894 | MGI:5826459 | lncRNA gene | predicted gene, 46822 |
| 3 | gene | 122.2114 | 122.2153 | positive | MGI_C57BL6J_5623022 | Gm40137 | NCBI_Gene:105244544 | MGI:5623022 | lncRNA gene | predicted gene, 40137 |
| 3 | pseudogene | 122.2300 | 122.2312 | negative | MGI_C57BL6J_3782792 | Gm4609 | NCBI_Gene:100043724,ENSEMBL:ENSMUSG00000078592 | MGI:3782792 | pseudogene | predicted gene 4609 |
| 3 | gene | 122.2325 | 122.2326 | negative | MGI_C57BL6J_5452680 | Gm22903 | ENSEMBL:ENSMUSG00000092934 | MGI:5452680 | miRNA gene | predicted gene, 22903 |
| 3 | gene | 122.2456 | 122.2707 | positive | MGI_C57BL6J_104995 | Gclm | NCBI_Gene:14630,ENSEMBL:ENSMUSG00000028124 | MGI:104995 | protein coding gene | glutamate-cysteine ligase, modifier subunit |
| 3 | gene | 122.2560 | 122.2937 | negative | MGI_C57BL6J_4937128 | Gm17494 | ENSEMBL:ENSMUSG00000057359 | MGI:4937128 | lncRNA gene | predicted gene, 17494 |
| 3 | pseudogene | 122.2567 | 122.2574 | positive | MGI_C57BL6J_3648621 | Gm6061 | NCBI_Gene:606529,ENSEMBL:ENSMUSG00000093392 | MGI:3648621 | pseudogene | predicted gene 6061 |
| 3 | gene | 122.2744 | 122.2853 | positive | MGI_C57BL6J_1923173 | Dnttip2 | NCBI_Gene:99480,ENSEMBL:ENSMUSG00000039756 | MGI:1923173 | protein coding gene | deoxynucleotidyltransferase, terminal, interacting protein 2 |
| 3 | gene | 122.2929 | 122.2930 | negative | MGI_C57BL6J_4413735 | n-TRtct2 | NCBI_Gene:102467253 | MGI:4413735 | tRNA gene | nuclear encoded tRNA arginine 2 (anticodon TCT) |
| 3 | gene | 122.2934 | 122.3052 | positive | MGI_C57BL6J_5623075 | Gm40190 | NCBI_Gene:105244600,ENSEMBL:ENSMUSG00000105057 | MGI:5623075 | lncRNA gene | predicted gene, 40190 |
| 3 | gene | 122.2936 | 122.2937 | negative | MGI_C57BL6J_3691613 | Mir760 | miRBase:MI0004605,NCBI_Gene:791077,ENSEMBL:ENSMUSG00000076456 | MGI:3691613 | miRNA gene | microRNA 760 |
| 3 | gene | 122.2941 | 122.5302 | positive | MGI_C57BL6J_1352501 | Bcar3 | NCBI_Gene:29815,ENSEMBL:ENSMUSG00000028121 | MGI:1352501 | protein coding gene | breast cancer anti-estrogen resistance 3 |
| 3 | gene | 122.3217 | 122.3317 | positive | MGI_C57BL6J_5623024 | Gm40139 | NCBI_Gene:105244546 | MGI:5623024 | lncRNA gene | predicted gene, 40139 |
| 3 | gene | 122.3771 | 122.3923 | negative | MGI_C57BL6J_3782793 | Gm4610 | NCBI_Gene:100043726,ENSEMBL:ENSMUSG00000104554 | MGI:3782793 | lncRNA gene | predicted gene 4610 |
| 3 | gene | 122.4264 | 122.4591 | negative | MGI_C57BL6J_5662973 | Gm42836 | ENSEMBL:ENSMUSG00000104574 | MGI:5662973 | lncRNA gene | predicted gene 42836 |
| 3 | gene | 122.4647 | 122.4649 | positive | MGI_C57BL6J_5454930 | Gm25153 | ENSEMBL:ENSMUSG00000087743 | MGI:5454930 | snRNA gene | predicted gene, 25153 |
| 3 | gene | 122.4765 | 122.4831 | negative | MGI_C57BL6J_5593508 | Gm34349 | NCBI_Gene:102637574 | MGI:5593508 | lncRNA gene | predicted gene, 34349 |
| 3 | gene | 122.5387 | 122.6197 | negative | MGI_C57BL6J_1925642 | Fnbp1l | NCBI_Gene:214459,ENSEMBL:ENSMUSG00000039735 | MGI:1925642 | protein coding gene | formin binding protein 1-like |
| 3 | gene | 122.5416 | 122.5417 | negative | MGI_C57BL6J_5562759 | Mir7657 | miRBase:MI0024997,NCBI_Gene:102465764,ENSEMBL:ENSMUSG00000104773 | MGI:5562759 | miRNA gene | microRNA 7657 |
| 3 | gene | 122.5650 | 122.5747 | positive | MGI_C57BL6J_5663706 | Gm43569 | ENSEMBL:ENSMUSG00000106194 | MGI:5663706 | lncRNA gene | predicted gene 43569 |
| 3 | gene | 122.5745 | 122.5771 | negative | MGI_C57BL6J_5663131 | Gm42994 | ENSEMBL:ENSMUSG00000106364 | MGI:5663131 | unclassified gene | predicted gene 42994 |
| 3 | gene | 122.6222 | 122.6302 | negative | MGI_C57BL6J_5663705 | Gm43568 | NCBI_Gene:108168895,ENSEMBL:ENSMUSG00000106365 | MGI:5663705 | lncRNA gene | predicted gene 43568 |
| 3 | pseudogene | 122.6326 | 122.6333 | negative | MGI_C57BL6J_3649175 | Gm5981 | NCBI_Gene:546810,ENSEMBL:ENSMUSG00000106562 | MGI:3649175 | pseudogene | predicted gene 5981 |
| 3 | gene | 122.6622 | 122.6841 | negative | MGI_C57BL6J_5623025 | Gm40140 | NCBI_Gene:105244547 | MGI:5623025 | lncRNA gene | predicted gene, 40140 |
| 3 | gene | 122.7289 | 122.8594 | positive | MGI_C57BL6J_2651499 | Pde5a | NCBI_Gene:242202,ENSEMBL:ENSMUSG00000053965 | MGI:2651499 | protein coding gene | phosphodiesterase 5A, cGMP-specific |
| 3 | gene | 122.7620 | 122.8020 | negative | MGI_C57BL6J_1918897 | 4930447N08Rik | NCBI_Gene:71647,ENSEMBL:ENSMUSG00000106583 | MGI:1918897 | lncRNA gene | RIKEN cDNA 4930447N08 gene |
| 3 | pseudogene | 122.7880 | 122.7887 | negative | MGI_C57BL6J_5593298 | Gm34139 | NCBI_Gene:102637287,ENSEMBL:ENSMUSG00000106376 | MGI:5593298 | pseudogene | predicted gene, 34139 |
| 3 | gene | 122.8125 | 122.8505 | negative | MGI_C57BL6J_5593736 | Gm34577 | NCBI_Gene:102637877 | MGI:5593736 | lncRNA gene | predicted gene, 34577 |
| 3 | pseudogene | 122.8619 | 122.8627 | positive | MGI_C57BL6J_3646838 | Gm9364 | NCBI_Gene:102637985,ENSEMBL:ENSMUSG00000106059 | MGI:3646838 | pseudogene | predicted gene 9364 |
| 3 | gene | 122.8647 | 122.9241 | negative | MGI_C57BL6J_1921307 | 4933405D12Rik | NCBI_Gene:74057,ENSEMBL:ENSMUSG00000097852 | MGI:1921307 | lncRNA gene | RIKEN cDNA 4933405D12 gene |
| 3 | gene | 122.8951 | 122.8995 | positive | MGI_C57BL6J_95478 | Fabp2 | NCBI_Gene:14079,ENSEMBL:ENSMUSG00000023057 | MGI:95478 | protein coding gene | fatty acid binding protein 2, intestinal |
| 3 | gene | 122.9131 | 122.9194 | negative | MGI_C57BL6J_3779169 | Gm10959 | ENSEMBL:ENSMUSG00000078590 | MGI:3779169 | protein coding gene | predicted gene 10959 |
| 3 | gene | 122.9242 | 122.9262 | positive | MGI_C57BL6J_1914954 | 1810037I17Rik | NCBI_Gene:67704,ENSEMBL:ENSMUSG00000054091 | MGI:1914954 | protein coding gene | RIKEN cDNA 1810037I17 gene |
| 3 | gene | 122.9315 | 122.9845 | negative | MGI_C57BL6J_2139607 | Usp53 | NCBI_Gene:99526,ENSEMBL:ENSMUSG00000039701 | MGI:2139607 | protein coding gene | ubiquitin specific peptidase 53 |
| 3 | gene | 122.9954 | 122.9990 | positive | MGI_C57BL6J_5663422 | Gm43285 | ENSEMBL:ENSMUSG00000105478 | MGI:5663422 | unclassified gene | predicted gene 43285 |
| 3 | gene | 123.0062 | 123.0350 | negative | MGI_C57BL6J_1913063 | Myoz2 | NCBI_Gene:59006,ENSEMBL:ENSMUSG00000028116 | MGI:1913063 | protein coding gene | myozenin 2 |
| 3 | gene | 123.0153 | 123.0154 | negative | MGI_C57BL6J_5451843 | Gm22066 | ENSEMBL:ENSMUSG00000076949 | MGI:5451843 | miRNA gene | predicted gene, 22066 |
| 3 | pseudogene | 123.0433 | 123.0437 | negative | MGI_C57BL6J_5663423 | Gm43286 | ENSEMBL:ENSMUSG00000106114 | MGI:5663423 | pseudogene | predicted gene 43286 |
| 3 | gene | 123.0765 | 123.2362 | negative | MGI_C57BL6J_2153070 | Synpo2 | NCBI_Gene:118449,ENSEMBL:ENSMUSG00000050315 | MGI:2153070 | protein coding gene | synaptopodin 2 |
| 3 | pseudogene | 123.1423 | 123.1430 | positive | MGI_C57BL6J_5663881 | Gm43744 | ENSEMBL:ENSMUSG00000106012 | MGI:5663881 | pseudogene | predicted gene 43744 |
| 3 | gene | 123.1639 | 123.1645 | negative | MGI_C57BL6J_5663424 | Gm43287 | ENSEMBL:ENSMUSG00000105562 | MGI:5663424 | unclassified gene | predicted gene 43287 |
| 3 | gene | 123.1901 | 123.1928 | negative | MGI_C57BL6J_5663425 | Gm43288 | ENSEMBL:ENSMUSG00000106220 | MGI:5663425 | unclassified gene | predicted gene 43288 |
| 3 | gene | 123.2094 | 123.2119 | negative | MGI_C57BL6J_5663420 | Gm43283 | ENSEMBL:ENSMUSG00000106490 | MGI:5663420 | unclassified gene | predicted gene 43283 |
| 3 | gene | 123.2675 | 123.3656 | positive | MGI_C57BL6J_1916858 | Sec24d | NCBI_Gene:69608,ENSEMBL:ENSMUSG00000039234 | MGI:1916858 | protein coding gene | Sec24 related gene family, member D (S. cerevisiae) |
| 3 | gene | 123.3212 | 123.3262 | negative | MGI_C57BL6J_5623027 | Gm40142 | NCBI_Gene:105244549 | MGI:5623027 | lncRNA gene | predicted gene, 40142 |
| 3 | gene | 123.3683 | 123.3861 | negative | MGI_C57BL6J_2442926 | Mettl14 | NCBI_Gene:210529,ENSEMBL:ENSMUSG00000028114 | MGI:2442926 | protein coding gene | methyltransferase like 14 |
| 3 | gene | 123.3911 | 123.3912 | negative | MGI_C57BL6J_5452490 | Gm22713 | ENSEMBL:ENSMUSG00000080545 | MGI:5452490 | snRNA gene | predicted gene, 22713 |
| 3 | gene | 123.4342 | 123.4379 | positive | MGI_C57BL6J_5623028 | Gm40143 | NCBI_Gene:105244550 | MGI:5623028 | lncRNA gene | predicted gene, 40143 |
| 3 | gene | 123.4412 | 123.4489 | negative | MGI_C57BL6J_5594224 | Gm35065 | NCBI_Gene:102638522,ENSEMBL:ENSMUSG00000105784 | MGI:5594224 | lncRNA gene | predicted gene, 35065 |
| 3 | gene | 123.4469 | 123.5066 | positive | MGI_C57BL6J_1100881 | Prss12 | NCBI_Gene:19142,ENSEMBL:ENSMUSG00000027978 | MGI:1100881 | protein coding gene | protease, serine 12 neurotrypsin (motopsin) |
| 3 | gene | 123.4720 | 123.4724 | positive | MGI_C57BL6J_5663727 | Gm43590 | ENSEMBL:ENSMUSG00000105058 | MGI:5663727 | unclassified gene | predicted gene 43590 |
| 3 | gene | 123.5076 | 123.5084 | negative | MGI_C57BL6J_1917145 | Snhg8 | NCBI_Gene:69895,ENSEMBL:ENSMUSG00000104960 | MGI:1917145 | lncRNA gene | small nucleolar RNA host gene 8 |
| 3 | gene | 123.5079 | 123.5081 | negative | MGI_C57BL6J_4360223 | Snora24 | NCBI_Gene:100306957,ENSEMBL:ENSMUSG00000092730 | MGI:4360223 | snoRNA gene | small nucleolar RNA, H/ACA box 24 |
| 3 | pseudogene | 123.5096 | 123.5107 | negative | MGI_C57BL6J_3645990 | Gm9369 | NCBI_Gene:668804,ENSEMBL:ENSMUSG00000104670 | MGI:3645990 | pseudogene | predicted pseudogene 9369 |
| 3 | gene | 123.5262 | 123.6909 | negative | MGI_C57BL6J_1932544 | Ndst3 | NCBI_Gene:83398,ENSEMBL:ENSMUSG00000027977 | MGI:1932544 | protein coding gene | N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 3 |
| 3 | gene | 123.6857 | 123.6880 | negative | MGI_C57BL6J_5662647 | Gm42510 | ENSEMBL:ENSMUSG00000104860 | MGI:5662647 | unclassified gene | predicted gene 42510 |
| 3 | gene | 123.7433 | 123.7434 | negative | MGI_C57BL6J_5454233 | Gm24456 | ENSEMBL:ENSMUSG00000065132 | MGI:5454233 | snRNA gene | predicted gene, 24456 |
| 3 | pseudogene | 123.7613 | 123.7619 | positive | MGI_C57BL6J_3705651 | Gm15400 | NCBI_Gene:108168856,ENSEMBL:ENSMUSG00000080842 | MGI:3705651 | pseudogene | predicted gene 15400 |
| 3 | pseudogene | 123.9841 | 123.9851 | negative | MGI_C57BL6J_3705526 | Gm15399 | ENSEMBL:ENSMUSG00000081939 | MGI:3705526 | pseudogene | predicted gene 15399 |
| 3 | gene | 124.0075 | 124.0290 | positive | MGI_C57BL6J_5623029 | Gm40144 | NCBI_Gene:105244551 | MGI:5623029 | lncRNA gene | predicted gene, 40144 |
| 3 | gene | 124.2462 | 124.3190 | positive | MGI_C57BL6J_5663868 | Gm43731 | ENSEMBL:ENSMUSG00000104531 | MGI:5663868 | lncRNA gene | predicted gene 43731 |
| 3 | pseudogene | 124.2920 | 124.2925 | positive | MGI_C57BL6J_3645996 | Gm9372 | NCBI_Gene:668809,ENSEMBL:ENSMUSG00000090474 | MGI:3645996 | pseudogene | predicted gene 9372 |
| 3 | pseudogene | 124.3042 | 124.3044 | negative | MGI_C57BL6J_5662961 | Gm42824 | ENSEMBL:ENSMUSG00000106238 | MGI:5662961 | pseudogene | predicted gene 42824 |
| 3 | gene | 124.3209 | 124.3247 | positive | MGI_C57BL6J_2443503 | Tram1l1 | NCBI_Gene:229801,ENSEMBL:ENSMUSG00000044528 | MGI:2443503 | protein coding gene | translocation associated membrane protein 1-like 1 |
| 3 | pseudogene | 124.3855 | 124.3867 | positive | MGI_C57BL6J_3704225 | Gm4617 | NCBI_Gene:100043740,ENSEMBL:ENSMUSG00000096544 | MGI:3704225 | pseudogene | predicted pseudogene 4617 |
| 3 | gene | 124.3981 | 124.3998 | negative | MGI_C57BL6J_1918179 | 4921527H02Rik | ENSEMBL:ENSMUSG00000106150 | MGI:1918179 | unclassified gene | RIKEN cDNA 4921527H02 gene |
| 3 | gene | 124.4010 | 124.4260 | negative | MGI_C57BL6J_1919074 | 1700006A11Rik | NCBI_Gene:71824,ENSEMBL:ENSMUSG00000027973 | MGI:1919074 | protein coding gene | RIKEN cDNA 1700006A11 gene |
| 3 | gene | 124.4107 | 124.4108 | positive | MGI_C57BL6J_5452352 | Gm22575 | ENSEMBL:ENSMUSG00000064915 | MGI:5452352 | snRNA gene | predicted gene, 22575 |
| 3 | gene | 124.4204 | 124.4221 | positive | MGI_C57BL6J_5662960 | Gm42823 | ENSEMBL:ENSMUSG00000106413 | MGI:5662960 | lncRNA gene | predicted gene 42823 |
| 3 | gene | 124.5016 | 124.5017 | positive | MGI_C57BL6J_5452677 | Gm22900 | ENSEMBL:ENSMUSG00000077726 | MGI:5452677 | snoRNA gene | predicted gene, 22900 |
| 3 | gene | 124.5564 | 124.5811 | negative | MGI_C57BL6J_1923637 | 1700003H04Rik | NCBI_Gene:384775,ENSEMBL:ENSMUSG00000039174 | MGI:1923637 | protein coding gene | RIKEN cDNA 1700003H04 gene |
| 3 | pseudogene | 124.5885 | 124.5892 | negative | MGI_C57BL6J_3643020 | Gm9379 | NCBI_Gene:668817,ENSEMBL:ENSMUSG00000106009 | MGI:3643020 | pseudogene | predicted gene 9379 |
| 3 | gene | 124.8395 | 124.9369 | positive | MGI_C57BL6J_5662962 | Gm42825 | ENSEMBL:ENSMUSG00000106423 | MGI:5662962 | lncRNA gene | predicted gene 42825 |
| 3 | gene | 124.8486 | 124.8489 | positive | MGI_C57BL6J_5453450 | Gm23673 | ENSEMBL:ENSMUSG00000088099 | MGI:5453450 | unclassified non-coding RNA gene | predicted gene, 23673 |
| 3 | pseudogene | 125.0421 | 125.0433 | negative | MGI_C57BL6J_3644106 | Gm9381 | NCBI_Gene:668820,ENSEMBL:ENSMUSG00000098033 | MGI:3644106 | pseudogene | predicted gene 9381 |
| 3 | gene | 125.2169 | 125.2196 | negative | MGI_C57BL6J_5662963 | Gm42826 | ENSEMBL:ENSMUSG00000106001 | MGI:5662963 | unclassified gene | predicted gene 42826 |
| 3 | gene | 125.4041 | 125.7289 | positive | MGI_C57BL6J_1932545 | Ndst4 | NCBI_Gene:64580,ENSEMBL:ENSMUSG00000027971 | MGI:1932545 | protein coding gene | N-deacetylase/N-sulfotransferase (heparin glucosaminyl) 4 |
| 3 | gene | 125.5201 | 125.5351 | negative | MGI_C57BL6J_5594396 | Gm35237 | NCBI_Gene:102638747 | MGI:5594396 | lncRNA gene | predicted gene, 35237 |
| 3 | gene | 125.6576 | 125.6813 | negative | MGI_C57BL6J_5594345 | Gm35186 | NCBI_Gene:102638681 | MGI:5594345 | lncRNA gene | predicted gene, 35186 |
| 3 | gene | 125.8653 | 125.9387 | negative | MGI_C57BL6J_109522 | Ugt8a | NCBI_Gene:22239,ENSEMBL:ENSMUSG00000032854 | MGI:109522 | protein coding gene | UDP galactosyltransferase 8A |
| 3 | gene | 125.8965 | 125.9006 | positive | MGI_C57BL6J_5623030 | Gm40145 | NCBI_Gene:105244552 | MGI:5623030 | lncRNA gene | predicted gene, 40145 |
| 3 | gene | 126.1067 | 126.1837 | positive | MGI_C57BL6J_5623031 | Gm40146 | NCBI_Gene:105244553 | MGI:5623031 | lncRNA gene | predicted gene, 40146 |
| 3 | gene | 126.1525 | 126.1527 | negative | MGI_C57BL6J_5456052 | Gm26275 | ENSEMBL:ENSMUSG00000087726 | MGI:5456052 | unclassified non-coding RNA gene | predicted gene, 26275 |
| 3 | gene | 126.2263 | 126.2284 | positive | MGI_C57BL6J_5662964 | Gm42827 | ENSEMBL:ENSMUSG00000106503 | MGI:5662964 | lncRNA gene | predicted gene 42827 |
| 3 | pseudogene | 126.2780 | 126.2793 | positive | MGI_C57BL6J_5011101 | Gm18916 | NCBI_Gene:100417950,ENSEMBL:ENSMUSG00000106094 | MGI:5011101 | pseudogene | predicted gene, 18916 |
| 3 | gene | 126.3637 | 126.4404 | positive | MGI_C57BL6J_2443513 | Arsj | NCBI_Gene:271970,ENSEMBL:ENSMUSG00000046561 | MGI:2443513 | protein coding gene | arylsulfatase J |
| 3 | gene | 126.3656 | 126.3808 | negative | MGI_C57BL6J_5594567 | Gm35408 | NCBI_Gene:102638976 | MGI:5594567 | lncRNA gene | predicted gene, 35408 |
| 3 | gene | 126.3863 | 126.4072 | negative | MGI_C57BL6J_5594510 | Gm35351 | NCBI_Gene:102638900 | MGI:5594510 | lncRNA gene | predicted gene, 35351 |
| 3 | gene | 126.4329 | 126.4393 | negative | MGI_C57BL6J_4938052 | Gm17225 | ENSEMBL:ENSMUSG00000090427 | MGI:4938052 | lncRNA gene | predicted gene 17225 |
| 3 | gene | 126.4410 | 126.4451 | positive | MGI_C57BL6J_5663464 | Gm43327 | ENSEMBL:ENSMUSG00000105695 | MGI:5663464 | unclassified gene | predicted gene 43327 |
| 3 | pseudogene | 126.4475 | 126.4480 | positive | MGI_C57BL6J_5295675 | Gm20568 | NCBI_Gene:108168897,ENSEMBL:ENSMUSG00000105233 | MGI:5295675 | pseudogene | predicted gene, 20568 |
| 3 | gene | 126.4683 | 126.5199 | positive | MGI_C57BL6J_5826460 | Gm46823 | NCBI_Gene:108168898 | MGI:5826460 | lncRNA gene | predicted gene, 46823 |
| 3 | gene | 126.5963 | 126.8463 | positive | MGI_C57BL6J_1341265 | Camk2d | NCBI_Gene:108058,ENSEMBL:ENSMUSG00000053819 | MGI:1341265 | protein coding gene | calcium/calmodulin-dependent protein kinase II, delta |
| 3 | gene | 126.6000 | 126.6045 | positive | MGI_C57BL6J_5663148 | Gm43011 | ENSEMBL:ENSMUSG00000106475 | MGI:5663148 | unclassified gene | predicted gene 43011 |
| 3 | gene | 126.6259 | 126.6288 | positive | MGI_C57BL6J_5663147 | Gm43010 | ENSEMBL:ENSMUSG00000104571 | MGI:5663147 | unclassified gene | predicted gene 43010 |
| 3 | gene | 126.6445 | 126.6480 | positive | MGI_C57BL6J_5663146 | Gm43009 | ENSEMBL:ENSMUSG00000105586 | MGI:5663146 | unclassified gene | predicted gene 43009 |
| 3 | gene | 126.6577 | 126.6593 | positive | MGI_C57BL6J_5663143 | Gm43006 | ENSEMBL:ENSMUSG00000105774 | MGI:5663143 | unclassified gene | predicted gene 43006 |
| 3 | gene | 126.6624 | 126.6630 | positive | MGI_C57BL6J_5663142 | Gm43005 | ENSEMBL:ENSMUSG00000104906 | MGI:5663142 | unclassified gene | predicted gene 43005 |
| 3 | gene | 126.6673 | 126.6699 | negative | MGI_C57BL6J_3783000 | Gm15551 | NCBI_Gene:102639086,ENSEMBL:ENSMUSG00000086679 | MGI:3783000 | lncRNA gene | predicted gene 15551 |
| 3 | gene | 126.7375 | 126.7411 | positive | MGI_C57BL6J_5662652 | Gm42515 | ENSEMBL:ENSMUSG00000105378 | MGI:5662652 | unclassified gene | predicted gene 42515 |
| 3 | gene | 126.7461 | 126.7480 | positive | MGI_C57BL6J_5662651 | Gm42514 | ENSEMBL:ENSMUSG00000105553 | MGI:5662651 | unclassified gene | predicted gene 42514 |
| 3 | gene | 126.8045 | 126.8069 | positive | MGI_C57BL6J_5633903 | C2dat1 | NA | NA | unclassified non-coding RNA gene | CAMK2D associated transcript 1 |
| 3 | gene | 126.9216 | 127.5003 | negative | MGI_C57BL6J_88025 | Ank2 | NCBI_Gene:109676,ENSEMBL:ENSMUSG00000032826 | MGI:88025 | protein coding gene | ankyrin 2, brain |
| 3 | gene | 126.9665 | 126.9675 | negative | MGI_C57BL6J_1925217 | A930036I15Rik | ENSEMBL:ENSMUSG00000105469 | MGI:1925217 | unclassified gene | RIKEN cDNA A930036I15 gene |
| 3 | gene | 126.9743 | 126.9759 | negative | MGI_C57BL6J_5663459 | Gm43322 | ENSEMBL:ENSMUSG00000105939 | MGI:5663459 | unclassified gene | predicted gene 43322 |
| 3 | gene | 126.9868 | 126.9877 | negative | MGI_C57BL6J_5663902 | Gm43765 | ENSEMBL:ENSMUSG00000105119 | MGI:5663902 | unclassified gene | predicted gene 43765 |
| 3 | gene | 127.0054 | 127.0055 | positive | MGI_C57BL6J_5455278 | Gm25501 | ENSEMBL:ENSMUSG00000077617 | MGI:5455278 | snoRNA gene | predicted gene, 25501 |
| 3 | gene | 127.0083 | 127.0108 | negative | MGI_C57BL6J_5663887 | Gm43750 | ENSEMBL:ENSMUSG00000106612 | MGI:5663887 | unclassified gene | predicted gene 43750 |
| 3 | gene | 127.0490 | 127.0520 | negative | MGI_C57BL6J_5663888 | Gm43751 | ENSEMBL:ENSMUSG00000105322 | MGI:5663888 | unclassified gene | predicted gene 43751 |
| 3 | gene | 127.1378 | 127.1405 | negative | MGI_C57BL6J_5663111 | Gm42974 | ENSEMBL:ENSMUSG00000104963 | MGI:5663111 | unclassified gene | predicted gene 42974 |
| 3 | gene | 127.1613 | 127.1683 | positive | MGI_C57BL6J_4439882 | Gm16958 | NCBI_Gene:100862268,ENSEMBL:ENSMUSG00000097456 | MGI:4439882 | lncRNA gene | predicted gene, 16958 |
| 3 | gene | 127.1803 | 127.1840 | negative | MGI_C57BL6J_2445192 | C230031I18Rik | ENSEMBL:ENSMUSG00000105202 | MGI:2445192 | unclassified gene | RIKEN cDNA C230031I18 gene |
| 3 | gene | 127.2406 | 127.2438 | negative | MGI_C57BL6J_5663110 | Gm42973 | ENSEMBL:ENSMUSG00000105733 | MGI:5663110 | unclassified gene | predicted gene 42973 |
| 3 | gene | 127.2667 | 127.2723 | negative | MGI_C57BL6J_5663109 | Gm42972 | ENSEMBL:ENSMUSG00000105445 | MGI:5663109 | unclassified gene | predicted gene 42972 |
| 3 | gene | 127.2835 | 127.2856 | negative | MGI_C57BL6J_5663108 | Gm42971 | ENSEMBL:ENSMUSG00000105568 | MGI:5663108 | unclassified gene | predicted gene 42971 |
| 3 | gene | 127.2983 | 127.2997 | negative | MGI_C57BL6J_5663107 | Gm42970 | ENSEMBL:ENSMUSG00000106446 | MGI:5663107 | unclassified gene | predicted gene 42970 |
| 3 | gene | 127.3901 | 127.3937 | negative | MGI_C57BL6J_5663106 | Gm42969 | ENSEMBL:ENSMUSG00000105822 | MGI:5663106 | unclassified gene | predicted gene 42969 |
| 3 | gene | 127.3939 | 127.4040 | negative | MGI_C57BL6J_5826461 | Gm46824 | NCBI_Gene:108168899 | MGI:5826461 | lncRNA gene | predicted gene, 46824 |
| 3 | gene | 127.4662 | 127.4708 | negative | MGI_C57BL6J_5594694 | Gm35535 | NCBI_Gene:102639156 | MGI:5594694 | lncRNA gene | predicted gene, 35535 |
| 3 | pseudogene | 127.4794 | 127.4814 | negative | MGI_C57BL6J_5663492 | Gm43355 | ENSEMBL:ENSMUSG00000106369 | MGI:5663492 | pseudogene | predicted gene 43355 |
| 3 | pseudogene | 127.4802 | 127.4812 | negative | MGI_C57BL6J_5662662 | Gm42525 | ENSEMBL:ENSMUSG00000105969 | MGI:5662662 | pseudogene | predicted gene 42525 |
| 3 | gene | 127.4811 | 127.4812 | positive | MGI_C57BL6J_5690886 | Gm44494 | ENSEMBL:ENSMUSG00000105116 | MGI:5690886 | miRNA gene | predicted gene, 44494 |
| 3 | pseudogene | 127.5129 | 127.5132 | positive | MGI_C57BL6J_5663491 | Gm43354 | ENSEMBL:ENSMUSG00000104957 | MGI:5663491 | pseudogene | predicted gene 43354 |
| 3 | gene | 127.5367 | 127.5533 | negative | MGI_C57BL6J_107634 | Larp7 | NCBI_Gene:28036,ENSEMBL:ENSMUSG00000027968 | MGI:107634 | protein coding gene | La ribonucleoprotein domain family, member 7 |
| 3 | gene | 127.5452 | 127.5453 | positive | MGI_C57BL6J_3619326 | Mir302b | miRBase:MI0003716,NCBI_Gene:723948,ENSEMBL:ENSMUSG00000076252 | MGI:3619326 | miRNA gene | microRNA 302b |
| 3 | gene | 127.5454 | 127.5454 | positive | MGI_C57BL6J_3619327 | Mir302c | miRBase:MI0003717,NCBI_Gene:723835,ENSEMBL:ENSMUSG00000065482 | MGI:3619327 | miRNA gene | microRNA 302c |
| 3 | gene | 127.5455 | 127.5456 | positive | MGI_C57BL6J_3619325 | Mir302a | miRBase:MI0000402,NCBI_Gene:723920,ENSEMBL:ENSMUSG00000065552 | MGI:3619325 | miRNA gene | microRNA 302a |
| 3 | gene | 127.5456 | 127.5457 | positive | MGI_C57BL6J_3619328 | Mir302d | miRBase:MI0003718,NCBI_Gene:723928,ENSEMBL:ENSMUSG00000065447 | MGI:3619328 | miRNA gene | microRNA 302d |
| 3 | gene | 127.5457 | 127.5458 | positive | MGI_C57BL6J_3619373 | Mir367 | miRBase:MI0003531,NCBI_Gene:723911,ENSEMBL:ENSMUSG00000065413 | MGI:3619373 | miRNA gene | microRNA 367 |
| 3 | gene | 127.5534 | 127.6180 | positive | MGI_C57BL6J_1918893 | Zgrf1 | NCBI_Gene:71643,ENSEMBL:ENSMUSG00000051278 | MGI:1918893 | protein coding gene | zinc finger, GRF-type containing 1 |
| 3 | gene | 127.5562 | 127.5576 | negative | MGI_C57BL6J_5663866 | Gm43729 | ENSEMBL:ENSMUSG00000105104 | MGI:5663866 | lncRNA gene | predicted gene 43729 |
| 3 | gene | 127.6262 | 127.6281 | negative | MGI_C57BL6J_5623032 | Gm40147 | NCBI_Gene:105244554 | MGI:5623032 | lncRNA gene | predicted gene, 40147 |
| 3 | gene | 127.6331 | 127.6356 | positive | MGI_C57BL6J_109619 | Neurog2 | NCBI_Gene:11924,ENSEMBL:ENSMUSG00000027967 | MGI:109619 | protein coding gene | neurogenin 2 |
| 3 | gene | 127.6703 | 127.7806 | negative | MGI_C57BL6J_1918731 | Alpk1 | NCBI_Gene:71481,ENSEMBL:ENSMUSG00000028028 | MGI:1918731 | protein coding gene | alpha-kinase 1 |
| 3 | pseudogene | 127.6919 | 127.6925 | negative | MGI_C57BL6J_3802104 | Gm16238 | NCBI_Gene:100191076,ENSEMBL:ENSMUSG00000089988 | MGI:3802104 | pseudogene | predicted gene 16238 |
| 3 | gene | 127.7146 | 127.7170 | negative | MGI_C57BL6J_1919269 | 1500005C15Rik | ENSEMBL:ENSMUSG00000104888 | MGI:1919269 | unclassified gene | RIKEN cDNA 1500005C15 gene |
| 3 | gene | 127.7172 | 127.7192 | negative | MGI_C57BL6J_5662724 | Gm42587 | ENSEMBL:ENSMUSG00000105701 | MGI:5662724 | unclassified gene | predicted gene 42587 |
| 3 | gene | 127.7316 | 127.7317 | positive | MGI_C57BL6J_5453056 | Gm23279 | ENSEMBL:ENSMUSG00000088537 | MGI:5453056 | rRNA gene | predicted gene, 23279 |
| 3 | gene | 127.7891 | 127.8322 | positive | MGI_C57BL6J_2182965 | Tifa | NCBI_Gene:211550,ENSEMBL:ENSMUSG00000046688 | MGI:2182965 | protein coding gene | TRAF-interacting protein with forkhead-associated domain |
| 3 | gene | 127.8070 | 127.8375 | negative | MGI_C57BL6J_2384822 | Ap1ar | NCBI_Gene:211556,ENSEMBL:ENSMUSG00000074238 | MGI:2384822 | protein coding gene | adaptor-related protein complex 1 associated regulatory protein |
| 3 | gene | 127.8296 | 127.8330 | negative | MGI_C57BL6J_5663791 | Gm43654 | ENSEMBL:ENSMUSG00000105804 | MGI:5663791 | unclassified gene | predicted gene 43654 |
| 3 | gene | 127.8369 | 127.8386 | positive | MGI_C57BL6J_5663790 | Gm43653 | ENSEMBL:ENSMUSG00000105447 | MGI:5663790 | lncRNA gene | predicted gene 43653 |
| 3 | gene | 127.8593 | 127.8676 | positive | MGI_C57BL6J_5594744 | Gm35585 | NCBI_Gene:102639226,ENSEMBL:ENSMUSG00000105550 | MGI:5594744 | lncRNA gene | predicted gene, 35585 |
| 3 | gene | 127.8632 | 127.8639 | negative | MGI_C57BL6J_5663789 | Gm43652 | ENSEMBL:ENSMUSG00000105008 | MGI:5663789 | unclassified gene | predicted gene 43652 |
| 3 | gene | 127.8691 | 127.8963 | negative | MGI_C57BL6J_1917867 | Fam241a | NCBI_Gene:70617,ENSEMBL:ENSMUSG00000050549 | MGI:1917867 | protein coding gene | family with sequence similarity 241, member A |
| 3 | gene | 127.9162 | 127.9552 | positive | MGI_C57BL6J_5439414 | 9830132P13Rik | NCBI_Gene:329753,ENSEMBL:ENSMUSG00000104763 | MGI:5439414 | lncRNA gene | RIKEN cDNA 9830132P13 gene |
| 3 | gene | 127.9707 | 127.9729 | positive | MGI_C57BL6J_5594826 | Gm35667 | NCBI_Gene:102639331,ENSEMBL:ENSMUSG00000106437 | MGI:5594826 | lncRNA gene | predicted gene, 35667 |
| 3 | gene | 128.0387 | 128.0407 | positive | MGI_C57BL6J_3642040 | Gm10650 | ENSEMBL:ENSMUSG00000074237 | MGI:3642040 | protein coding gene | predicted gene 10650 |
| 3 | gene | 128.1170 | 128.2316 | negative | MGI_C57BL6J_3028083 | D030025E07Rik | NCBI_Gene:402774,ENSEMBL:ENSMUSG00000105816 | MGI:3028083 | lncRNA gene | RIKEN cDNA D030025E07 gene |
| 3 | pseudogene | 128.2807 | 128.2811 | negative | MGI_C57BL6J_5663793 | Gm43656 | ENSEMBL:ENSMUSG00000106119 | MGI:5663793 | pseudogene | predicted gene 43656 |
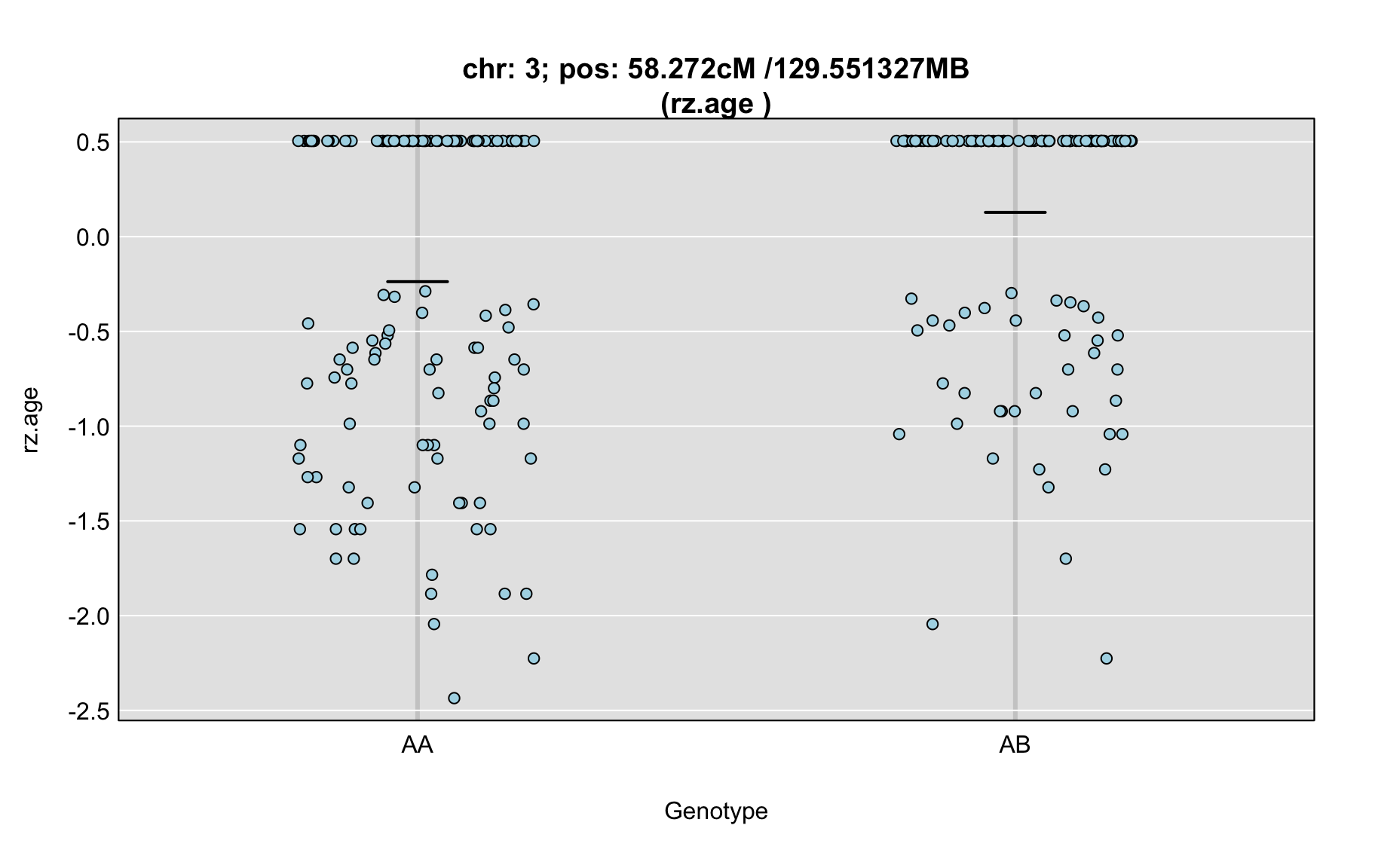
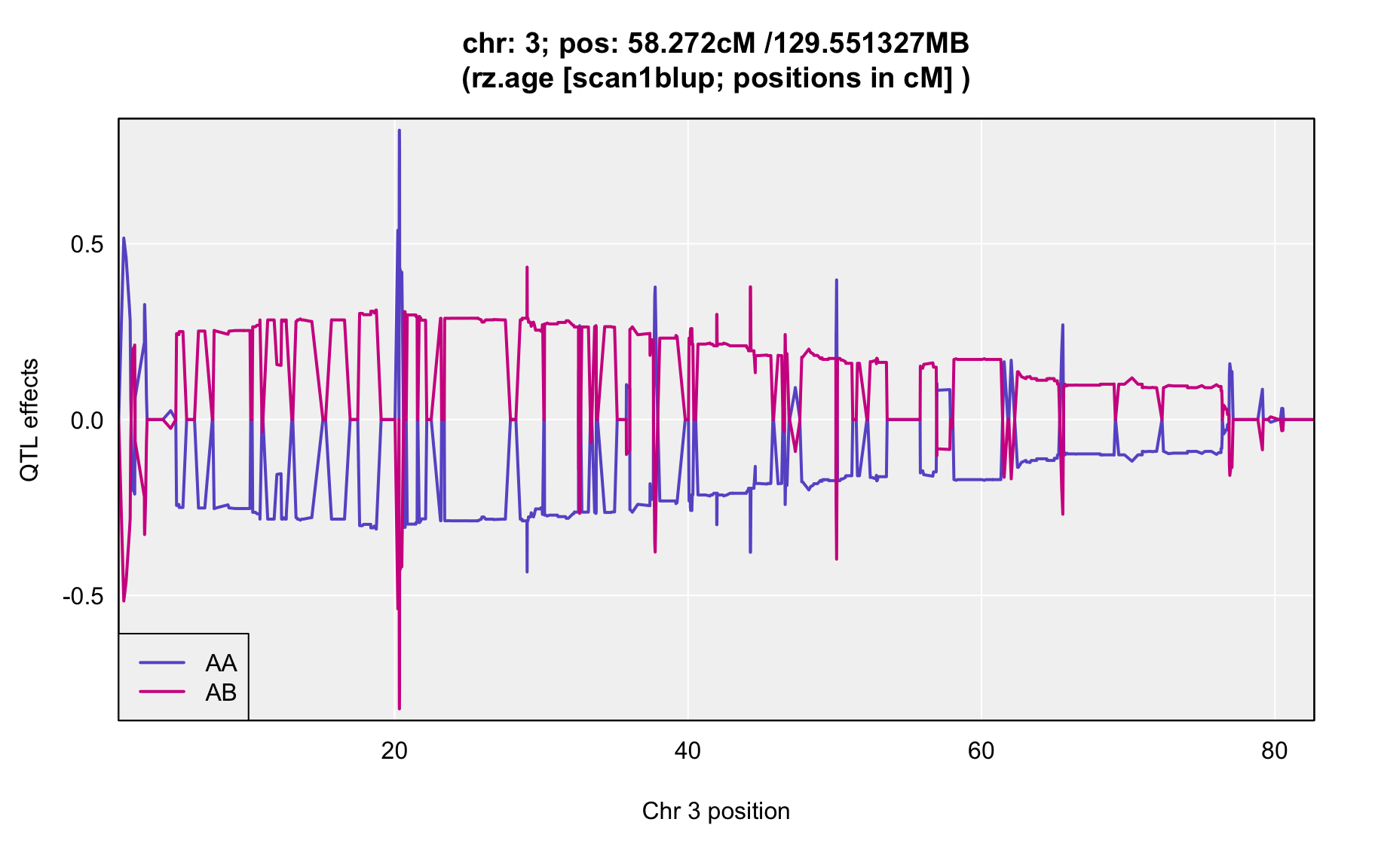
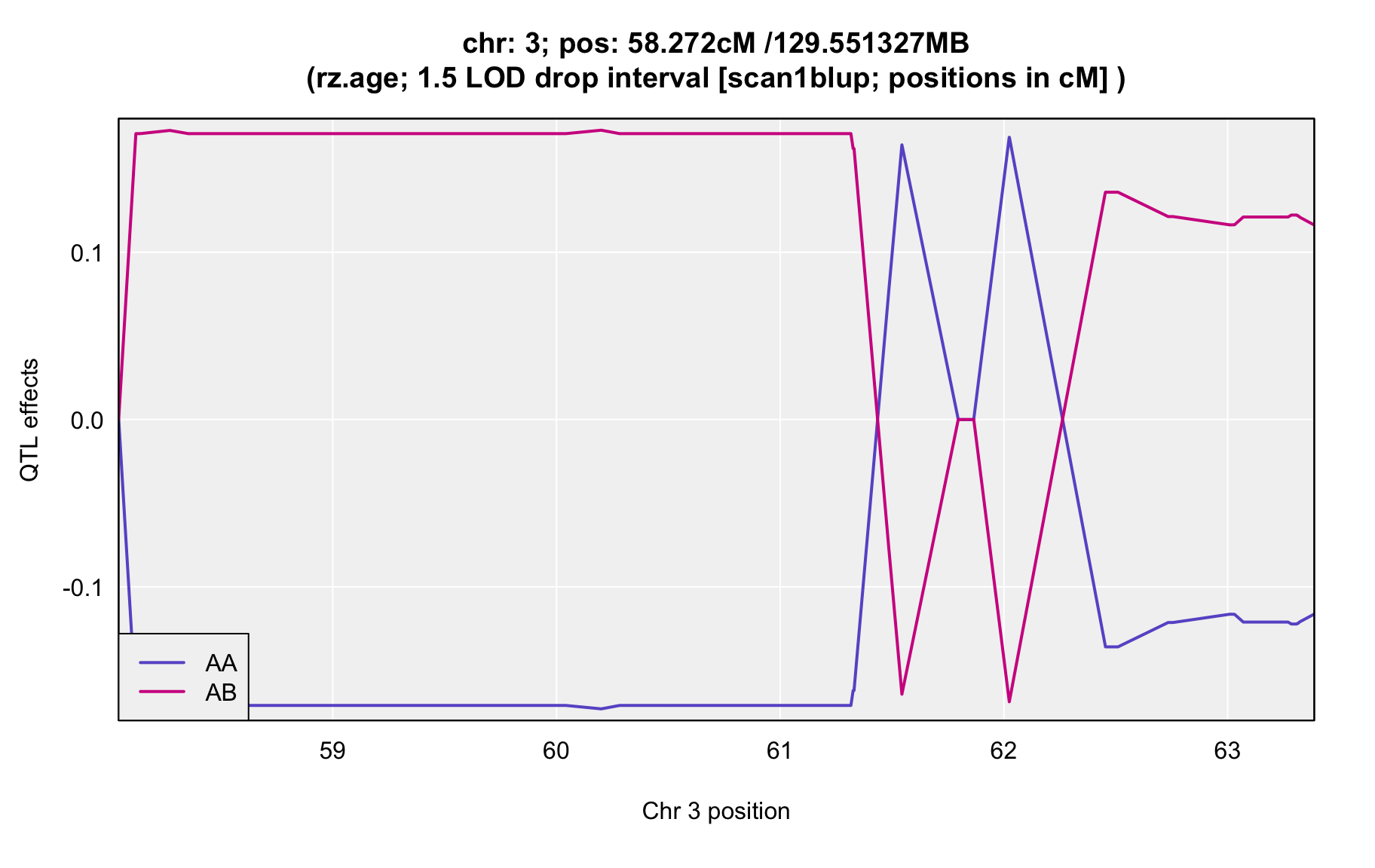
| chr | type | start | stop | strand | ID | Name | Dbxref | gene_id | mgi_type | description |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | gene | 129.5274 | 129.5324 | negative | MGI_C57BL6J_5595145 | Gm35986 | NCBI_Gene:102639747,ENSEMBL:ENSMUSG00000105837 | MGI:5595145 | lncRNA gene | predicted gene, 35986 |
| 3 | gene | 129.5324 | 129.6385 | positive | MGI_C57BL6J_2156528 | Elovl6 | NCBI_Gene:170439,ENSEMBL:ENSMUSG00000041220 | MGI:2156528 | protein coding gene | ELOVL family member 6, elongation of long chain fatty acids (yeast) |
| 3 | gene | 129.5381 | 129.5398 | positive | MGI_C57BL6J_5663209 | Gm43072 | ENSEMBL:ENSMUSG00000106092 | MGI:5663209 | unclassified gene | predicted gene 43072 |
| 3 | gene | 129.6776 | 129.7554 | negative | MGI_C57BL6J_95290 | Egf | NCBI_Gene:13645,ENSEMBL:ENSMUSG00000028017 | MGI:95290 | protein coding gene | epidermal growth factor |
| 3 | gene | 129.6841 | 129.6842 | negative | MGI_C57BL6J_5454292 | Gm24515 | ENSEMBL:ENSMUSG00000092701 | MGI:5454292 | miRNA gene | predicted gene, 24515 |
| 3 | gene | 129.7288 | 129.7402 | positive | MGI_C57BL6J_5662834 | Gm42697 | ENSEMBL:ENSMUSG00000106120 | MGI:5662834 | lncRNA gene | predicted gene 42697 |
| 3 | gene | 129.7453 | 129.7502 | positive | MGI_C57BL6J_5662787 | Gm42650 | ENSEMBL:ENSMUSG00000105596 | MGI:5662787 | lncRNA gene | predicted gene 42650 |
| 3 | gene | 129.7538 | 129.7639 | positive | MGI_C57BL6J_2441710 | 6330410L21Rik | NCBI_Gene:319224,ENSEMBL:ENSMUSG00000105960 | MGI:2441710 | lncRNA gene | RIKEN cDNA 6330410L21 gene |
| 3 | gene | 129.7879 | 129.8040 | negative | MGI_C57BL6J_2685267 | Lrit3 | NCBI_Gene:242235,ENSEMBL:ENSMUSG00000093865 | MGI:2685267 | protein coding gene | leucine-rich repeat, immunoglobulin-like and transmembrane domains 3 |
| 3 | gene | 129.7976 | 129.8156 | positive | MGI_C57BL6J_5595289 | Gm36130 | NCBI_Gene:102639932 | MGI:5595289 | lncRNA gene | predicted gene, 36130 |
| 3 | gene | 129.8044 | 129.8226 | negative | MGI_C57BL6J_1097709 | Rrh | NCBI_Gene:20132,ENSEMBL:ENSMUSG00000028012 | MGI:1097709 | protein coding gene | retinal pigment epithelium derived rhodopsin homolog |
| 3 | gene | 129.8249 | 129.8314 | negative | MGI_C57BL6J_1930948 | Gar1 | NCBI_Gene:68147,ENSEMBL:ENSMUSG00000028010 | MGI:1930948 | protein coding gene | GAR1 ribonucleoprotein |
| 3 | gene | 129.8359 | 129.8753 | positive | MGI_C57BL6J_105937 | Cfi | NCBI_Gene:12630,ENSEMBL:ENSMUSG00000058952 | MGI:105937 | protein coding gene | complement component factor i |
| 3 | gene | 129.8442 | 129.8472 | negative | MGI_C57BL6J_5595339 | Gm36180 | NCBI_Gene:102639999 | MGI:5595339 | lncRNA gene | predicted gene, 36180 |
| 3 | gene | 129.8786 | 129.8958 | positive | MGI_C57BL6J_1913600 | Pla2g12a | NCBI_Gene:66350,ENSEMBL:ENSMUSG00000027999 | MGI:1913600 | protein coding gene | phospholipase A2, group XIIA |
| 3 | gene | 129.8959 | 129.9014 | negative | MGI_C57BL6J_5826462 | Gm46825 | NCBI_Gene:108168900 | MGI:5826462 | lncRNA gene | predicted gene, 46825 |
| 3 | gene | 129.9014 | 129.9141 | positive | MGI_C57BL6J_1312921 | Casp6 | NCBI_Gene:12368,ENSEMBL:ENSMUSG00000027997 | MGI:1312921 | protein coding gene | caspase 6 |
| 3 | gene | 129.9065 | 129.9103 | positive | MGI_C57BL6J_5623035 | Gm40150 | NCBI_Gene:105244557 | MGI:5623035 | lncRNA gene | predicted gene, 40150 |
| 3 | gene | 129.9150 | 129.9702 | negative | MGI_C57BL6J_1914065 | Mcub | NCBI_Gene:66815,ENSEMBL:ENSMUSG00000027994 | MGI:1914065 | protein coding gene | mitochondrial calcium uniporter dominant negative beta subunit |
| 3 | gene | 129.9465 | 129.9472 | negative | MGI_C57BL6J_5623036 | Gm40151 | NCBI_Gene:105244558 | MGI:5623036 | lncRNA gene | predicted gene, 40151 |
| 3 | gene | 129.9828 | 130.0616 | negative | MGI_C57BL6J_2139764 | Sec24b | NCBI_Gene:99683,ENSEMBL:ENSMUSG00000001052 | MGI:2139764 | protein coding gene | Sec24 related gene family, member B (S. cerevisiae) |
| 3 | pseudogene | 130.0684 | 130.0690 | positive | MGI_C57BL6J_3645563 | Gm9396 | NCBI_Gene:668844,ENSEMBL:ENSMUSG00000063328 | MGI:3645563 | pseudogene | predicted gene 9396 |
| 3 | pseudogene | 130.0808 | 130.0811 | positive | MGI_C57BL6J_5662800 | Gm42663 | ENSEMBL:ENSMUSG00000106072 | MGI:5662800 | pseudogene | predicted gene 42663 |
| 3 | gene | 130.1315 | 130.1829 | positive | MGI_C57BL6J_1922224 | 4930502M04Rik | NA | NA | unclassified non-coding RNA gene | RIKEN cDNA 4930502M04 gene |
| 3 | gene | 130.1315 | 130.5999 | positive | MGI_C57BL6J_1924268 | Col25a1 | NCBI_Gene:77018,ENSEMBL:ENSMUSG00000058897 | MGI:1924268 | protein coding gene | collagen, type XXV, alpha 1 |
| 3 | pseudogene | 130.1404 | 130.1453 | positive | MGI_C57BL6J_3645564 | Gm9397 | NCBI_Gene:668845,ENSEMBL:ENSMUSG00000106276 | MGI:3645564 | pseudogene | predicted gene 9397 |
| 3 | gene | 130.2055 | 130.2056 | positive | MGI_C57BL6J_5452459 | Gm22682 | ENSEMBL:ENSMUSG00000065788 | MGI:5452459 | snRNA gene | predicted gene, 22682 |
| 3 | gene | 130.5395 | 130.5456 | positive | MGI_C57BL6J_5663347 | Gm43210 | ENSEMBL:ENSMUSG00000105011 | MGI:5663347 | unclassified gene | predicted gene 43210 |
| 3 | gene | 130.5619 | 130.5638 | positive | MGI_C57BL6J_5663348 | Gm43211 | ENSEMBL:ENSMUSG00000105760 | MGI:5663348 | unclassified gene | predicted gene 43211 |
| 3 | gene | 130.5921 | 130.5949 | positive | MGI_C57BL6J_5662974 | Gm42837 | ENSEMBL:ENSMUSG00000105520 | MGI:5662974 | unclassified gene | predicted gene 42837 |
| 3 | gene | 130.6174 | 130.6375 | positive | MGI_C57BL6J_1919010 | Etnppl | NCBI_Gene:71760,ENSEMBL:ENSMUSG00000019232 | MGI:1919010 | protein coding gene | ethanolamine phosphate phospholyase |
| 3 | gene | 130.6320 | 130.6334 | negative | MGI_C57BL6J_5595481 | Gm36322 | NCBI_Gene:102640195 | MGI:5595481 | lncRNA gene | predicted gene, 36322 |
| 3 | gene | 130.6459 | 130.6953 | negative | MGI_C57BL6J_5595532 | Gm36373 | NCBI_Gene:102640261 | MGI:5595532 | lncRNA gene | predicted gene, 36373 |
| 3 | gene | 130.6959 | 130.7094 | negative | MGI_C57BL6J_1913607 | Ostc | NCBI_Gene:66357,ENSEMBL:ENSMUSG00000041084 | MGI:1913607 | protein coding gene | oligosaccharyltransferase complex subunit (non-catalytic) |
| 3 | gene | 130.7036 | 130.7059 | negative | MGI_C57BL6J_5663135 | Gm42998 | ENSEMBL:ENSMUSG00000105466 | MGI:5663135 | unclassified gene | predicted gene 42998 |
| 3 | gene | 130.7268 | 130.7304 | negative | MGI_C57BL6J_1915686 | Rpl34 | NCBI_Gene:68436,ENSEMBL:ENSMUSG00000062006 | MGI:1915686 | protein coding gene | ribosomal protein L34 |
| 3 | gene | 130.7305 | 130.7716 | positive | MGI_C57BL6J_5663134 | Gm42997 | ENSEMBL:ENSMUSG00000105789 | MGI:5663134 | lncRNA gene | predicted gene 42997 |
| 3 | pseudogene | 130.7575 | 130.7578 | positive | MGI_C57BL6J_5663661 | Gm43524 | ENSEMBL:ENSMUSG00000106260 | MGI:5663661 | pseudogene | predicted gene 43524 |
| 3 | gene | 130.7593 | 130.7621 | positive | MGI_C57BL6J_5663659 | Gm43522 | ENSEMBL:ENSMUSG00000105716 | MGI:5663659 | lncRNA gene | predicted gene 43522 |
| 3 | gene | 130.7921 | 130.8040 | negative | MGI_C57BL6J_5595602 | Gm36443 | NCBI_Gene:102640363 | MGI:5595602 | lncRNA gene | predicted gene, 36443 |
| 3 | gene | 130.8385 | 130.9373 | negative | MGI_C57BL6J_1917148 | 2010110G14Rik | ENSEMBL:ENSMUSG00000106095 | MGI:1917148 | lncRNA gene | RIKEN cDNA 2010110G14 gene |
| 3 | pseudogene | 130.8464 | 130.8470 | negative | MGI_C57BL6J_3649176 | Gm5982 | NCBI_Gene:546811,ENSEMBL:ENSMUSG00000104984 | MGI:3649176 | pseudogene | predicted gene 5982 |
| 3 | gene | 130.8518 | 130.8688 | positive | MGI_C57BL6J_5595679 | Gm36520 | NCBI_Gene:102640465,ENSEMBL:ENSMUSG00000104557 | MGI:5595679 | lncRNA gene | predicted gene, 36520 |
| 3 | gene | 130.8709 | 130.8711 | negative | MGI_C57BL6J_5455867 | Gm26090 | ENSEMBL:ENSMUSG00000088522 | MGI:5455867 | snoRNA gene | predicted gene, 26090 |
| 3 | gene | 130.8741 | 130.8768 | positive | MGI_C57BL6J_5663488 | Gm43351 | ENSEMBL:ENSMUSG00000105769 | MGI:5663488 | lncRNA gene | predicted gene 43351 |
| 3 | gene | 130.8849 | 130.8856 | positive | MGI_C57BL6J_5663487 | Gm43350 | ENSEMBL:ENSMUSG00000105628 | MGI:5663487 | unclassified gene | predicted gene 43350 |
| 3 | gene | 130.9017 | 130.9030 | positive | MGI_C57BL6J_1923318 | 5830437K03Rik | ENSEMBL:ENSMUSG00000104579 | MGI:1923318 | unclassified gene | RIKEN cDNA 5830437K03 gene |
| 3 | gene | 130.9235 | 130.9248 | negative | MGI_C57BL6J_5663486 | Gm43349 | ENSEMBL:ENSMUSG00000106249 | MGI:5663486 | lncRNA gene | predicted gene 43349 |
| 3 | pseudogene | 130.9291 | 130.9308 | negative | MGI_C57BL6J_3645472 | Gm5855 | NCBI_Gene:545568,ENSEMBL:ENSMUSG00000105558 | MGI:3645472 | pseudogene | predicted gene 5855 |
| 3 | gene | 131.0238 | 131.0264 | negative | MGI_C57BL6J_2442445 | A430072C10Rik | ENSEMBL:ENSMUSG00000104859 | MGI:2442445 | unclassified gene | RIKEN cDNA A430072C10 gene |
| 3 | gene | 131.1016 | 131.1080 | negative | MGI_C57BL6J_5595762 | Gm36603 | NCBI_Gene:102640569 | MGI:5595762 | lncRNA gene | predicted gene, 36603 |
| 3 | gene | 131.1090 | 131.1121 | negative | MGI_C57BL6J_5663489 | Lef1os1 | ENSEMBL:ENSMUSG00000106086 | MGI:5663489 | antisense lncRNA gene | LEF1 opposite strand RNA 1 |
| 3 | gene | 131.1103 | 131.2244 | positive | MGI_C57BL6J_96770 | Lef1 | NCBI_Gene:16842,ENSEMBL:ENSMUSG00000027985 | MGI:96770 | protein coding gene | lymphoid enhancer binding factor 1 |
| 3 | gene | 131.1368 | 131.1390 | positive | MGI_C57BL6J_5662586 | Gm42449 | ENSEMBL:ENSMUSG00000104612 | MGI:5662586 | unclassified gene | predicted gene 42449 |
| 3 | gene | 131.1918 | 131.1930 | positive | MGI_C57BL6J_1925142 | 6720418B01Rik | NA | NA | unclassified gene | RIKEN cDNA 6720418B01 gene |
| 3 | gene | 131.2334 | 131.2721 | negative | MGI_C57BL6J_96009 | Hadh | NCBI_Gene:15107,ENSEMBL:ENSMUSG00000027984 | MGI:96009 | protein coding gene | hydroxyacyl-Coenzyme A dehydrogenase |
| 3 | pseudogene | 131.2841 | 131.2851 | negative | MGI_C57BL6J_5010677 | Gm18492 | NCBI_Gene:100417267,ENSEMBL:ENSMUSG00000106268 | MGI:5010677 | pseudogene | predicted gene, 18492 |
| 3 | gene | 131.2884 | 131.3032 | negative | MGI_C57BL6J_1918769 | Cyp2u1 | NCBI_Gene:71519,ENSEMBL:ENSMUSG00000027983 | MGI:1918769 | protein coding gene | cytochrome P450, family 2, subfamily u, polypeptide 1 |
| 3 | gene | 131.3026 | 131.3172 | positive | MGI_C57BL6J_1925437 | 4930534D22Rik | NCBI_Gene:100503594,ENSEMBL:ENSMUSG00000097761 | MGI:1925437 | lncRNA gene | RIKEN cDNA 4930534D22 gene |
| 3 | gene | 131.3190 | 131.4914 | negative | MGI_C57BL6J_1921692 | Sgms2 | NCBI_Gene:74442,ENSEMBL:ENSMUSG00000050931 | MGI:1921692 | protein coding gene | sphingomyelin synthase 2 |
| 3 | pseudogene | 131.3605 | 131.3609 | positive | MGI_C57BL6J_5663253 | Gm43116 | NCBI_Gene:108168901,ENSEMBL:ENSMUSG00000104754 | MGI:5663253 | pseudogene | predicted gene 43116 |
| 3 | gene | 131.4906 | 131.4992 | positive | MGI_C57BL6J_5595912 | Gm36753 | NCBI_Gene:102640761 | MGI:5595912 | lncRNA gene | predicted gene, 36753 |
| 3 | gene | 131.5091 | 131.5111 | positive | MGI_C57BL6J_5596022 | Gm36863 | NCBI_Gene:102640912 | MGI:5596022 | lncRNA gene | predicted gene, 36863 |
| 3 | gene | 131.5648 | 131.6437 | positive | MGI_C57BL6J_1330587 | Papss1 | NCBI_Gene:23971,ENSEMBL:ENSMUSG00000028032 | MGI:1330587 | protein coding gene | 3’-phosphoadenosine 5’-phosphosulfate synthase 1 |
| 3 | gene | 131.5876 | 131.5895 | positive | MGI_C57BL6J_5596076 | Gm36917 | NCBI_Gene:102640987 | MGI:5596076 | lncRNA gene | predicted gene, 36917 |
| 3 | gene | 131.6104 | 131.6138 | negative | MGI_C57BL6J_5663018 | Gm42881 | ENSEMBL:ENSMUSG00000105973 | MGI:5663018 | unclassified gene | predicted gene 42881 |
| 3 | gene | 131.7598 | 132.0307 | positive | MGI_C57BL6J_5589024 | Gm29865 | NCBI_Gene:102631558,ENSEMBL:ENSMUSG00000105331 | MGI:5589024 | lncRNA gene | predicted gene, 29865 |
| 3 | pseudogene | 131.8991 | 131.8995 | negative | MGI_C57BL6J_3030209 | Olfr375-ps1 | NCBI_Gene:257879,ENSEMBL:ENSMUSG00000104853 | MGI:3030209 | pseudogene | olfactory receptor 375, pseudogene 1 |
| 3 | gene | 131.9183 | 131.9184 | positive | MGI_C57BL6J_5455880 | Gm26103 | ENSEMBL:ENSMUSG00000087877 | MGI:5455880 | snRNA gene | predicted gene, 26103 |
| 3 | gene | 132.0073 | 132.0582 | positive | MGI_C57BL6J_5589108 | Gm29949 | NCBI_Gene:102631666 | MGI:5589108 | lncRNA gene | predicted gene, 29949 |
| 3 | gene | 132.0823 | 132.0824 | negative | MGI_C57BL6J_5452198 | Gm22421 | ENSEMBL:ENSMUSG00000064992 | MGI:5452198 | snRNA gene | predicted gene, 22421 |
| 3 | gene | 132.0852 | 132.1803 | positive | MGI_C57BL6J_1890663 | Dkk2 | NCBI_Gene:56811,ENSEMBL:ENSMUSG00000028031 | MGI:1890663 | protein coding gene | dickkopf WNT signaling pathway inhibitor 2 |
| 3 | gene | 132.4437 | 132.4455 | negative | MGI_C57BL6J_5663017 | Gm42880 | ENSEMBL:ENSMUSG00000106280 | MGI:5663017 | lncRNA gene | predicted gene 42880 |
| 3 | gene | 132.4796 | 132.5004 | positive | MGI_C57BL6J_5589306 | Gm30147 | NCBI_Gene:102631945 | MGI:5589306 | lncRNA gene | predicted gene, 30147 |
| 3 | gene | 132.5376 | 132.5377 | negative | MGI_C57BL6J_5455084 | Gm25307 | ENSEMBL:ENSMUSG00000089510 | MGI:5455084 | snRNA gene | predicted gene, 25307 |
| 3 | gene | 132.6298 | 132.6478 | positive | MGI_C57BL6J_3647547 | Gimd1 | NCBI_Gene:433653,ENSEMBL:ENSMUSG00000091721 | MGI:3647547 | protein coding gene | GIMAP family P-loop NTPase domain containing 1 |
| 3 | pseudogene | 132.6551 | 132.6563 | negative | MGI_C57BL6J_3648832 | Gm9402 | NCBI_Gene:668857,ENSEMBL:ENSMUSG00000106313 | MGI:3648832 | pseudogene | predicted gene 9402 |
| 3 | gene | 132.6605 | 132.6844 | negative | MGI_C57BL6J_102774 | Aimp1 | NCBI_Gene:13722,ENSEMBL:ENSMUSG00000028029 | MGI:102774 | protein coding gene | aminoacyl tRNA synthetase complex-interacting multifunctional protein 1 |
| 3 | gene | 132.6840 | 132.8417 | positive | MGI_C57BL6J_2445052 | Tbck | NCBI_Gene:271981,ENSEMBL:ENSMUSG00000028030 | MGI:2445052 | protein coding gene | TBC1 domain containing kinase |
| 3 | gene | 132.7674 | 132.7694 | positive | MGI_C57BL6J_5663009 | Gm42872 | ENSEMBL:ENSMUSG00000105719 | MGI:5663009 | unclassified gene | predicted gene 42872 |
| 3 | gene | 132.7721 | 132.7829 | negative | MGI_C57BL6J_5589453 | Gm30294 | NCBI_Gene:102632141 | MGI:5589453 | lncRNA gene | predicted gene, 30294 |
| 3 | gene | 132.7872 | 132.7873 | negative | MGI_C57BL6J_5452099 | Gm22322 | ENSEMBL:ENSMUSG00000087776 | MGI:5452099 | snoRNA gene | predicted gene, 22322 |
| 3 | gene | 132.8063 | 132.8095 | positive | MGI_C57BL6J_5663013 | Gm42876 | ENSEMBL:ENSMUSG00000105691 | MGI:5663013 | unclassified gene | predicted gene 42876 |
| 3 | gene | 132.8422 | 132.8786 | negative | MGI_C57BL6J_5588970 | Gm29811 | NCBI_Gene:101243624,ENSEMBL:ENSMUSG00000106230 | MGI:5588970 | lncRNA gene | predicted gene, 29811 |
| 3 | pseudogene | 132.8423 | 132.8450 | positive | MGI_C57BL6J_3648825 | Gm9403 | NCBI_Gene:668858,ENSEMBL:ENSMUSG00000106499 | MGI:3648825 | pseudogene | predicted gene 9403 |
| 3 | gene | 132.8817 | 132.9503 | negative | MGI_C57BL6J_2148811 | Npnt | NCBI_Gene:114249,ENSEMBL:ENSMUSG00000040998 | MGI:2148811 | protein coding gene | nephronectin |
| 3 | gene | 132.9399 | 132.9400 | positive | MGI_C57BL6J_5452973 | Gm23196 | ENSEMBL:ENSMUSG00000089431 | MGI:5452973 | snRNA gene | predicted gene, 23196 |
| 3 | gene | 132.9818 | 133.0920 | negative | MGI_C57BL6J_1914803 | Gstcd | NCBI_Gene:67553,ENSEMBL:ENSMUSG00000028018 | MGI:1914803 | protein coding gene | glutathione S-transferase, C-terminal domain containing |
| 3 | pseudogene | 133.0293 | 133.0299 | positive | MGI_C57BL6J_5011333 | Gm19148 | NCBI_Gene:100418338,ENSEMBL:ENSMUSG00000104787 | MGI:5011333 | pseudogene | predicted gene, 19148 |
| 3 | gene | 133.0506 | 133.0524 | negative | MGI_C57BL6J_5663011 | Gm42874 | ENSEMBL:ENSMUSG00000105259 | MGI:5663011 | unclassified gene | predicted gene 42874 |
| 3 | gene | 133.0525 | 133.0543 | negative | MGI_C57BL6J_5663010 | Gm42873 | ENSEMBL:ENSMUSG00000105217 | MGI:5663010 | unclassified gene | predicted gene 42873 |
| 3 | gene | 133.0918 | 133.1110 | positive | MGI_C57BL6J_1919043 | Ints12 | NCBI_Gene:71793,ENSEMBL:ENSMUSG00000028016 | MGI:1919043 | protein coding gene | integrator complex subunit 12 |
| 3 | gene | 133.1123 | 133.2349 | negative | MGI_C57BL6J_1924919 | Arhgef38 | NCBI_Gene:77669,ENSEMBL:ENSMUSG00000040969 | MGI:1924919 | protein coding gene | Rho guanine nucleotide exchange factor (GEF) 38 |
| 3 | gene | 133.3075 | 133.3100 | negative | MGI_C57BL6J_5663391 | Gm43254 | ENSEMBL:ENSMUSG00000105957 | MGI:5663391 | lncRNA gene | predicted gene 43254 |
| 3 | gene | 133.3101 | 133.3782 | positive | MGI_C57BL6J_1922026 | Ppa2 | NCBI_Gene:74776,ENSEMBL:ENSMUSG00000028013 | MGI:1922026 | protein coding gene | pyrophosphatase (inorganic) 2 |
| 3 | gene | 133.4317 | 133.4318 | negative | MGI_C57BL6J_5454819 | Gm25042 | ENSEMBL:ENSMUSG00000080506 | MGI:5454819 | miRNA gene | predicted gene, 25042 |
| 3 | gene | 133.4637 | 133.5452 | negative | MGI_C57BL6J_2443298 | Tet2 | NCBI_Gene:214133,ENSEMBL:ENSMUSG00000040943 | MGI:2443298 | protein coding gene | tet methylcytosine dioxygenase 2 |
| 3 | gene | 133.5451 | 133.5851 | positive | MGI_C57BL6J_5662704 | Gm42567 | ENSEMBL:ENSMUSG00000105085 | MGI:5662704 | lncRNA gene | predicted gene 42567 |
| 3 | pseudogene | 133.5790 | 133.5911 | negative | MGI_C57BL6J_6118967 | Gm49316 | NCBI_Gene:102632793,ENSEMBL:ENSMUSG00000105595 | MGI:6118967 | pseudogene | predicted gene 49316 |
| 3 | gene | 133.5884 | 133.5885 | negative | MGI_C57BL6J_5452788 | Gm23011 | ENSEMBL:ENSMUSG00000077336 | MGI:5452788 | snoRNA gene | predicted gene, 23011 |
| 3 | gene | 133.6237 | 133.6348 | negative | MGI_C57BL6J_5610456 | Gm37228 | ENSEMBL:ENSMUSG00000102530 | MGI:5610456 | unclassified gene | predicted gene, 37228 |
| 3 | gene | 133.6457 | 133.6638 | negative | MGI_C57BL6J_5623038 | Gm40153 | NCBI_Gene:105244560,ENSEMBL:ENSMUSG00000105952 | MGI:5623038 | lncRNA gene | predicted gene, 40153 |
| 3 | gene | 133.7892 | 133.7915 | negative | MGI_C57BL6J_3643755 | Gm6135 | NCBI_Gene:620205,ENSEMBL:ENSMUSG00000106069 | MGI:3643755 | lncRNA gene | prediticted gene 6135 |
| 3 | gene | 133.8159 | 133.8186 | negative | MGI_C57BL6J_5589643 | Gm30484 | NCBI_Gene:102632401,ENSEMBL:ENSMUSG00000104645 | MGI:5589643 | lncRNA gene | predicted gene, 30484 |
| 3 | gene | 133.8306 | 134.2406 | negative | MGI_C57BL6J_5477185 | Gm26691 | NCBI_Gene:102632484,ENSEMBL:ENSMUSG00000097911 | MGI:5477185 | lncRNA gene | predicted gene, 26691 |
| 3 | gene | 134.1470 | 134.1471 | negative | MGI_C57BL6J_5690753 | Gm44361 | ENSEMBL:ENSMUSG00000106130 | MGI:5690753 | miRNA gene | predicted gene, 44361 |
| 3 | gene | 134.1604 | 134.1920 | positive | MGI_C57BL6J_5663329 | Gm43192 | ENSEMBL:ENSMUSG00000104556 | MGI:5663329 | lncRNA gene | predicted gene 43192 |
| 3 | pseudogene | 134.1809 | 134.1814 | positive | MGI_C57BL6J_5663330 | Gm43193 | ENSEMBL:ENSMUSG00000105523 | MGI:5663330 | pseudogene | predicted gene 43193 |
| 3 | gene | 134.1999 | 134.2132 | positive | MGI_C57BL6J_5589807 | Gm30648 | NCBI_Gene:102632621,ENSEMBL:ENSMUSG00000106139 | MGI:5589807 | lncRNA gene | predicted gene, 30648 |
| 3 | gene | 134.2364 | 134.2622 | positive | MGI_C57BL6J_2442112 | Cxxc4 | NCBI_Gene:319478,ENSEMBL:ENSMUSG00000044365 | MGI:2442112 | protein coding gene | CXXC finger 4 |
| 3 | gene | 134.2405 | 134.2406 | negative | MGI_C57BL6J_3811410 | Mir1895 | miRBase:MI0008316,NCBI_Gene:100316832,ENSEMBL:ENSMUSG00000084564 | MGI:3811410 | miRNA gene | microRNA 1895 |
| 3 | pseudogene | 134.2720 | 134.2725 | negative | MGI_C57BL6J_5663696 | Gm43559 | ENSEMBL:ENSMUSG00000104779 | MGI:5663696 | pseudogene | predicted gene 43559 |
| 3 | gene | 134.3573 | 134.3575 | positive | MGI_C57BL6J_5663695 | Gm43558 | ENSEMBL:ENSMUSG00000106628 | MGI:5663695 | unclassified gene | predicted gene 43558 |
| 3 | pseudogene | 134.3654 | 134.3661 | negative | MGI_C57BL6J_3648504 | Gm6077 | NCBI_Gene:606718,ENSEMBL:ENSMUSG00000104530 | MGI:3648504 | pseudogene | predicted gene 6077 |
| 3 | gene | 134.4041 | 134.4236 | negative | MGI_C57BL6J_1922458 | 4930539C22Rik | NCBI_Gene:75208,ENSEMBL:ENSMUSG00000105765 | MGI:1922458 | lncRNA gene | RIKEN cDNA 4930539C22 gene |
| 3 | gene | 134.4621 | 134.6308 | negative | MGI_C57BL6J_5477314 | Gm26820 | ENSEMBL:ENSMUSG00000097031 | MGI:5477314 | lncRNA gene | predicted gene, 26820 |
| 3 | gene | 134.8290 | 134.9346 | positive | MGI_C57BL6J_892968 | Tacr3 | NCBI_Gene:21338,ENSEMBL:ENSMUSG00000028172 | MGI:892968 | protein coding gene | tachykinin receptor 3 |
| 3 | pseudogene | 134.9630 | 134.9654 | positive | MGI_C57BL6J_3647155 | Gm4860 | NCBI_Gene:229837,ENSEMBL:ENSMUSG00000105559 | MGI:3647155 | pseudogene | predicted gene 4860 |
| 3 | pseudogene | 134.9659 | 134.9672 | positive | MGI_C57BL6J_5663173 | Gm43036 | ENSEMBL:ENSMUSG00000106569 | MGI:5663173 | pseudogene | predicted gene 43036 |
| 3 | pseudogene | 134.9715 | 134.9720 | positive | MGI_C57BL6J_5663172 | Gm43035 | ENSEMBL:ENSMUSG00000104539 | MGI:5663172 | pseudogene | predicted gene 43035 |
| 3 | pseudogene | 135.1219 | 135.1224 | positive | MGI_C57BL6J_3780311 | Gm2142 | NCBI_Gene:100039290,ENSEMBL:ENSMUSG00000105538 | MGI:3780311 | pseudogene | predicted gene 2142 |
| 3 | gene | 135.1474 | 135.1475 | positive | MGI_C57BL6J_5453825 | Gm24048 | ENSEMBL:ENSMUSG00000076366 | MGI:5453825 | miRNA gene | predicted gene, 24048 |
| 3 | pseudogene | 135.1706 | 135.1713 | positive | MGI_C57BL6J_5010731 | Gm18546 | NCBI_Gene:100417343,ENSEMBL:ENSMUSG00000105266 | MGI:5010731 | pseudogene | predicted gene, 18546 |
| 3 | pseudogene | 135.2090 | 135.2094 | positive | MGI_C57BL6J_5663170 | Gm43033 | ENSEMBL:ENSMUSG00000106079 | MGI:5663170 | pseudogene | predicted gene 43033 |
| 3 | gene | 135.2125 | 135.2736 | positive | MGI_C57BL6J_1098230 | Cenpe | NCBI_Gene:229841,ENSEMBL:ENSMUSG00000045328 | MGI:1098230 | protein coding gene | centromere protein E |
| 3 | gene | 135.2812 | 135.3044 | positive | MGI_C57BL6J_1917022 | Bdh2 | NCBI_Gene:69772,ENSEMBL:ENSMUSG00000028167 | MGI:1917022 | protein coding gene | 3-hydroxybutyrate dehydrogenase, type 2 |
| 3 | gene | 135.2820 | 135.3163 | negative | MGI_C57BL6J_5826463 | Gm46826 | NCBI_Gene:108168902 | MGI:5826463 | lncRNA gene | predicted gene, 46826 |
| 3 | gene | 135.3077 | 135.3454 | positive | MGI_C57BL6J_2140077 | Slc9b2 | NCBI_Gene:97086,ENSEMBL:ENSMUSG00000037994 | MGI:2140077 | protein coding gene | solute carrier family 9, subfamily B (NHA2, cation proton antiporter 2), member 2 |
| 3 | gene | 135.3480 | 135.3978 | positive | MGI_C57BL6J_1921696 | Slc9b1 | NCBI_Gene:74446,ENSEMBL:ENSMUSG00000050150 | MGI:1921696 | protein coding gene | solute carrier family 9, subfamily B (NHA1, cation proton antiporter 1), member 1 |
| 3 | gene | 135.4064 | 135.4241 | negative | MGI_C57BL6J_1914256 | Cisd2 | NCBI_Gene:67006,ENSEMBL:ENSMUSG00000028165 | MGI:1914256 | protein coding gene | CDGSH iron sulfur domain 2 |
| 3 | gene | 135.4252 | 135.4279 | negative | MGI_C57BL6J_1914308 | 2810428J06Rik | ENSEMBL:ENSMUSG00000105981 | MGI:1914308 | unclassified gene | RIKEN cDNA 2810428J06 gene |
| 3 | gene | 135.4282 | 135.4359 | negative | MGI_C57BL6J_5663516 | Gm43379 | ENSEMBL:ENSMUSG00000105449 | MGI:5663516 | unclassified gene | predicted gene 43379 |
| 3 | gene | 135.4362 | 135.4387 | negative | MGI_C57BL6J_1922477 | 4930539J05Rik | NCBI_Gene:319587,ENSEMBL:ENSMUSG00000097032 | MGI:1922477 | lncRNA gene | RIKEN cDNA 4930539J05 gene |
| 3 | gene | 135.4381 | 135.4682 | positive | MGI_C57BL6J_1913355 | Ube2d3 | NCBI_Gene:66105,ENSEMBL:ENSMUSG00000078578 | MGI:1913355 | protein coding gene | ubiquitin-conjugating enzyme E2D 3 |
| 3 | gene | 135.4717 | 135.4855 | negative | MGI_C57BL6J_5590151 | Gm30992 | NCBI_Gene:102633078 | MGI:5590151 | lncRNA gene | predicted gene, 30992 |
| 3 | pseudogene | 135.4836 | 135.4840 | positive | MGI_C57BL6J_5662944 | Gm42807 | ENSEMBL:ENSMUSG00000105404 | MGI:5662944 | pseudogene | predicted gene 42807 |
| 3 | gene | 135.4856 | 135.5714 | positive | MGI_C57BL6J_88175 | Manba | NCBI_Gene:110173,ENSEMBL:ENSMUSG00000028164 | MGI:88175 | protein coding gene | mannosidase, beta A, lysosomal |
| 3 | pseudogene | 135.4957 | 135.4959 | positive | MGI_C57BL6J_3802060 | Gm16100 | ENSEMBL:ENSMUSG00000083045 | MGI:3802060 | pseudogene | predicted gene 16100 |
| 3 | pseudogene | 135.5250 | 135.5268 | negative | MGI_C57BL6J_3644634 | Oaz2-ps | NCBI_Gene:102633210,ENSEMBL:ENSMUSG00000083610 | MGI:3644634 | pseudogene | ornithine decarboxylase antizyme 2, pseudogene |
| 3 | gene | 135.5847 | 135.6920 | negative | MGI_C57BL6J_97312 | Nfkb1 | NCBI_Gene:18033,ENSEMBL:ENSMUSG00000028163 | MGI:97312 | protein coding gene | nuclear factor of kappa light polypeptide gene enhancer in B cells 1, p105 |
| 3 | gene | 135.6116 | 135.6128 | positive | MGI_C57BL6J_5826464 | Gm46827 | NCBI_Gene:108168903 | MGI:5826464 | lncRNA gene | predicted gene, 46827 |
| 3 | gene | 135.6507 | 135.6546 | negative | MGI_C57BL6J_5663567 | Gm43430 | ENSEMBL:ENSMUSG00000106149 | MGI:5663567 | unclassified gene | predicted gene 43430 |
| 3 | gene | 135.7273 | 135.7287 | positive | MGI_C57BL6J_3642121 | Gm9799 | NA | NA | protein coding gene | predicted gene 9799 |
| 3 | gene | 135.8161 | 135.8161 | negative | MGI_C57BL6J_1923004 | 9030607L02Rik | NA | NA | unclassified gene | RIKEN cDNA 9030607L02 gene |
| 3 | gene | 135.8169 | 135.8886 | positive | MGI_C57BL6J_1914797 | Slc39a8 | NCBI_Gene:67547,ENSEMBL:ENSMUSG00000053897 | MGI:1914797 | protein coding gene | solute carrier family 39 (metal ion transporter), member 8 |
| 3 | gene | 135.8337 | 135.8403 | negative | MGI_C57BL6J_1918288 | 4933401H06Rik | NCBI_Gene:71038,ENSEMBL:ENSMUSG00000106306 | MGI:1918288 | lncRNA gene | RIKEN cDNA 4933401H06 gene |
| 3 | gene | 135.8603 | 135.8626 | positive | MGI_C57BL6J_5663566 | Gm43429 | ENSEMBL:ENSMUSG00000106290 | MGI:5663566 | unclassified gene | predicted gene 43429 |
| 3 | gene | 135.8721 | 135.8760 | positive | MGI_C57BL6J_5663565 | Gm43428 | ENSEMBL:ENSMUSG00000105681 | MGI:5663565 | unclassified gene | predicted gene 43428 |
| 3 | gene | 135.9018 | 136.1672 | positive | MGI_C57BL6J_5590402 | Gm31243 | NCBI_Gene:102633410,ENSEMBL:ENSMUSG00000104710 | MGI:5590402 | lncRNA gene | predicted gene, 31243 |
| 3 | pseudogene | 135.9023 | 135.9028 | negative | MGI_C57BL6J_5010515 | Gm18330 | NCBI_Gene:100416947,ENSEMBL:ENSMUSG00000106141 | MGI:5010515 | pseudogene | predicted gene, 18330 |
| 3 | pseudogene | 135.9632 | 135.9728 | negative | MGI_C57BL6J_3704226 | Gm16500 | NCBI_Gene:100039350 | MGI:3704226 | pseudogene | predicted gene 16500 |
| 3 | pseudogene | 135.9635 | 135.9638 | negative | MGI_C57BL6J_5663739 | Gm43602 | ENSEMBL:ENSMUSG00000105937 | MGI:5663739 | pseudogene | predicted gene 43602 |
| 3 | pseudogene | 136.0492 | 136.0517 | negative | MGI_C57BL6J_3779483 | Gm5281 | NCBI_Gene:383923,ENSEMBL:ENSMUSG00000105244 | MGI:3779483 | pseudogene | predicted gene 5281 |
| 3 | gene | 136.0534 | 136.3261 | negative | MGI_C57BL6J_2442120 | Bank1 | NCBI_Gene:242248,ENSEMBL:ENSMUSG00000037922 | MGI:2442120 | protein coding gene | B cell scaffold protein with ankyrin repeats 1 |
| 3 | pseudogene | 136.3315 | 136.3330 | positive | MGI_C57BL6J_3645775 | Gm9408 | NCBI_Gene:668868,ENSEMBL:ENSMUSG00000105869 | MGI:3645775 | pseudogene | predicted gene 9408 |
| 3 | pseudogene | 136.3462 | 136.3464 | positive | MGI_C57BL6J_5663539 | Gm43402 | ENSEMBL:ENSMUSG00000106363 | MGI:5663539 | pseudogene | predicted gene 43402 |
| 3 | gene | 136.4353 | 136.4493 | negative | MGI_C57BL6J_1917305 | 1700030L20Rik | NCBI_Gene:70055,ENSEMBL:ENSMUSG00000099508 | MGI:1917305 | lncRNA gene | RIKEN cDNA 1700030L20 gene |
| 3 | gene | 136.6701 | 136.9377 | positive | MGI_C57BL6J_107164 | Ppp3ca | NCBI_Gene:19055,ENSEMBL:ENSMUSG00000028161 | MGI:107164 | protein coding gene | protein phosphatase 3, catalytic subunit, alpha isoform |
| 3 | gene | 136.7012 | 136.7027 | positive | MGI_C57BL6J_5662959 | Gm42822 | ENSEMBL:ENSMUSG00000106596 | MGI:5662959 | unclassified gene | predicted gene 42822 |
| 3 | gene | 136.7590 | 136.7600 | negative | MGI_C57BL6J_1922633 | 4930599N24Rik | ENSEMBL:ENSMUSG00000105545 | MGI:1922633 | unclassified gene | RIKEN cDNA 4930599N24 gene |
| 3 | gene | 136.7983 | 136.7984 | negative | MGI_C57BL6J_5690719 | Gm44327 | ENSEMBL:ENSMUSG00000106439 | MGI:5690719 | miRNA gene | predicted gene, 44327 |
Additive covariates
Significance thresholds
operm <- scan1perm(pr.qc, covars["rz.age"], model="normal", addcovar=addcovar, n_perm=10, perm_Xsp=TRUE, chr_lengths=chr_lengths(gm$gmap))
summary_table<-data.frame(unclass(summary(operm, alpha=c(0.01, 0.05, 0.1))))
names(summary_table) <- c("autosomes","X")
summary_table$significance.level <- rownames(summary_table)
rownames(summary_table) <- NULL
summary_table[c(3,1:2)] %>%
kable(escape = F,align = c("ccc")) %>%
kable_styling("striped", full_width = T) %>%
column_spec(1, bold=TRUE)| significance.level | autosomes | X |
|---|---|---|
| 0.01 | 6.215670 | 6.980791 |
| 0.05 | 5.415470 | 6.832073 |
| 0.1 | 4.412724 | 6.637599 |
Genome-wide scan
plot_lod<-function(out,map){
for (i in 1:dim(out)[2]){
#par(mar=c(5.1, 6.1, 1.1, 1.1))
ymx <- maxlod(out) # overall maximum LOD score
plot(out, map, lodcolumn=i, col="slateblue", ylim=c(0, ymx+0.5))
#legend("topright", lwd=2, colnames(out)[i], bg="gray90")
title(main = paste0(colnames(out)[i], " - ICI vs. EOI (with additive covariates) [positions in cM]"))
add_threshold(map, summary(operm,alpha=0.1), col = 'purple')
add_threshold(map, summary(operm, alpha=0.05), col = 'red')
add_threshold(map, summary(operm, alpha=0.01), col = 'blue')
}
}
print("with no kinship")[1] “with no kinship”
outfa <- scan1(pr.qc, covars["rz.age"], Xcovar=Xcovar, model="normal", addcovar=addcovar)
plot_lod(outfa,gm$gmap)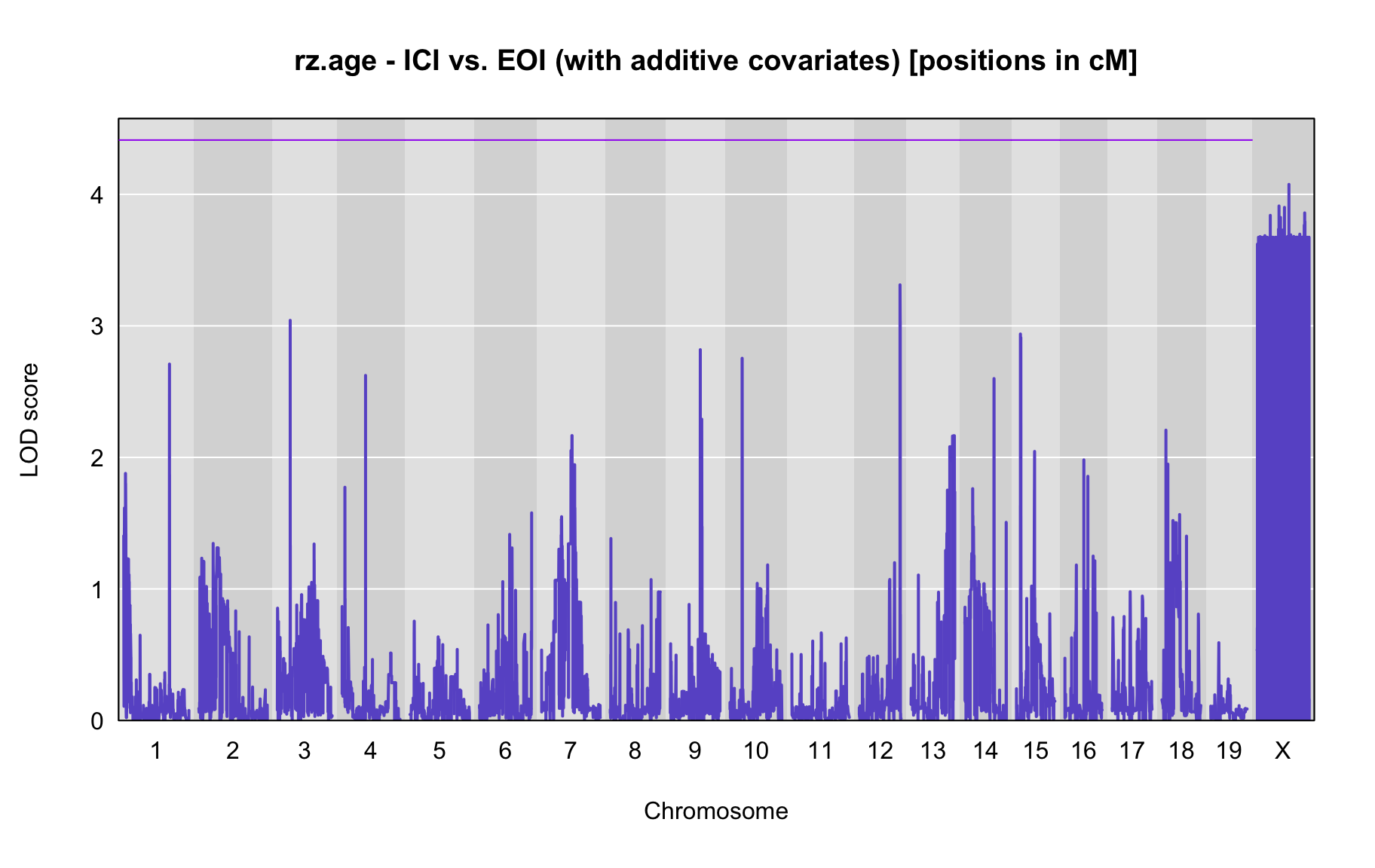
print("with normal kinship")[1] “with normal kinship”
outka <- scan1(pr.qc, covars["rz.age"], Xcovar=Xcovar, model="normal", addcovar=addcovar, kinship=kinship)
plot_lod(outka,gm$gmap)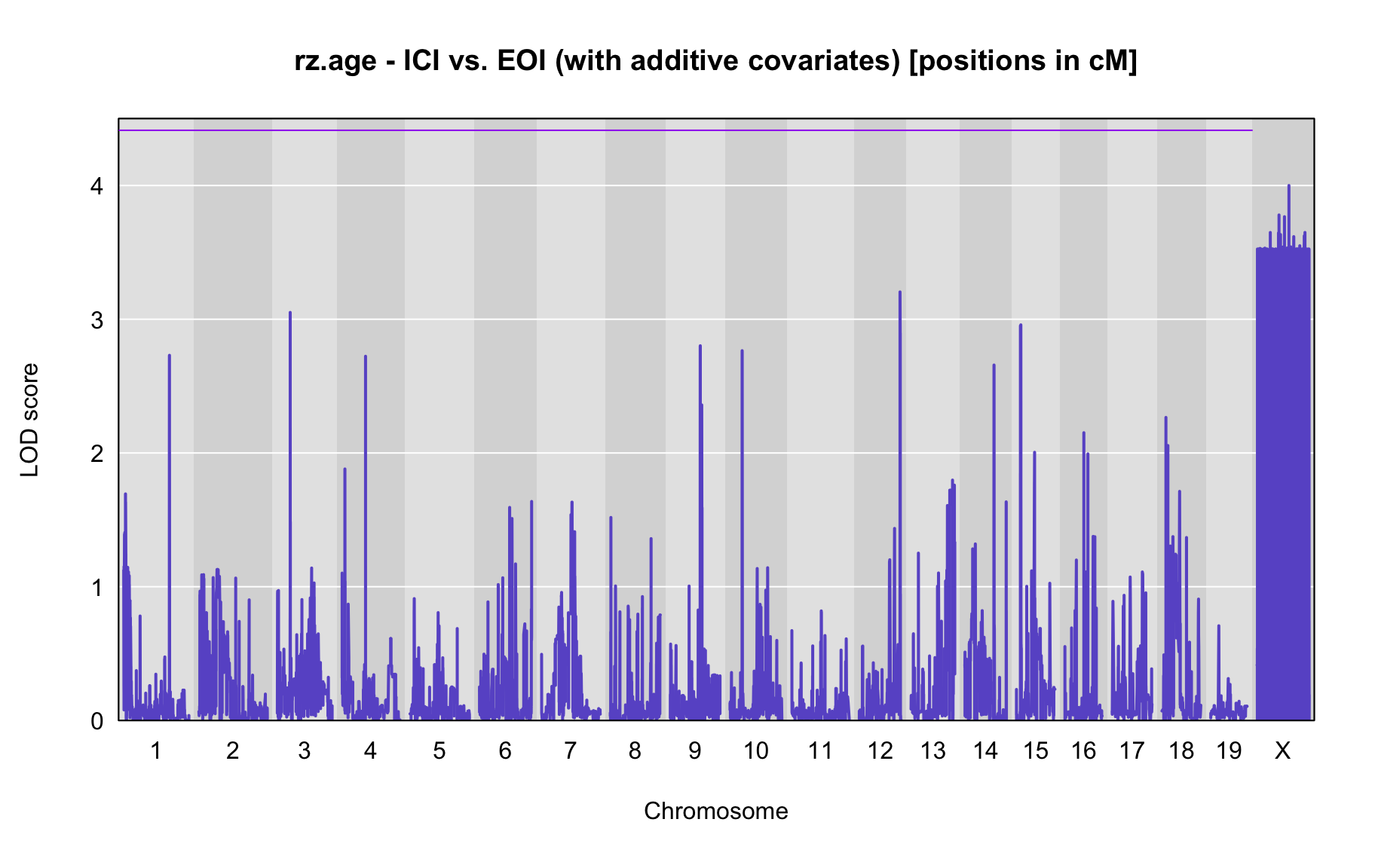
print("with loco kinship")[1] “with loco kinship”
outa <- scan1(pr.qc, covars["rz.age"], Xcovar=Xcovar, model="normal", addcovar=addcovar, kinship=K)
plot_lod(outa,gm$gmap)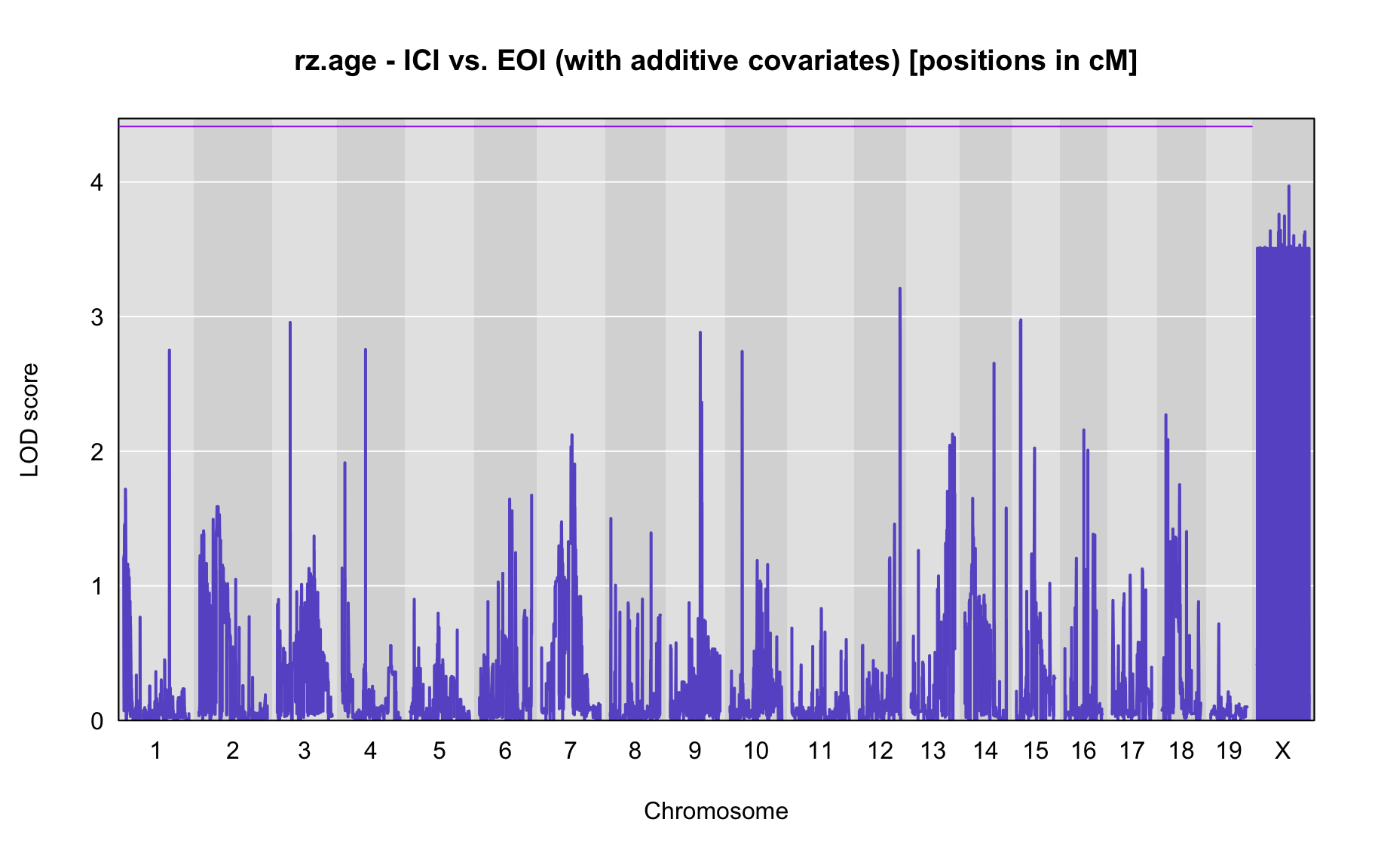
out = outa
print(paste0("number of samples in analysis = ", as.data.frame(attributes(out)$sample_size)[1,]))[1] “number of samples in analysis = 267”
LOD peaks
Centimorgan (cM)
peaks<-find_peaks(out, gm$gmap, threshold=summary(operm,alpha=0.05)$A, thresholdX = summary(operm,alpha=0.05)$X, peakdrop=3, drop=1.5)
if(nrow(peaks) > 0){
peaks$marker <- find_marker(gm$gmap, chr=peaks$chr,pos=peaks$pos)
names(peaks)[2] <- c("phenotype")
peaks <- peaks[-1]
rownames(peaks) <- NULL
print(kable(peaks, escape = F, align = c("cccccccc"), "html")
%>% kable_styling("striped", full_width = T)%>%
column_spec(1, bold=TRUE)
)
#plot only peak chromosomes
plot_lod_chr<-function(out,map,chrom){
for (i in 1:dim(out)[2]){
#png(filename=paste0("/Users/chenm/Documents/qtl/Jai/",colnames(out)[i], "_lod.png"))
#par(mar=c(5.1, 6.1, 1.1, 1.1))
ymx <- maxlod(out) # overall maximum LOD score
plot(out, map, chr = chrom, lodcolumn=i, col="slateblue", ylim=c(0, ymx+0.5))
#legend("topright", lwd=2, colnames(out)[i], bg="gray90")
title(main = paste0(colnames(out)[i], " - chr", chrom, " - ICI vs. EOI (with additive covariates) [positions in cM]"))
add_threshold(map, summary(operm,alpha=0.1), col = 'purple')
add_threshold(map, summary(operm, alpha=0.05), col = 'red')
add_threshold(map, summary(operm, alpha=0.01), col = 'blue')
#for (j in 1: dim(summary_table)[1]){
# abline(h=summary_table[j, i],col="red")
# text(x=400, y =summary_table[j, i]+0.12, labels = paste("p=", row.names(summary_table)[j]))
#}
#dev.off()
}
}
} else {
print(paste0("There are no peaks that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}[1] “There are no peaks that have a LOD that reaches suggestive (p<0.05) level of 5.41546972581755 [autosomes]/6.83207325744769 [x-chromosome]”
if(nrow(peaks) < 50 & nrow(peaks) > 0){
for(i in unique(peaks$chr)){
#for (i in 1:nrow(peaks)){
#plot_lod_chr(out,gm$gmap, peaks$chr[i])
plot_lod_chr(out,gm$gmap, i)
}
} else {
print(paste0("There are too many peaks (",nrow(peaks)," peaks) that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}[1] “There are too many peaks (0 peaks) that have a LOD that reaches suggestive (p<0.05) level of 5.41546972581755 [autosomes]/6.83207325744769 [x-chromosome]”
Megabase (MB)
peaks_mba<-find_peaks(out, gm$pmap, threshold=summary(operm,alpha=0.05)$A, thresholdX = summary(operm,alpha=0.05)$X, peakdrop=3, drop=1.5)
if(nrow(peaks) > 0){
peaks_mba$marker <- find_marker(gm$pmap, chr=peaks_mba$chr,pos=peaks_mba$pos)
names(peaks_mba)[2] <- c("phenotype")
peaks_mba <- peaks_mba[-1]
rownames(peaks_mba) <- NULL
print(kable(peaks_mba, escape = F, align = c("cccccccc"), "html")
%>% kable_styling("striped", full_width = T)%>%
column_spec(1, bold=TRUE)
)
plot_lod_chr_mb<-function(out,map,chrom){
for (i in 1:dim(out)[2]){
#png(filename=paste0("/Users/chenm/Documents/qtl/Jai/",colnames(out)[i], "_lod.png"))
#par(mar=c(5.1, 6.1, 1.1, 1.1))
ymx <- maxlod(out) # overall maximum LOD score
plot(out, map, chr = chrom, lodcolumn=i, col="slateblue", ylim=c(0, ymx+0.5))
#legend("topright", lwd=2, colnames(out)[i], bg="gray90")
title(main = paste0(colnames(out)[i], " - chr", chrom, " - ICI vs. EOI (with additive covariates) [positions in cM]"))
add_threshold(map, summary(operm,alpha=0.1), col = 'purple')
add_threshold(map, summary(operm, alpha=0.05), col = 'red')
add_threshold(map, summary(operm, alpha=0.01), col = 'blue')
#for (j in 1: dim(summary_table)[1]){
# abline(h=summary_table[j, i],col="red")
# text(x=400, y =summary_table[j, i]+0.12, labels = paste("p=", row.names(summary_table)[j]))
#}
#dev.off()
}
}
} else {
print(paste0("There are no peaks that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}[1] “There are no peaks that have a LOD that reaches suggestive (p<0.05) level of 5.41546972581755 [autosomes]/6.83207325744769 [x-chromosome]”
if(nrow(peaks_mba) < 50 & nrow(peaks_mba) > 0){
for(i in unique(peaks_mba$chr)){
#for (i in 1:nrow(peaks_mba)){
#plot_lod_chr_mb(out,gm$pmap, peaks_mba$chr[i])
plot_lod_chr_mb(out,gm$pmap,i)
}
} else {
print(paste0("There are too many peaks (",nrow(peaks_mba)," peaks) that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}[1] “There are too many peaks (0 peaks) that have a LOD that reaches suggestive (p<0.05) level of 5.41546972581755 [autosomes]/6.83207325744769 [x-chromosome]”
QTL effects
For each peak LOD location we give a list of gene
if(nrow(peaks) < 50 & nrow(peaks) > 0){
for (i in 1:nrow(peaks)){
#for (i in 1:1){
#Plot 1
g <- maxmarg(pr.qc, gm$gmap, chr=peaks$chr[i], pos=peaks$pos[i], return_char=TRUE)
#png(filename=paste0("/Users/chenm/Documents/qtl/Jai/","qtl_effect_", i, ".png"))
#par(mar=c(4.1, 4.1, 1.5, 0.6))
plot_pxg(g, covars[,peaks$phenotype[i]], ylab=peaks$phenotype[i], sort=FALSE)
title(main = paste0("chr: ", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$phenotype[i]," )"), line=0.2)
##dev.off()
chr = peaks$chr[i]
# Plot 2
pr_sub <- pull_genoprobint(pr.qc, gm$gmap, chr, c(peaks$ci_lo[i], peaks$ci_hi[i]))
#coeff <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]], addcovar = addcovar)
#coeff <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]], Xcovar=Xcovar)
#coeff <- scan1coef(pr.qc[,chr], gm$covar[peaks$lodcolumn[i]], model="binary")
#coeff_sub <- scan1coef(pr_sub[,chr], gm$covar[peaks$lodcolumn[i]], model="binary")
blup <- scan1blup(pr.qc[,chr], covars[peaks$phenotype[i]])
blup_sub <- scan1blup(pr_sub[,chr], covars[peaks$phenotype[i]])
write.csv(as.data.frame(blup_sub), paste0("data/ici.vs.eoi_",peaks$phenotype[i],"-additive.covariates_blup_sub_chr-",chr,"_peak.marker-",peaks$marker[i],"_lod.drop-1.5_5.batches_52.csv"), quote=F)
#plot_coef(coeff,
# gm$gmap, columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$lodcolumn[i]," [scan1coeff; positions in cM] )")
# )
#plot_coef(coeff_sub,
# gm$gmap, columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$lodcolumn[i],"; 1.5 LOD drop interval [scan1coeff; positions in cM] ) ")
# )
plot_coef(blup,
gm$gmap, columns=1:2,
bgcolor="gray95", legend="bottomleft",
main = paste0("chr: ", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$phenotype[i]," [scan1blup; positions in cM] )")
)
plot_coef(blup_sub,
gm$gmap, columns=1:2,
bgcolor="gray95", legend="bottomleft",
main = paste0("chr: ", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$phenotype[i],"; 1.5 LOD drop interval [scan1blup; positions in cM] )")
)
# Plot 3
#c2effB <- scan1coef(pr.qc[,chr], gm$covar[peaks$lodcolumn[i]], model="binary", contrasts=cbind(a=c(-1, 0), d=c(0, -1)))
#c2effBb <- scan1blup(pr.qc[,chr], gm$covar[peaks$lodcolumn[i]], contrasts=cbind(a=c(-1, 0), d=c(0, -1)))
##c2effB <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]], addcovar = addcovar, contrasts=cbind(mu=c(1,1,1), a=c(-1, 0, 1), d=c(0, 1, 0)))
##c2effB <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]],Xcovar=Xcovar, contrasts=cbind(mu=c(1,1,1), a=c(-1, 0, 1), d=c(0, 1, 0)))
#plot(c2effB, gm$gmap[chr], columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i], "pos", peaks$pos[i], "(",peaks$lodcolumn[i],")")
# )
#plot(c2effBb, gm$gmap[chr], columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i], "pos", peaks$pos[i], "(",peaks$lodcolumn[i],")")
# )
##last_coef <- unclass(c2effB)[nrow(c2effB),2:3] # last two coefficients
##for(t in seq(along=last_coef))
## axis(side=4, at=last_coef[t], names(last_coef)[t], tick=FALSE)
#Table 1
chr = peaks_mba$chr[i]
start=as.numeric(peaks_mba$ci_lo[i])
end=as.numeric(peaks_mba$ci_hi[i])
genesgss = query_genes(chr, start, end)
write.csv(genesgss, file=paste0("data/ici.vs.eoi_",peaks$phenotype[i],"-additive.covariates_genes_chr-",chr,"_peak.marker-",peaks$marker[i],"_lod.drop-1.5_5.batches_52.csv"), quote=F)
rownames(genesgss) <- NULL
genesgss$strand_old = genesgss$strand
genesgss$strand[genesgss$strand=="+"] <- "positive"
genesgss$strand[genesgss$strand=="-"] <- "negative"
#genesgss <-
#table <-
#genesgss[,c("chr","type","start","stop","strand","ID","Name","Dbxref","gene_id","mgi_type","description")] %>%
#kable(escape = F,align = c("ccccccccccc")) %>%
#kable_styling("striped", full_width = T) #%>%
#cat #%>%
#column_spec(1, bold=TRUE)
#
#print(kable(genesgss[,c("chr","type","start","stop","strand","ID","Name","Dbxref","gene_id","mgi_type","description")], escape = F,align = c("ccccccccccc")))
print(kable(genesgss[,c("chr","type","start","stop","strand","ID","Name","Dbxref","gene_id","mgi_type","description")], "html") %>% kable_styling("striped", full_width = T))
#table
}
} else {
print(paste0("There are too many peaks (",nrow(peaks_mba)," peaks) that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}[1] “There are too many peaks (0 peaks) that have a LOD that reaches suggestive (p<0.05) level of 5.41546972581755 [autosomes]/6.83207325744769 [x-chromosome]”
Interactive term
Significance thresholds
operm <- scan1perm(pr.qc, covars["rz.age"], model="normal", addcovar=addcovar, intcovar=addcovar, n_perm=10, perm_Xsp=TRUE, chr_lengths=chr_lengths(gm$gmap))
summary_table<-data.frame(unclass(summary(operm, alpha=c(0.01, 0.05, 0.1))))
names(summary_table) <- c("autosomes","X")
summary_table$significance.level <- rownames(summary_table)
rownames(summary_table) <- NULL
summary_table[c(3,1:2)] %>%
kable(escape = F,align = c("ccc")) %>%
kable_styling("striped", full_width = T) %>%
column_spec(1, bold=TRUE)| significance.level | autosomes | X |
|---|---|---|
| 0.01 | 8.172135 | 9.315865 |
| 0.05 | 7.675942 | 9.240424 |
| 0.1 | 7.054153 | 9.141771 |
Genome-wide scan
plot_lod<-function(out,map){
for (i in 1:dim(out)[2]){
#par(mar=c(5.1, 6.1, 1.1, 1.1))
ymx <- maxlod(out) # overall maximum LOD score
plot(out, map, lodcolumn=i, col="slateblue", ylim=c(0, ymx+0.5))
#legend("topright", lwd=2, colnames(out)[i], bg="gray90")
title(main = paste0(colnames(out)[i], " - ICI vs. EOI (with interactive covariate) [positions in cM]"))
add_threshold(map, summary(operm,alpha=0.1), col = 'purple')
add_threshold(map, summary(operm, alpha=0.05), col = 'red')
add_threshold(map, summary(operm, alpha=0.01), col = 'blue')
}
}
print("with no kinship")[1] “with no kinship”
outfi <- scan1(pr.qc, covars["rz.age"], Xcovar=Xcovar, model="normal", addcovar=addcovar, intcovar=addcovar)
plot_lod(outfi,gm$gmap)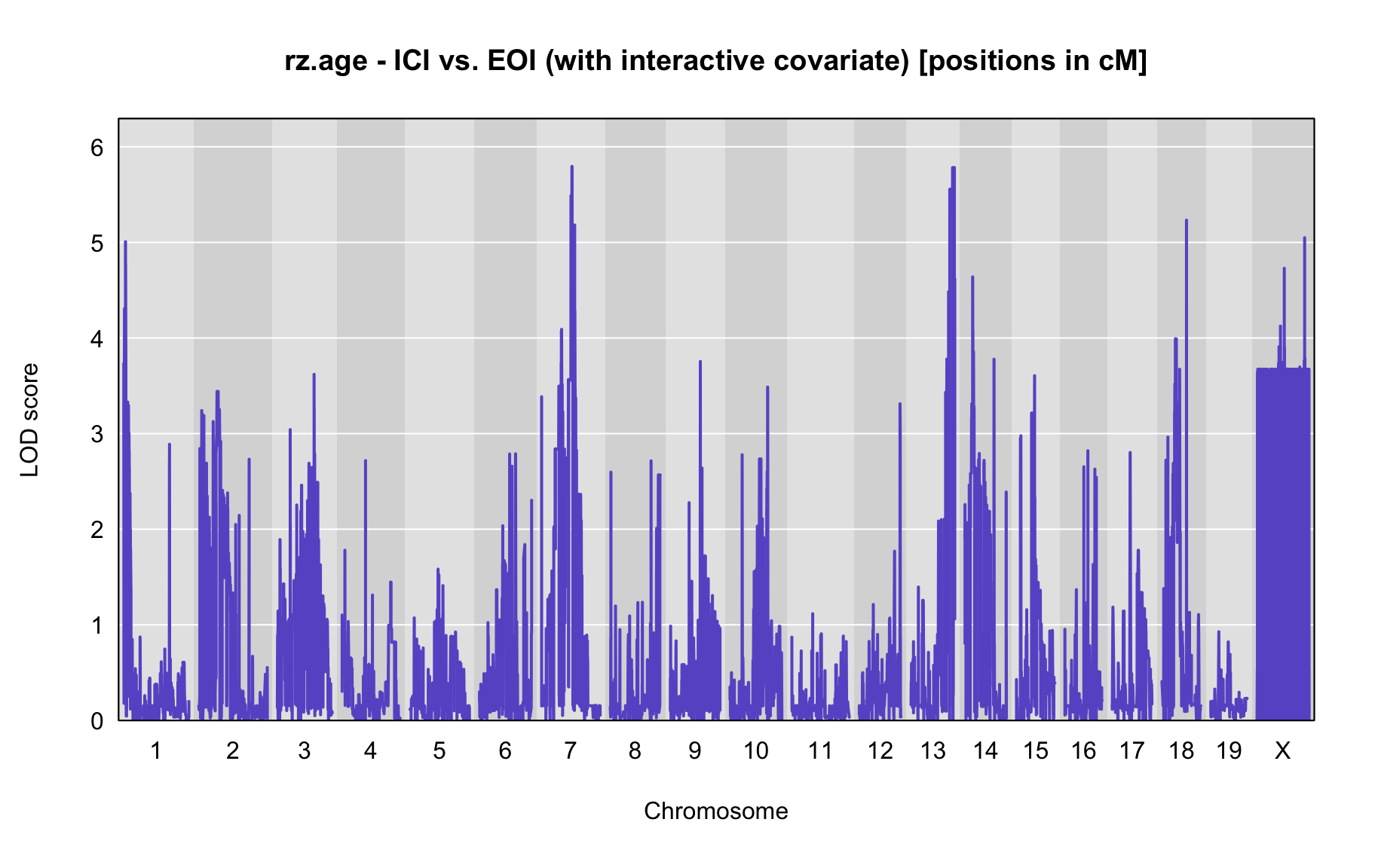
outf.new <- outfi-outfa
plot_lod(outf.new,gm$gmap)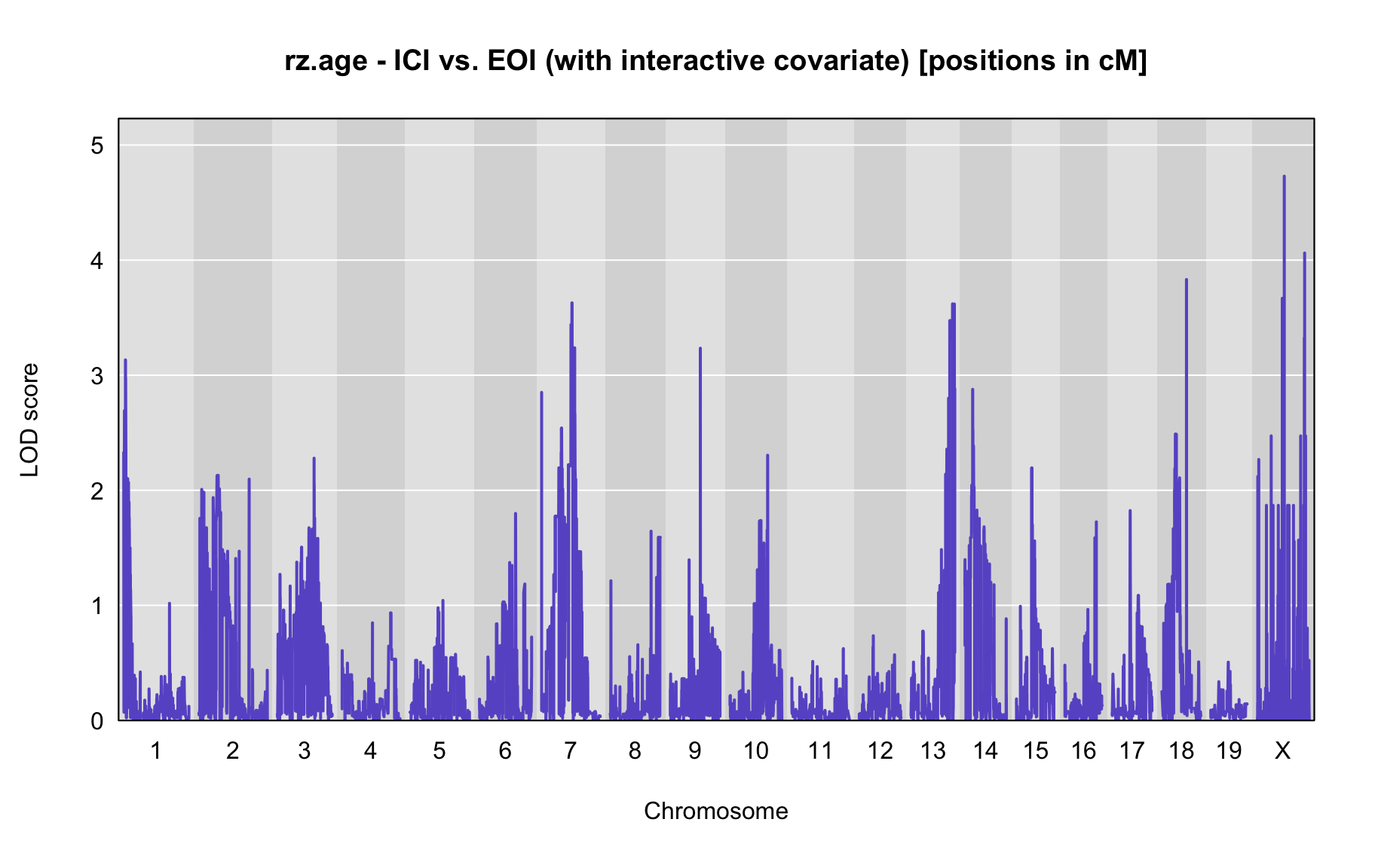
print("with normal kinship")[1] “with normal kinship”
outki <- scan1(pr.qc, covars["rz.age"], Xcovar=Xcovar, model="normal", addcovar=addcovar, intcovar=addcovar, kinship=kinship)
plot_lod(outki,gm$gmap)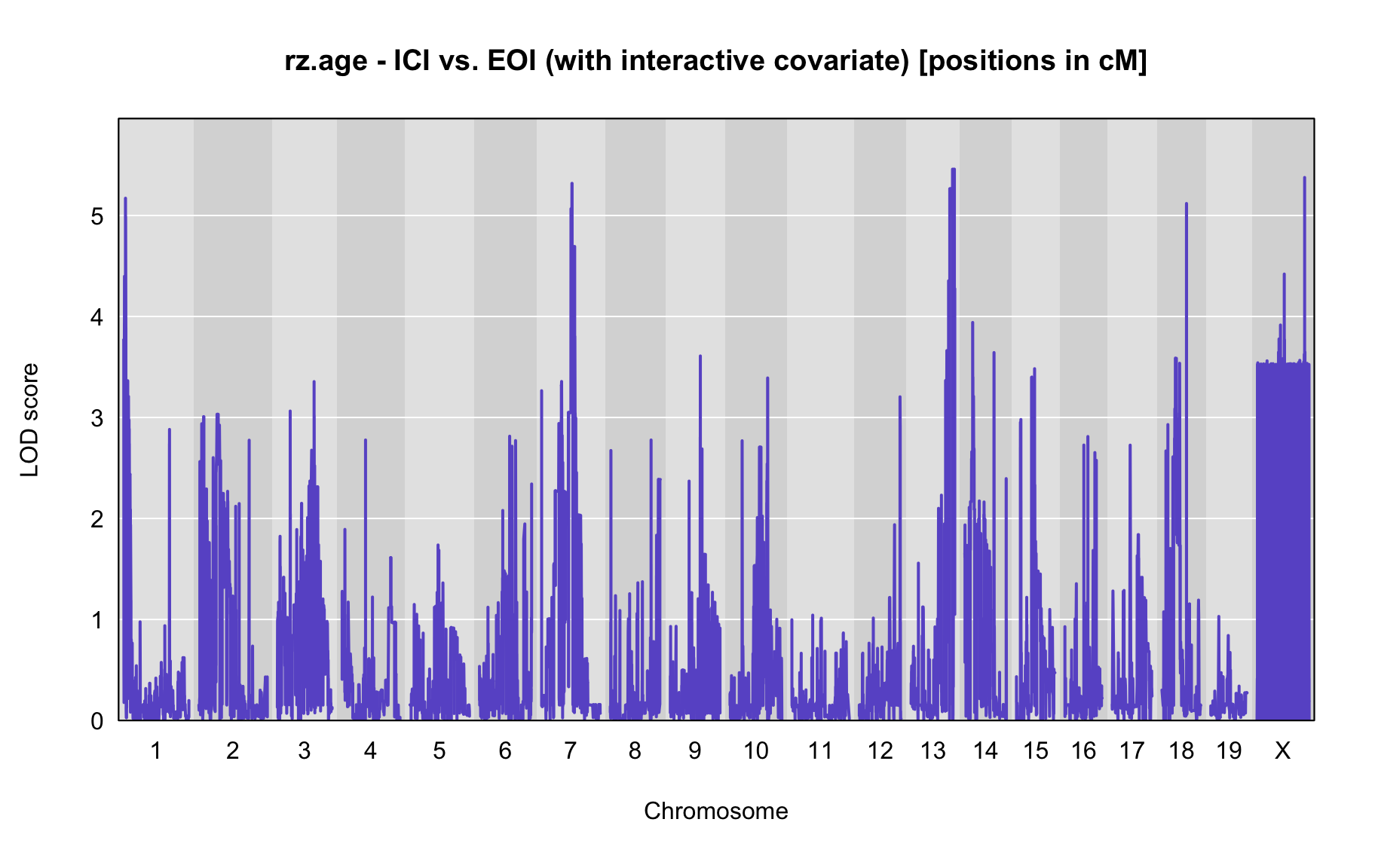
outk.new <- outki-outka
plot_lod(outk.new,gm$gmap)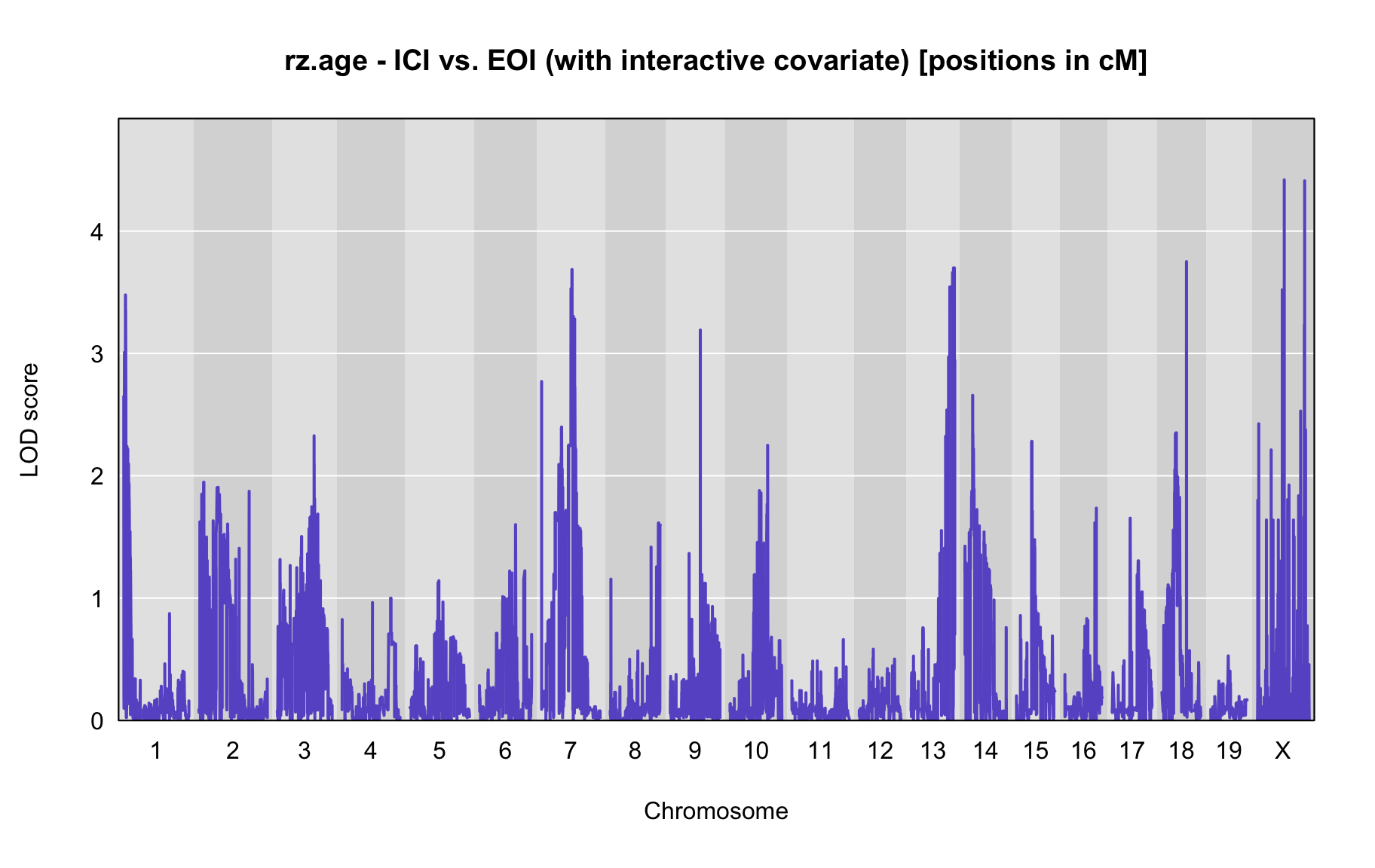
print("with loco kinship")[1] “with loco kinship”
outi <- scan1(pr.qc, covars["rz.age"], Xcovar=Xcovar, model="normal", addcovar=addcovar, intcovar=addcovar, kinship=K)
plot_lod(outi,gm$gmap)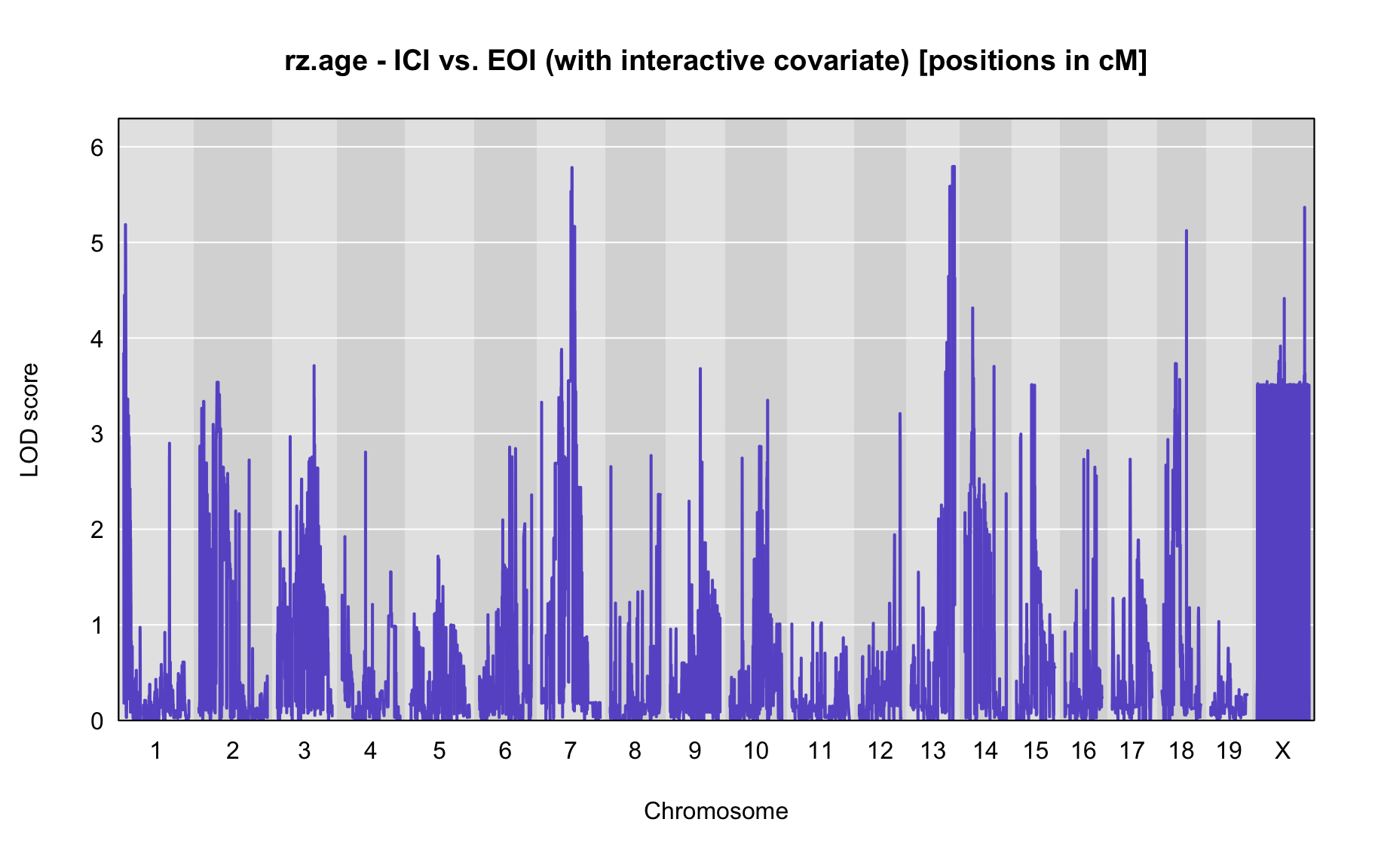
outi.new <- outi-outa
plot_lod(outi.new,gm$gmap)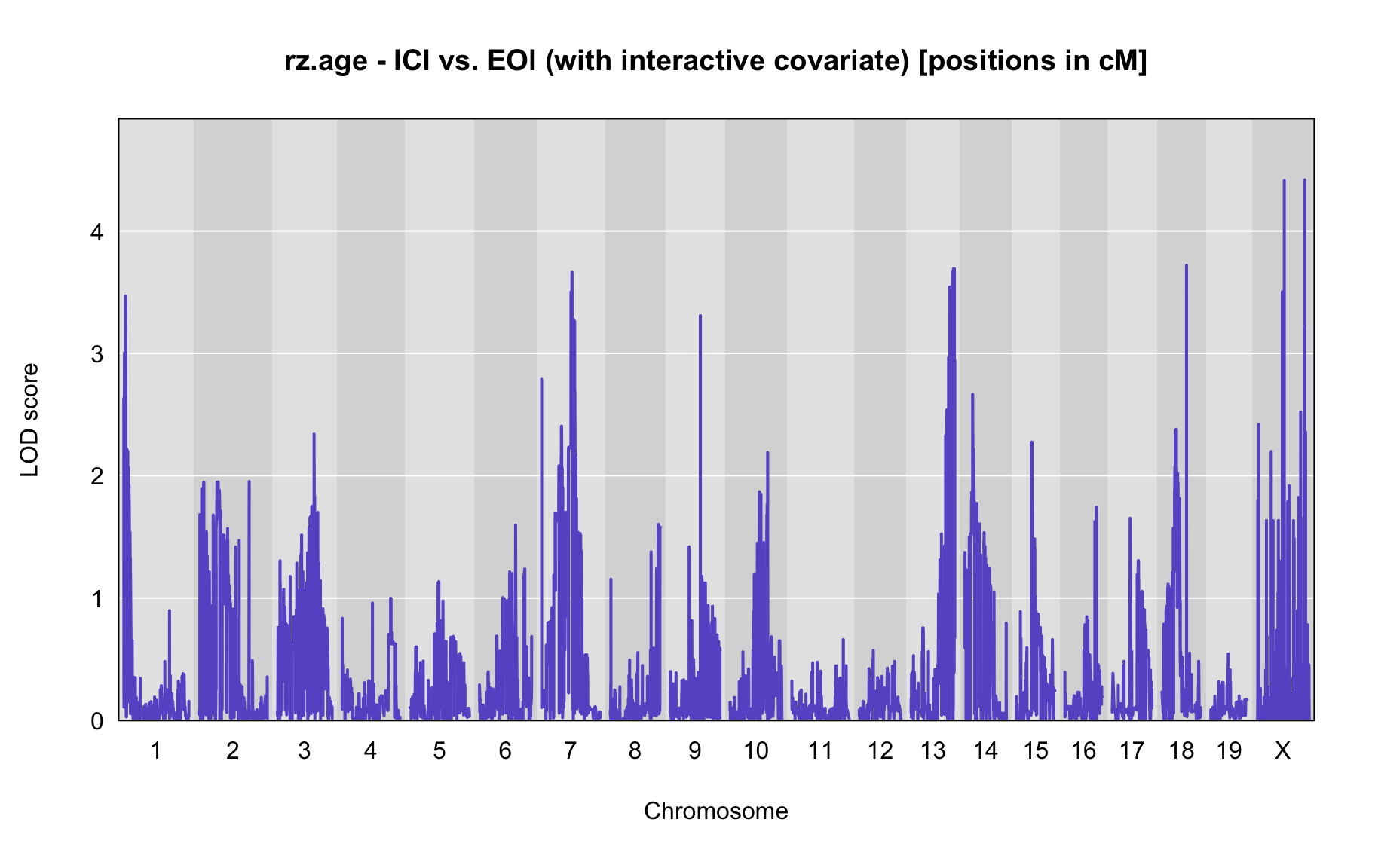
out = outi.new
print(paste0("number of samples in analysis = ", as.data.frame(attributes(out)$sample_size)[1,]))[1] “number of samples in analysis = 267”
LOD peaks
Centimorgan (cM)
peaks<-find_peaks(out, gm$gmap, threshold=summary(operm,alpha=0.05)$A, thresholdX = summary(operm,alpha=0.05)$X, peakdrop=3, drop=1.5)
if(nrow(peaks) > 0){
peaks$marker <- find_marker(gm$gmap, chr=peaks$chr,pos=peaks$pos)
names(peaks)[2] <- c("phenotype")
peaks <- peaks[-1]
rownames(peaks) <- NULL
print(kable(peaks, escape = F, align = c("cccccccc"), "html")
%>% kable_styling("striped", full_width = T)%>%
column_spec(1, bold=TRUE)
)
#plot only peak chromosomes
plot_lod_chr<-function(out,map,chrom){
for (i in 1:dim(out)[2]){
#png(filename=paste0("/Users/chenm/Documents/qtl/Jai/",colnames(out)[i], "_lod.png"))
#par(mar=c(5.1, 6.1, 1.1, 1.1))
ymx <- maxlod(out) # overall maximum LOD score
plot(out, map, chr = chrom, lodcolumn=i, col="slateblue", ylim=c(0, ymx+0.5))
#legend("topright", lwd=2, colnames(out)[i], bg="gray90")
title(main = paste0(colnames(out)[i], " - chr", chrom, " - ICI vs. EOI (with interactive covariate) [positions in cM]"))
add_threshold(map, summary(operm,alpha=0.1), col = 'purple')
add_threshold(map, summary(operm, alpha=0.05), col = 'red')
add_threshold(map, summary(operm, alpha=0.01), col = 'blue')
#for (j in 1: dim(summary_table)[1]){
# abline(h=summary_table[j, i],col="red")
# text(x=400, y =summary_table[j, i]+0.12, labels = paste("p=", row.names(summary_table)[j]))
#}
#dev.off()
}
}
} else {
print(paste0("There are no peaks that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}[1] “There are no peaks that have a LOD that reaches suggestive (p<0.05) level of 7.67594187344621 [autosomes]/9.24042402889532 [x-chromosome]”
if(nrow(peaks) < 50 & nrow(peaks) > 0){
for(i in unique(peaks$chr)){
#for (i in 1:nrow(peaks)){
#plot_lod_chr(out,gm$gmap, peaks$chr[i])
plot_lod_chr(out,gm$gmap, i)
}
} else {
print(paste0("There are too many peaks (",nrow(peaks)," peaks) that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}[1] “There are too many peaks (0 peaks) that have a LOD that reaches suggestive (p<0.05) level of 7.67594187344621 [autosomes]/9.24042402889532 [x-chromosome]”
Megabase (MB)
peaks_mba<-find_peaks(out, gm$pmap, threshold=summary(operm,alpha=0.05)$A, thresholdX = summary(operm,alpha=0.05)$X, peakdrop=3, drop=1.5)
if(nrow(peaks) > 0){
peaks_mba$marker <- find_marker(gm$pmap, chr=peaks_mba$chr,pos=peaks_mba$pos)
names(peaks_mba)[2] <- c("phenotype")
peaks_mba <- peaks_mba[-1]
rownames(peaks_mba) <- NULL
print(kable(peaks_mba, escape = F, align = c("cccccccc"), "html")
%>% kable_styling("striped", full_width = T)%>%
column_spec(1, bold=TRUE)
)
plot_lod_chr_mb<-function(out,map,chrom){
for (i in 1:dim(out)[2]){
#png(filename=paste0("/Users/chenm/Documents/qtl/Jai/",colnames(out)[i], "_lod.png"))
#par(mar=c(5.1, 6.1, 1.1, 1.1))
ymx <- maxlod(out) # overall maximum LOD score
plot(out, map, chr = chrom, lodcolumn=i, col="slateblue", ylim=c(0, ymx+0.5))
#legend("topright", lwd=2, colnames(out)[i], bg="gray90")
title(main = paste0(colnames(out)[i], " - chr", chrom, " - ICI vs. EOI (with interactive covariate) [positions in cM]"))
add_threshold(map, summary(operm,alpha=0.1), col = 'purple')
add_threshold(map, summary(operm, alpha=0.05), col = 'red')
add_threshold(map, summary(operm, alpha=0.01), col = 'blue')
#for (j in 1: dim(summary_table)[1]){
# abline(h=summary_table[j, i],col="red")
# text(x=400, y =summary_table[j, i]+0.12, labels = paste("p=", row.names(summary_table)[j]))
#}
#dev.off()
}
}
} else {
print(paste0("There are no peaks that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}[1] “There are no peaks that have a LOD that reaches suggestive (p<0.05) level of 7.67594187344621 [autosomes]/9.24042402889532 [x-chromosome]”
if(nrow(peaks_mba) < 50 & nrow(peaks_mba) > 0){
for(i in unique(peaks_mba$chr)){
#for (i in 1:nrow(peaks_mba)){
#plot_lod_chr_mb(out,gm$pmap, peaks_mba$chr[i])
plot_lod_chr_mb(out,gm$pmap,i)
}
} else {
print(paste0("There are too many peaks (",nrow(peaks_mba)," peaks) that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}[1] “There are too many peaks (0 peaks) that have a LOD that reaches suggestive (p<0.05) level of 7.67594187344621 [autosomes]/9.24042402889532 [x-chromosome]”
QTL effects
For each peak LOD location we give a list of gene
if(nrow(peaks) < 50 & nrow(peaks) > 0){
for (i in 1:nrow(peaks)){
#for (i in 1:1){
#Plot 1
g <- maxmarg(pr.qc, gm$gmap, chr=peaks$chr[i], pos=peaks$pos[i], return_char=TRUE)
#png(filename=paste0("/Users/chenm/Documents/qtl/Jai/","qtl_effect_", i, ".png"))
#par(mar=c(4.1, 4.1, 1.5, 0.6))
plot_pxg(g, covars[,peaks$phenotype[i]], ylab=peaks$phenotype[i], sort=FALSE)
title(main = paste0("chr: ", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$phenotype[i]," )"), line=0.2)
##dev.off()
chr = peaks$chr[i]
# Plot 2
pr_sub <- pull_genoprobint(pr.qc, gm$gmap, chr, c(peaks$ci_lo[i], peaks$ci_hi[i]))
#coeff <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]], addcovar = addcovar)
#coeff <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]], Xcovar=Xcovar)
#coeff <- scan1coef(pr.qc[,chr], gm$covar[peaks$lodcolumn[i]], model="binary")
#coeff_sub <- scan1coef(pr_sub[,chr], gm$covar[peaks$lodcolumn[i]], model="binary")
blup <- scan1blup(pr.qc[,chr], covars[peaks$phenotype[i]])
blup_sub <- scan1blup(pr_sub[,chr], covars[peaks$phenotype[i]])
write.csv(as.data.frame(blup_sub), paste0("data/ici.vs.eoi_",peaks$phenotype[i],"-interactive.covariate_blup_sub_chr-",chr,"_peak.marker-",peaks$marker[i],"_lod.drop-1.5_5.batches_52.csv"), quote=F)
#plot_coef(coeff,
# gm$gmap, columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$lodcolumn[i]," [scan1coeff; positions in cM] )")
# )
#plot_coef(coeff_sub,
# gm$gmap, columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$lodcolumn[i],"; 1.5 LOD drop interval [scan1coeff; positions in cM] ) ")
# )
plot_coef(blup,
gm$gmap, columns=1:2,
bgcolor="gray95", legend="bottomleft",
main = paste0("chr: ", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$phenotype[i]," [scan1blup; positions in cM] )")
)
plot_coef(blup_sub,
gm$gmap, columns=1:2,
bgcolor="gray95", legend="bottomleft",
main = paste0("chr: ", chr=peaks$chr[i],"; pos: ", peaks$pos[i], "cM /",peaks_mba$pos[i],"MB\n(",peaks$phenotype[i],"; 1.5 LOD drop interval [scan1blup; positions in cM] )")
)
# Plot 3
#c2effB <- scan1coef(pr.qc[,chr], gm$covar[peaks$lodcolumn[i]], model="binary", contrasts=cbind(a=c(-1, 0), d=c(0, -1)))
#c2effBb <- scan1blup(pr.qc[,chr], gm$covar[peaks$lodcolumn[i]], contrasts=cbind(a=c(-1, 0), d=c(0, -1)))
##c2effB <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]], addcovar = addcovar, contrasts=cbind(mu=c(1,1,1), a=c(-1, 0, 1), d=c(0, 1, 0)))
##c2effB <- scan1coef(pr[,chr], cross$pheno[,peaks$lodcolumn[i]],Xcovar=Xcovar, contrasts=cbind(mu=c(1,1,1), a=c(-1, 0, 1), d=c(0, 1, 0)))
#plot(c2effB, gm$gmap[chr], columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i], "pos", peaks$pos[i], "(",peaks$lodcolumn[i],")")
# )
#plot(c2effBb, gm$gmap[chr], columns=1:2,
# bgcolor="gray95", legend="bottomleft",
# main = paste("chr", chr=peaks$chr[i], "pos", peaks$pos[i], "(",peaks$lodcolumn[i],")")
# )
##last_coef <- unclass(c2effB)[nrow(c2effB),2:3] # last two coefficients
##for(t in seq(along=last_coef))
## axis(side=4, at=last_coef[t], names(last_coef)[t], tick=FALSE)
#Table 1
chr = peaks_mba$chr[i]
start=as.numeric(peaks_mba$ci_lo[i])
end=as.numeric(peaks_mba$ci_hi[i])
genesgss = query_genes(chr, start, end)
write.csv(genesgss, file=paste0("data/ici.vs.eoi_",peaks$phenotype[i],"-interactive.covariate_genes_chr-",chr,"_peak.marker-",peaks$marker[i],"_lod.drop-1.5_5.batches_52.csv"), quote=F)
rownames(genesgss) <- NULL
genesgss$strand_old = genesgss$strand
genesgss$strand[genesgss$strand=="+"] <- "positive"
genesgss$strand[genesgss$strand=="-"] <- "negative"
#genesgss <-
#table <-
#genesgss[,c("chr","type","start","stop","strand","ID","Name","Dbxref","gene_id","mgi_type","description")] %>%
#kable(escape = F,align = c("ccccccccccc")) %>%
#kable_styling("striped", full_width = T) #%>%
#cat #%>%
#column_spec(1, bold=TRUE)
#
#print(kable(genesgss[,c("chr","type","start","stop","strand","ID","Name","Dbxref","gene_id","mgi_type","description")], escape = F,align = c("ccccccccccc")))
print(kable(genesgss[,c("chr","type","start","stop","strand","ID","Name","Dbxref","gene_id","mgi_type","description")], "html") %>% kable_styling("striped", full_width = T))
#table
}
} else {
print(paste0("There are too many peaks (",nrow(peaks_mba)," peaks) that have a LOD that reaches suggestive (p<0.05) level of ",summary(operm,alpha=0.05)$A, " [autosomes]/",summary(operm,alpha=0.05)$X, " [x-chromosome]"))
}[1] “There are too many peaks (0 peaks) that have a LOD that reaches suggestive (p<0.05) level of 7.67594187344621 [autosomes]/9.24042402889532 [x-chromosome]”
R version 3.6.2 (2019-12-12)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS Catalina 10.15.7
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRlapack.dylib
locale:
[1] en_AU.UTF-8/en_AU.UTF-8/en_AU.UTF-8/C/en_AU.UTF-8/en_AU.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] abind_1.4-5 qtl2_0.22 reshape2_1.4.4 ggplot2_3.3.5
[5] tibble_3.1.2 psych_2.0.7 readxl_1.3.1 cluster_2.1.0
[9] dplyr_1.0.8 optparse_1.6.6 rhdf5_2.28.1 mclust_5.4.6
[13] tidyr_1.0.2 data.table_1.14.0 knitr_1.33 kableExtra_1.1.0
[17] workflowr_1.6.2
loaded via a namespace (and not attached):
[1] httr_1.4.1 bit64_4.0.5 viridisLite_0.4.0 assertthat_0.2.1
[5] highr_0.9 blob_1.2.1 cellranger_1.1.0 yaml_2.2.1
[9] pillar_1.6.1 RSQLite_2.2.7 backports_1.2.1 lattice_0.20-38
[13] glue_1.4.2 digest_0.6.27 promises_1.1.0 rvest_0.3.5
[17] colorspace_2.0-2 htmltools_0.5.1.1 httpuv_1.5.2 plyr_1.8.6
[21] pkgconfig_2.0.3 purrr_0.3.4 scales_1.1.1 webshot_0.5.2
[25] getopt_1.20.3 later_1.0.0 git2r_0.26.1 generics_0.0.2
[29] ellipsis_0.3.2 cachem_1.0.5 withr_2.4.2 cli_3.0.0
[33] mnormt_1.5-7 magrittr_2.0.1 crayon_1.4.1 memoise_2.0.0
[37] evaluate_0.14 fs_1.4.1 fansi_0.5.0 nlme_3.1-142
[41] xml2_1.3.1 tools_3.6.2 hms_0.5.3 lifecycle_1.0.1
[45] stringr_1.4.0 Rhdf5lib_1.6.3 munsell_0.5.0 compiler_3.6.2
[49] rlang_1.0.2 grid_3.6.2 rstudioapi_0.13 rmarkdown_2.1
[53] gtable_0.3.0 DBI_1.1.1 R6_2.5.0 fastmap_1.1.0
[57] bit_4.0.4 utf8_1.2.1 rprojroot_1.3-2 readr_1.3.1
[61] stringi_1.7.2 parallel_3.6.2 Rcpp_1.0.7 vctrs_0.3.8
[65] tidyselect_1.1.2 xfun_0.24